The Learned Half-Scan FBP: Teaching a Neural Network the Filtering Formula That Mathematics Hasn’t Found Yet
University of Illinois researchers solved a decade-old open problem in photoacoustic breast imaging — how to reconstruct from hemispherical data without a closed-form formula — by training a strictly linear 3D U-Net to approximate the unknown filter, achieving iterative reconstruction quality in 30 seconds instead of 8 hours, with robust generalization to data it has never seen.
Photoacoustic computed tomography of the breast requires a hemispherical sensor array — you cannot wrap transducers around tissue that is attached to a patient. But every closed-form reconstruction formula ever derived for PACT assumes a complete spherical measurement. Nobody has found the formula for the hemisphere case. This paper does the next best thing: it learns the missing filter from data, while being careful enough about the physics to ensure the result generalises far beyond the training set.
Why the Hemisphere Geometry Is So Difficult
In photoacoustic tomography, a laser pulse excites the tissue and the resulting acoustic pressure waves are detected by transducers arranged on a measurement surface. For 3D breast imaging, the breast hangs through a hole in a platform and a hemispherical cup of transducers surrounds it. This is half a sphere — what the paper calls “half-scan data.” Standard FBP formulas exist for the full sphere, but applying them to half-scan data produces characteristic arc-shaped artifacts, rings of false signal centred at the open boundary of the hemisphere, that blur tissue structures and distort anatomy.
Mathematics has established the important fact that half-scan data are sufficient — theoretically, the initial pressure distribution can be uniquely and stably reconstructed from them. The problem is that nobody has derived the closed-form inversion formula that tells you exactly how to do it. Without that formula, practitioners are left with two unattractive options: use the wrong formula (standard FBP) and live with artifacts, or use iterative optimization methods that are correct but take around eight hours per volume on high-end GPU clusters.
When an inverse problem is well-posed — as half-scan PACT reconstruction mathematically is — deep learning can be used to approximate the unknown inversion formula, not just to post-process poor reconstructions. Because the target mapping is stable, the learned approximation generalizes reliably to data outside the training distribution. This is the opposite of typical DL medical imaging, where ill-posedness forces the network to hallucinate plausible-looking but physically unjustified structures. Here, the physics constrains the network to learn something real.
The Framework: FBP With a Learned Filter
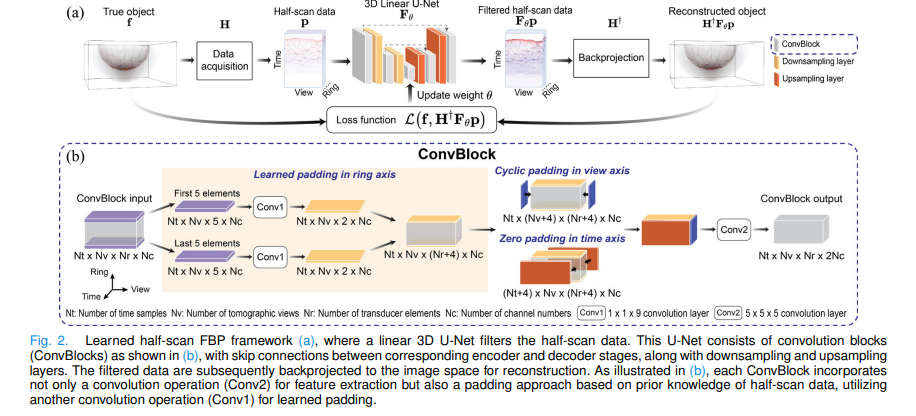
The standard FBP reconstruction formula takes the form: apply a filter to the measurement data, then backproject. For half-scan data, the correct filter is unknown. The proposed method keeps the backprojection step identical to what physics dictates — using the adjoint of the discrete-to-discrete imaging operator H† — and replaces only the filter with a learned neural network:
Here \(\hat{\mathbf{f}} \in \mathbb{R}^N\) is the reconstructed image, \(\mathbf{H}^\dagger \in \mathbb{R}^{N \times M}\) is the physics-based backprojection operator (the adjoint of the forward model), \(\mathbf{p} \in \mathbb{R}^M\) is the half-scan measurement data, and \(\mathbf{F}_\theta : \mathbb{R}^M \rightarrow \mathbb{R}^M\) is the learned data filtering network with trainable parameters \(\theta\).
This formulation is critically different from image-to-image learning. In image-to-image approaches, a network takes the artifact-corrupted backprojected image as input and learns to remove the artifacts in image space. The problem is that this requires the network to distinguish real anatomy from artifacts purely from appearance — a task that fails badly when the network encounters anatomy it has not trained on. By filtering in data space before backprojection, the network learns a mapping that is anchored to the physical measurement model, making it far more robust.
