The Deep Learning System That Learned to Map Eloquent Brain Circuits from Electrical Stimulation and Anatomy
ESM-AnatTractNet integrates electrophysiological validation with anatomical context to accurately classify true positive white matter pathways — enabling safer pediatric epilepsy surgery through precise, non-invasive functional mapping.
Every millimeter matters when removing epileptic brain tissue from a child. The seizure onset zone — the network of neurons generating pathological electrical discharges — often sits perilously close to regions controlling movement, speech, or vision. Resect too conservatively, and seizures persist. Resect too aggressively, and the child wakes up unable to move a hand, form a sentence, or see half the visual world. The surgical dilemma is ancient, but the tools for resolving it have remained frustratingly imprecise.
Electrical stimulation mapping (ESM) has long been the clinical gold standard for localizing eloquent cortex. By applying brief electrical pulses through implanted subdural electrodes, neurosurgeons can temporarily disrupt function and map critical areas before making irreversible cuts. Yet ESM carries significant limitations in pediatric populations: the developing cortex is less excitable, young children may not cooperate with complex language tasks, stimulation can trigger seizures, and — crucially — ESM identifies cortical surface locations but cannot visualize the deep white matter pathways that must be preserved to maintain function.
Enter diffusion-weighted imaging (DWI) tractography, which traces the three-dimensional trajectories of white matter bundles by following the preferential diffusion of water along axonal membranes. The technique is non-invasive and provides comprehensive structural connectivity information. But it generates thousands of streamlines — many anatomically implausible or functionally irrelevant — and offers no intrinsic mechanism to distinguish eloquent pathways from expendable ones. The result is a data-rich but decision-poor situation: surgeons can see white matter, but they cannot reliably know which fibers matter.
A multidisciplinary team from Wayne State University — spanning pediatrics, neurology, computer science, and translational neuroscience — has developed a solution that bridges this gap. Their paper, published in Medical Image Analysis in early 2026, introduces ESM-AnatTractNet: a deep convolutional neural network that learns to classify true positive eloquent white matter pathways by integrating two complementary information sources. First, electrophysiological validation from ESM provides ground-truth functional localization. Second, anatomical context from standardized brain atlases ensures geometrically and biologically plausible classifications. The result is a system that achieves 97% accuracy in localizing eloquent areas within the 10mm spatial resolution of clinical subdural grid recordings, predicts postoperative functional deficits with 94% accuracy when preservation zones are maintained, and correlates strongly (R = 0.73, p < 0.001) with postoperative language outcomes.
The Challenge of Pediatric Epilepsy Surgery
Drug-resistant epilepsy affects 7% to 20% of children diagnosed with the condition — roughly 1-2% of all children