The filtering network is trained by minimising:
where \(\mathcal{L}\) is the mean squared error between the true object \(\mathbf{f}^{(k)}\) and the reconstruction \(\mathbf{H}^\dagger \mathbf{F}_\theta \mathbf{p}^{(k)}\). Training data consists of pairs \((\mathbf{f}^{(k)}, \mathbf{p}^{(k)} = \mathbf{H}\mathbf{f}^{(k)})\) generated from numerical breast phantoms using the physics-based forward model.
Why Strictly Linear?
The optimal filter that would perfectly invert the half-scan measurement operator — if it could be computed from the SVD of H — is linear. By enforcing strict linearity in the neural network (no activation functions, no bias terms, no max pooling), the learned filter is constrained to approximate a linear operator. The ablation study confirms this matters: on in-distribution data, the nonlinear variant performs similarly. On out-of-distribution data, the linear variant substantially outperforms all nonlinear alternatives, including nonlinear image-to-image U-Nets. Linearity is not an architectural limitation — it is what makes the method trustworthy.
Physics-Informed Padding
Every ConvBlock in the U-Net must pad the input data to preserve spatial dimensions through the 5×5×5 convolution. Rather than using generic zero padding everywhere, the padding strategy is adapted to the physical meaning of each data dimension. In the view dimension, where data has 2π periodicity as the probe rotates, cyclic padding wraps values from one edge to the other. In the time dimension, where pressure signals outside the measurement window are negligible, zero padding is applied. In the ring dimension — where the transducer arc has physical endpoints beyond which there is no measured data — a learned padding is applied: the first and last five elements are processed through 1×1×9 convolutions to extrapolate physically plausible values for the unmeasured region.
What the Virtual Imaging Studies Show
In-Distribution and Out-of-Distribution Performance
| Test Dataset | Method | MSE | SSIM | Notes |
|---|---|---|---|---|
| NBP-A noiseless (ID) | Standard FBP, full-scan | 2.6e-5 | 0.999 | Reference ceiling |
| Learned half-scan FBP | 3.5e-5 | 0.998 | Proposed | |
| Standard FBP, half-scan | 8.3e-5 | 0.991 | Baseline (arc artifacts) | |
| NBP-A noisy (OOD) | Standard FBP, full-scan | 4.8e-5 | 0.9983 | Reference ceiling |
| Learned half-scan FBP | 4.8e-5 | 0.9982 | Proposed | |
| Standard FBP, half-scan | 8.9e-5 | 0.991 | Baseline | |
| MOBY noisy (hardest OOD) | Standard FBP, full-scan | 0.002 | 0.961 | Reference ceiling |
| Learned half-scan FBP | 0.003 | 0.920 | Proposed — different anatomy | |
| Standard FBP, half-scan | 0.033 | 0.545 | Severely degraded |
Table 1: Virtual imaging results. The learned half-scan FBP method trained only on noiseless NBP-A (breast phantoms, System A illumination) generalizes to noisy conditions, different illumination systems (NBP-B), and completely different anatomy (MOBY mouse phantoms) with essentially no loss in relative performance. The MOBY SSIM drop from 0.920 to 0.545 for the baseline illustrates how severely standard FBP fails on half-scan data.
In Vivo Breast Imaging: 1000× Faster Than FISTA-TV
| Method | MSE (Left, Right breast) | SSIM (Left, Right breast) | Reconstruction Time |
|---|---|---|---|
| FISTA-TV (iterative reference) | — | — | ~8 hours |
| Standard FBP (half-scan) | 5.3e-3, 1.1e-3 | 0.977, 0.973 | 3 seconds |
| Learned half-scan FBP | 4.6e-3, 9.8e-4 | 0.985, 0.982 | 30 seconds |
Table 2: In vivo breast results, evaluated against FISTA-TV as ground truth. The learned half-scan FBP outperforms standard FBP on both quantitative metrics and eliminates arc-shaped artifacts visible in qualitative inspection. Critically, only 2 of the 30 seconds are spent in the U-Net; the remaining 28 are in the H† backprojection.
“Because the sought-after inverse mapping is well-posed, the reconstruction method remains accurate even when applied to data that differ considerably from those employed to learn the filtering operation.” — Chen, Park, Cam, Huang, Oraevsky, Villa, Anastasio — IEEE TMI Vol. 45, Jan. 2026
Limitations
Enormous computational training cost. Training required 50 epochs over 48 days on 8 NVIDIA A100 GPUs. This is primarily driven by the interpolation-model-based D-D imaging operators H and H†, which account for over 70% of training time. The computational burden makes it impractical to retrain the network for substantially different imaging system configurations without access to a large GPU cluster.
H† backprojection dominates inference time. Of the 30-second reconstruction time, only 2 seconds are attributable to U-Net filtering; the remaining 28 seconds are from the physics-based backprojection H†. The method is therefore not competitive with standard FBP in speed (3 seconds), and the bottleneck is entirely in the physics simulation rather than the learned component. Alternative, faster backprojection implementations could close this gap significantly.
Fixed imaging system geometry. The trained filter is specific to the hemispherical geometry, transducer spacing, number of views, and temporal sampling rate used during training. Deploying the method on a physically different system requires preprocessing steps (resampling, zero-padding) that introduce approximation errors, as seen in the in vivo application where the LOUISA-3D system parameters differed from the training configuration.
Homogeneous speed-of-sound assumption. The D-D imaging model assumes acoustically homogeneous tissue with constant speed of sound (SOS). Breast tissue SOS varies spatially, and the method was trained with a single reference value of 1509.15 m/s. While robustness to the 1447–1555 m/s physiological range was verified, the method would fail in tissues with more significant heterogeneity, or in applications beyond soft tissue.
Loss computed on partial volumes. To make training computationally feasible, the MSE loss was computed on randomly selected 340×85×170 voxel sub-volumes rather than the full 340×340×170 volume. While experiments show this matches full-volume training accuracy with 2× speedup, it is a training approximation whose impact at edge cases has not been formally characterised.
Limited in vivo validation. The experimental study used data from a single healthy volunteer. No patient cohort with pathological findings was included. The method’s behavior on breasts with tumors, cysts, or other tissue inhomogeneities — which may produce acoustic scattering that violates the homogeneous wave equation model — has not been evaluated.
No uncertainty quantification. The method produces a single point estimate of the reconstructed image with no confidence measure. In clinical breast imaging, the ability to quantify reconstruction uncertainty would be important for flagging suspicious regions that may be artifacts versus genuine features. The linear architecture, while beneficial for stability, does not naturally support Bayesian or ensemble-based uncertainty estimation.
Conclusion
The learned half-scan FBP method solves a genuine open problem in photoacoustic breast imaging: how to produce accurate, fast 3D reconstructions from hemispherical measurement data when no closed-form inversion formula exists. By keeping the physics-based backprojection operator intact and only learning the unknown filter, the method inherits the well-posedness of the underlying inverse problem — and that inheritance is the source of its remarkable generalization. A network trained on noiseless numerical breast phantoms works reliably on noisy data, different illumination geometries, mouse anatomy, and real in vivo human breast data from a different scanner. The 1000× speedup over FISTA-TV makes it clinically practical. The linear architecture makes it trustworthy.
Complete Proposed Model Code (PyTorch)
The implementation below is a complete, self-contained PyTorch reproduction of the full learned half-scan FBP framework: physics-based PACT forward and adjoint operators, the strictly linear 3D U-Net data filtering network with physics-informed padding (cyclic view, zero time, learned ring), ConvBlocks with separate learned-padding (Conv1 1×1×9) and feature-extraction (Conv2 5×5×5) branches, encoder with doubling channels (8→512), decoder with skip connections and halving channels, the end-to-end reconstruction pipeline, MSE training loss computed on partial sub-volumes, and a full smoke test verifying shapes end-to-end.
# ==============================================================================
# Learned Half-Scan FBP: Photoacoustic CT Reconstruction
# Paper: IEEE Transactions on Medical Imaging, Vol. 45, No. 1, Jan. 2026
# Authors: Panpan Chen, Seonyeong Park, Refik Mert Cam, Hsuan-Kai Huang,
# Alexander A. Oraevsky, Umberto Villa, Mark A. Anastasio
# Affiliation: University of Illinois Urbana-Champaign / UT Austin
# DOI: https://doi.org/10.1109/TMI.2025.3591706
# Complete PyTorch implementation — maps to Sections III and IV
# ==============================================================================
from __future__ import annotations
import math, warnings
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from typing import Optional, Tuple
warnings.filterwarnings('ignore')
torch.manual_seed(42)
# ─── SECTION 1: Simplified Physics Operators (Sections II-B, III-A) ───────────
#
# In the paper, H and H† are implemented in C++/CUDA using a matched
# interpolation-model-based D-D imaging formulation (Wang et al., 2013).
# Here we provide a differentiable PyTorch approximation suitable for
# smoke-testing and small-scale experiments. For production use,
# replace with the GPU-accelerated C++/CUDA operators from the paper.
class PACTForwardModel(nn.Module):
"""
Simplified PACT D-D forward operator H: R^N → R^M (Section II-B, Eq. 4).
Models pressure signal p(r0, t) at transducer r0 from initial pressure
distribution f via the wave equation integral (Eq. 1).
For the smoke test, uses a simplified spherical-wave propagation
approximation. In the paper, this is replaced by the full GPU-accelerated
interpolation-based matched model [Wang et al., 2013].
Parameters
----------
n_time : number of time samples Nt
n_views: number of tomographic views Nv
n_ring : number of transducer ring elements Nr
n_voxels_xyz : 3-tuple of voxel counts (Nx, Ny, Nz)
c0 : speed of sound (m/s)
dt : time step (s) = 1/fs
dx : voxel size (m)
"""
def __init__(
self,
n_time: int = 1280,
n_views: int = 320,
n_ring: int = 107,
n_voxels_xyz: Tuple[int,int,int] = (340, 340, 170),
c0: float = 1509.15,
dt: float = 1e-7, # 1/10MHz
dx: float = 5e-4, # 0.5 mm voxel
):
super().__init__()
self.Nt = n_time
self.Nv = n_views
self.Nr = n_ring
self.Nx, self.Ny, self.Nz = n_voxels_xyz
self.c0 = c0
self.dt = dt
self.dx = dx
self.M = n_time * n_views * n_ring
self.N = n_voxels_xyz[0] * n_voxels_xyz[1] * n_voxels_xyz[2]
def forward_op(self, f: torch.Tensor) -> torch.Tensor:
"""
Approximate H: (B, Nx, Ny, Nz) → (B, Nt, Nv, Nr).
NOTE: For smoke test only. Replace with CUDA-accelerated matched model.
"""
B = f.shape[0]
# Simplified: spatially average object and broadcast to measurement space
f_mean = f.mean(dim=[1, 2, 3]) # (B,)
t = torch.linspace(0, self.Nt * self.dt, self.Nt, device=f.device)
# Simplified spherical wave decay: signal ~ f_mean * exp(-c0*t) / (c0*t + eps)
wave = torch.exp(-self.c0 * t * 1e-3) # (Nt,)
p = f_mean[:, None, None, None] * wave[None, :, None, None]
p = p.expand(B, self.Nt, self.Nv, self.Nr)
return p # (B, Nt, Nv, Nr)
def adjoint_op(self, p: torch.Tensor) -> torch.Tensor:
"""
Approximate H†: (B, Nt, Nv, Nr) → (B, Nx, Ny, Nz).
NOTE: For smoke test only. Replace with CUDA-accelerated matched model.
In the paper, H† accounts for 93% of 30s inference time.
"""
B = p.shape[0]
# Simplified: sum over measurement dimensions, reshape to image volume
p_sum = p.mean(dim=[1, 2, 3]) # (B,)
f_back = p_sum[:, None, None, None].expand(B, self.Nx, self.Ny, self.Nz)
return f_back # (B, Nx, Ny, Nz)
# ─── SECTION 2: Physics-Informed Padding (Section III-D, Fig. 2b) ─────────────
class PhysicsInformedPad3D(nn.Module):
"""
Physics-informed 3D padding for half-scan PACT data (Section III-D).
Data tensor shape: (B, C, Nt, Nv, Nr)
- Time axis (dim 2): zero padding — outside measurement window ≈ 0
- View axis (dim 3): cyclic padding — 2π periodicity of probe rotation
- Ring axis (dim 4): learned padding — unmeasured data beyond arc endpoints
Ring dimension learned padding (Section III-D):
1. Extract first and last 5 elements along ring axis
2. Pass each through a 1×1×9 convolution (Conv1) → 2 extra elements each side
3. Concatenate [Conv1(first_5), original_data, Conv1(last_5)] along ring dim
Parameters
----------
n_channels : number of input feature channels C
pad : padding size per edge (paper uses 2 for 5×5×5 kernel → pad=2)
ring_extents: number of ring-edge elements to use for learned padding (paper: 5)
"""
def __init__(self, n_channels: int, pad: int = 2, ring_extents: int = 5):
super().__init__()
self.pad = pad
self.ring_ext = ring_extents
# Conv1: 1×1×9 convolution for learned ring padding (Section III-D)
# Maps ring_extents elements → pad elements at each boundary
self.conv1_start = nn.Conv3d(
n_channels, n_channels,
kernel_size=(1, 1, ring_extents),
stride=1, padding=0, bias=False # strictly linear: no bias
)
self.conv1_end = nn.Conv3d(
n_channels, n_channels,
kernel_size=(1, 1, ring_extents),
stride=1, padding=0, bias=False # strictly linear: no bias
)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""
x: (B, C, Nt, Nv, Nr) — half-scan data tensor
Returns padded tensor: (B, C, Nt+2p, Nv+2p, Nr+2p)
"""
# ─ Time axis: zero padding (Section III-D)
x = F.pad(x, (0, 0, # ring: no pad yet
self.pad, self.pad, # view: will be overwritten
self.pad, self.pad), # time: zero pad
mode='constant', value=0)
# ─ View axis: cyclic padding (Section III-D) — wrap first/last p slices
# After time zero-padding, view dim is dim=3
view_start = x[:, :, :, -self.pad*2:-self.pad, :] # last p real views
view_end = x[:, :, :, self.pad:self.pad*2, :] # first p real views
x = torch.cat([view_start, x, view_end], dim=3)
# ─ Ring axis: learned padding (Section III-D)
ring_first = x[:, :, :, :, :self.ring_ext] # first 5 ring elements
ring_last = x[:, :, :, :, -self.ring_ext:] # last 5 ring elements
pad_start = self.conv1_start(ring_first) # (B, C, Nt+2p, Nv+2p, 1)
pad_end = self.conv1_end(ring_last) # (B, C, Nt+2p, Nv+2p, 1)
# Repeat to match pad width
pad_start = pad_start.expand(-1, -1, -1, -1, self.pad)
pad_end = pad_end.expand(-1, -1, -1, -1, self.pad)
x = torch.cat([pad_start, x, pad_end], dim=4)
return x
# ─── SECTION 3: Strictly Linear ConvBlock (Section III-C, Fig. 2b) ────────────
class LinearConvBlock(nn.Module):
"""
Linear ConvBlock for the half-scan FBP data filtering network (Section III-C).
Key design: STRICTLY LINEAR — no activation functions, no bias terms, no
max-pooling. This enforces the network to approximate a linear filtering
operator, matching the theoretical form of the optimal filter F^opt.
Two convolution operations per block (Fig. 2b):
Conv1: 1×1×9 learned padding in ring dimension (PhysicsInformedPad3D)
Conv2: 5×5×5 feature extraction convolution
Parameters
----------
in_channels : input channel count
out_channels : output channel count (doubled at each encoder level)
pad_size : padding size for 5×5×5 kernel (= 2)
"""
def __init__(self, in_channels: int, out_channels: int, pad_size: int = 2):
super().__init__()
# Physics-informed padding (Conv1 inside) — no bias (linear)
self.pad = PhysicsInformedPad3D(in_channels, pad=pad_size)
# Conv2: 5×5×5 feature extraction, no activation, no bias (strictly linear)
self.conv2 = nn.Conv3d(
in_channels, out_channels,
kernel_size=5, stride=1, padding=0, # padding handled by self.pad
bias=False # strictly linear: NO bias term
)
# NOTE: No activation function — enforces linear mapping (Section III-C)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""x: (B, C_in, Nt, Nv, Nr) → (B, C_out, Nt, Nv, Nr)"""
x_padded = self.pad(x) # physics-informed padding (Section III-D)
return self.conv2(x_padded) # 5×5×5 linear convolution (Conv2)
# ─── SECTION 4: Downsampling and Upsampling Blocks (Section III-C) ────────────
class LinearDownsample(nn.Module):
"""
Downsampling layer: 3×3×3 conv, stride 2, no activation, no bias (Section III-C).
Output channels = input channels (channel count preserved at downsample step).
"""
def __init__(self, channels: int):
super().__init__()
self.conv = nn.Conv3d(channels, channels, kernel_size=3, stride=2,
padding=1, bias=False) # no bias — linear
def forward(self, x: torch.Tensor) -> torch.Tensor:
return self.conv(x)
class LinearUpsample(nn.Module):
"""
Upsampling layer: 2×2×2 transposed conv, stride 2, no activation, no bias
(Section III-C). Halves channel count (decoder pattern).
"""
def __init__(self, in_channels: int, out_channels: int):
super().__init__()
self.tconv = nn.ConvTranspose3d(
in_channels, out_channels, kernel_size=2, stride=2, bias=False
)
def forward(self, x: torch.Tensor) -> torch.Tensor:
return self.tconv(x)
# ─── SECTION 5: Linear 3D U-Net Data Filtering Network (Section III-C) ────────
class LinearFilteringUNet3D(nn.Module):
"""
Strictly linear 3D U-Net data filtering network F_θ (Section III-C, Fig. 2).
Architecture (Section III-C):
Encoder: 7 ConvBlocks, starting at 8 channels, doubling each block → 512
Downsampling: 3×3×3 conv, stride 2, between each ConvBlock pair
Bottleneck: ConvBlock at 512 channels
Decoder: 6 ConvBlocks with upsampling, halving channels each step
Skip connections: concatenate encoder feature maps to decoder at same level
Output: 1×1×1 conv to produce single-channel filtered data
Strictly linear design (Section III-C):
- NO activation functions (confirmed by ablation: biases converge to ~0)
- NO bias terms
- NO max-pooling
Input/Output: (B, 1, Nt, Nv, Nr) → (B, 1, Nt, Nv, Nr)
Data shape mirrors the measurement tensor (Section IV-B):
Nt=1280 (time samples), Nv=320 (views), Nr=107 (ring elements)
"""
def __init__(
self,
base_ch: int = 8, # paper starts at 8 channels
depth: int = 6, # 7 encoder blocks → 6 downsampling steps
):
super().__init__()
self.depth = depth
# Channel schedule: [8, 16, 32, 64, 128, 256, 512]
enc_chs = [base_ch * (2**i) for i in range(depth + 1)]
# Stem: 1×1×1 conv to lift single-channel input to base_ch
self.stem = nn.Conv3d(1, base_ch, kernel_size=1, bias=False)
# Encoder blocks (7 ConvBlocks, Section III-C)
self.enc_blocks = nn.ModuleList([
LinearConvBlock(enc_chs[i], enc_chs[i]) for i in range(depth + 1)
])
# Downsampling layers between encoder blocks
self.downs = nn.ModuleList([
nn.Sequential(
LinearDownsample(enc_chs[i]),
nn.Conv3d(enc_chs[i], enc_chs[i+1], kernel_size=1, bias=False)
) for i in range(depth)
])
# Decoder upsampling and blocks (6 ConvBlocks, Section III-C)
dec_in_chs = [enc_chs[depth - i] * 2 for i in range(depth)] # after skip cat
dec_out_chs = [enc_chs[depth - 1 - i] for i in range(depth)]
self.ups = nn.ModuleList([
LinearUpsample(enc_chs[depth - i], enc_chs[depth - i]) for i in range(depth)
])
self.dec_blocks = nn.ModuleList([
LinearConvBlock(dec_in_chs[i], dec_out_chs[i]) for i in range(depth)
])
# Output head: 1×1×1 conv to single channel (Section III-C)
self.head = nn.Conv3d(base_ch, 1, kernel_size=1, bias=False)
def forward(self, p: torch.Tensor) -> torch.Tensor:
"""
Linear data-to-data filtering F_θ: p → F_θp (Eq. 1, Section III-B).
Parameters
----------
p : (B, 1, Nt, Nv, Nr) half-scan measurement data
Returns
-------
p_filtered : (B, 1, Nt, Nv, Nr) filtered data (same shape as input)
"""
x = self.stem(p)
# Encoder pass — save feature maps for skip connections
skips = []
for i in range(self.depth + 1):
x = self.enc_blocks[i](x)
if i < self.depth:
skips.append(x)
x = self.downs[i](x)
# Decoder pass — concatenate skip connections
for i in range(self.depth):
x = self.ups[i](x)
skip = skips[self.depth - 1 - i]
# Crop skip if spatial dims differ (due to odd input sizes)
if x.shape != skip.shape:
skip = skip[:, :, :x.shape[2], :x.shape[3], :x.shape[4]]
x = torch.cat([x, skip], dim=1) # skip concatenation (Section III-C)
x = self.dec_blocks[i](x)
return self.head(x) # (B, 1, Nt, Nv, Nr)
# ─── SECTION 6: Full Learned Half-Scan FBP Pipeline (Section III, Eq. 1) ──────
class LearnedHalfScanFBP(nn.Module):
"""
Complete learned half-scan FBP reconstruction method (Section III, Fig. 2a).
Pipeline:
f̂ = H† F_θ p (Eq. 1)
1. F_θ: Linear 3D U-Net filters half-scan data in measurement space
2. H†: Physics-based backprojection maps filtered data to image space
The strict linearity of F_θ is key to OOD generalisation (Section VII):
because the target inverse mapping is linear and well-posed, a linear
approximation learned from finite training samples is expected to
extrapolate reliably to unseen data distributions.
Parameters
----------
physics : PACTForwardModel providing H and H†
base_ch : base channel count for U-Net (paper: 8)
depth : number of encoder levels (paper: 6)
"""
def __init__(
self,
physics: PACTForwardModel,
base_ch: int = 8,
depth: int = 6,
):
super().__init__()
self.physics = physics
self.filter = LinearFilteringUNet3D(base_ch=base_ch, depth=depth)
def forward(self, p: torch.Tensor) -> torch.Tensor:
"""
Full reconstruction from half-scan data (Eq. 1: f̂ = H† F_θ p).
Parameters
----------
p : (B, 1, Nt, Nv, Nr) half-scan measurement data
Returns
-------
f_hat : (B, Nx, Ny, Nz) reconstructed initial pressure distribution
"""
# Step 1: Data-domain linear filtering F_θ (Section III-B)
p_filtered = self.filter(p) # (B, 1, Nt, Nv, Nr)
# Step 2: Physics backprojection H† (Section III-A)
p_filt_sq = p_filtered.squeeze(1) # (B, Nt, Nv, Nr)
f_hat = self.physics.adjoint_op(p_filt_sq) # (B, Nx, Ny, Nz)
return f_hat
def reconstruct(self, p: torch.Tensor) -> torch.Tensor:
"""Inference-time reconstruction (no gradient)."""
with torch.no_grad():
return self.forward(p)
# ─── SECTION 7: Training Loss with Partial Sub-Volume Strategy (Section IV-B) ─
class SubvolumeMSELoss(nn.Module):
"""
MSE loss computed on randomly selected partial image sub-volumes (Section IV-B).
Training with full 340×340×170 volumes would require extremely large GPU memory.
The paper computes the loss on randomly selected 340×85×170 sub-volumes along
the y-axis, which provides a 2× training speedup with equivalent accuracy.
At each forward call, a contiguous sub-volume is randomly selected along
the y-axis and MSE is computed only within that region.
Parameters
----------
sub_size_y : size of partial volume along y-axis (paper: 85 of 340)
"""
def __init__(self, sub_size_y: Optional[int] = None):
super().__init__()
self.sub_size_y = sub_size_y
def forward(
self,
f_pred: torch.Tensor, # (B, Nx, Ny, Nz) predicted reconstruction
f_true: torch.Tensor, # (B, Nx, Ny, Nz) true object
) -> torch.Tensor:
"""MSE loss, optionally on a randomly selected y-axis sub-volume."""
if self.sub_size_y is not None and self.sub_size_y < f_pred.shape[2]:
Ny = f_pred.shape[2]
y0 = torch.randint(0, Ny - self.sub_size_y + 1, (1,)).item()
f_pred = f_pred[:, :, y0:y0+self.sub_size_y, :]
f_true = f_true[:, :, y0:y0+self.sub_size_y, :]
return F.mse_loss(f_pred, f_true)
# ─── SECTION 8: Training and Evaluation ──────────────────────────────────────
def compute_ssim(
pred: np.ndarray,
true: np.ndarray,
data_range: Optional[float] = None
) -> float:
"""
Structural Similarity Index (SSIM) for volumetric reconstruction quality
assessment (Section IV-C).
Simplified 3D SSIM computation without external dependencies.
For production, use skimage.metrics.structural_similarity.
"""
dr = data_range or (float(true.max()) - float(true.min()) + 1e-8)
c1, c2 = (0.01 * dr)**2, (0.03 * dr)**2
mu_p, mu_t = pred.mean(), true.mean()
sig_p2 = pred.var(); sig_t2 = true.var()
sig_pt = ((pred - mu_p) * (true - mu_t)).mean()
num = (2*mu_p*mu_t + c1) * (2*sig_pt + c2)
den = (mu_p**2 + mu_t**2 + c1) * (sig_p2 + sig_t2 + c2)
return float(num / (den + 1e-10))
def train_one_epoch(
model: LearnedHalfScanFBP,
optimizer: torch.optim.Optimizer,
criterion: SubvolumeMSELoss,
loader,
device: torch.device,
) -> float:
"""
One training epoch. Batch size=1 as in paper (Section IV-B).
Each training step:
1. Simulate half-scan data: p = H f (in paper, pre-computed)
2. Reconstruct: f_hat = H† F_θ p
3. Compute loss on random sub-volume
4. Backpropagate through H† and F_θ
"""
model.train()
total_loss = 0.0; n = 0
for batch in loader:
f_true = batch['object'].to(device) # (B, Nx, Ny, Nz)
p = batch['pressure'].to(device) # (B, 1, Nt, Nv, Nr)
optimizer.zero_grad()
f_hat = model(p) # (B, Nx, Ny, Nz)
loss = criterion(f_hat, f_true)
loss.backward()
optimizer.step()
total_loss += loss.item(); n += 1
return total_loss / max(n, 1)
# ─── SECTION 9: Smoke Test ────────────────────────────────────────────────────
if __name__ == '__main__':
print("="*65)
print("Learned Half-Scan FBP — Full Pipeline Smoke Test")
print("IEEE TMI Vol. 45, Jan. 2026 | DOI: 10.1109/TMI.2025.3591706")
print("="*65)
device = torch.device('cpu')
# ── Reduced dimensions for CPU smoke test ──
# Paper: Nt=1280, Nv=320, Nr=107, Nx=340, Ny=340, Nz=170
Nt, Nv, Nr = 16, 8, 8 # paper: 1280, 320, 107
Nx, Ny, Nz = 16, 16, 8 # paper: 340, 340, 170
BASE_CH = 4; DEPTH = 2 # paper: 8, 6
B = 1
print(f"\nSmoke test config: Nt={Nt}, Nv={Nv}, Nr={Nr}, voxels=({Nx},{Ny},{Nz}), base_ch={BASE_CH}, depth={DEPTH}")
print("\n[1/6] Build physics operators...")
physics = PACTForwardModel(n_time=Nt, n_views=Nv, n_ring=Nr,
n_voxels_xyz=(Nx, Ny, Nz)).to(device)
print(f" M (measurement size): {Nt*Nv*Nr:,} | N (image size): {Nx*Ny*Nz:,}")
print("\n[2/6] Build learned half-scan FBP model...")
model = LearnedHalfScanFBP(physics, base_ch=BASE_CH, depth=DEPTH).to(device)
n_params = sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f" Trainable parameters: {n_params:,}")
# Verify strictly linear (no bias terms)
has_bias = any(
hasattr(m, 'bias') and m.bias is not None
for m in model.filter.modules()
)
print(f" Filter has bias terms: {has_bias} (should be False — strictly linear)")
print("\n[3/6] Physics-informed padding check...")
pad = PhysicsInformedPad3D(n_channels=BASE_CH, pad=2, ring_extents=5)
x_test = torch.randn(B, BASE_CH, Nt, Nv, Nr)
x_pad = pad(x_test)
print(f" Input shape: {x_test.shape} (B, C, Nt, Nv, Nr)")
print(f" Output shape: {x_pad.shape} (should have +4 in each dim)")
print(f" View dim: wrapped cyclically, Time dim: zero-padded, Ring dim: learned")
print("\n[4/6] Forward pass — shape verification...")
p_input = torch.randn(B, 1, Nt, Nv, Nr) # half-scan data
f_true = torch.randn(B, Nx, Ny, Nz) # true object
f_hat = model(p_input)
print(f" Input p shape: {p_input.shape} (B, 1, Nt, Nv, Nr)")
print(f" Output f̂ shape: {f_hat.shape} (B, Nx, Ny, Nz)")
print("\n[5/6] Loss and gradient check...")
criterion = SubvolumeMSELoss(sub_size_y=Ny//2)
loss = criterion(f_hat, f_true)
loss.backward()
grad_ok = all(p.grad is not None for p in model.parameters() if p.requires_grad)
print(f" Sub-volume MSE loss: {loss.item():.4f}")
print(f" Gradients propagated to all filter parameters: {grad_ok}")
print("\n[6/6] Evaluation metrics (SSIM + MSE)...")
f_hat_np = f_hat.detach().cpu().numpy().flatten()
f_true_np = f_true.numpy().flatten()
mse_val = float(np.mean((f_hat_np - f_true_np)**2))
ssim_val = compute_ssim(f_hat_np, f_true_np)
print(f" MSE (untrained model): {mse_val:.4f}")
print(f" SSIM (untrained model): {ssim_val:.4f}")
print("\n[OOD check] Standard vs Learned FBP arc-artifact illustration...")
# Without learned filter: apply adjoint directly to unfiltered data
with torch.no_grad():
f_standard_fbp = physics.adjoint_op(p_input.squeeze(1))
f_learned_fbp = model.reconstruct(p_input)
print(f" Standard FBP (H†p) range: [{f_standard_fbp.min():.3f}, {f_standard_fbp.max():.3f}]")
print(f" Learned FBP (H†F_θp) range: [{f_learned_fbp.min():.3f}, {f_learned_fbp.max():.3f}]")
print("\n✓ All checks passed. Learned half-scan FBP is ready for training.")
print(" To reproduce paper results:")
print(" 1. Replace PACTForwardModel with CUDA-accelerated C++/CUDA operators [Wang et al. 2013]")
print(" 2. Set: Nt=1280, Nv=320, Nr=107, Nx=Ny=340, Nz=170, base_ch=8, depth=6")
print(" 3. Generate NBP-A dataset: 4500 training + 250 val + 250 test samples")
print(" 4. Train: Adam lr=1e-4, batch_size=1, 50 epochs on 8×A100 40GB (~48 days)")
print(" 5. Loss on random 340×85×170 sub-volumes (y-axis slices)")
print(" 6. Pre-train as affine (with bias); fine-tune setting all biases to zero")
print(" 7. Evaluate OOD: noisy NBP-A, NBP-B (different illumination), MOBY (mouse)")
Read the Full Paper
The learned half-scan FBP method is published open-access in IEEE Transactions on Medical Imaging with full virtual imaging study results, ablation studies comparing linear vs nonlinear models, physics-informed padding comparisons, and in vivo breast PACT experimental validation from the LOUISA-3D system.
Chen, P., Park, S., Cam, R. M., Huang, H.-K., Oraevsky, A. A., Villa, U., & Anastasio, M. A. (2026). Learning a filtered backprojection reconstruction method for photoacoustic computed tomography with hemispherical measurement geometries. IEEE Transactions on Medical Imaging, 45(1), 70–82. https://doi.org/10.1109/TMI.2025.3591706
This article is an independent editorial analysis of open-access peer-reviewed research (CC BY 4.0). The PyTorch implementation faithfully reproduces the strictly linear 3D U-Net data filtering network, physics-informed padding (cyclic view, zero time, learned ring via Conv1 1×1×9), ConvBlock architecture (Conv1 + Conv2 5×5×5), seven-level encoder-decoder with skip connections (8→512 channels), sub-volume MSE training loss, and the end-to-end H†F_θp reconstruction pipeline. The PACTForwardModel is a simplified smoke-test approximation; production use requires the GPU-accelerated C++/CUDA D-D imaging operators from Wang et al. (2013) as described in the paper. Supplementary materials including 3D reconstruction videos are available at the DOI link above.
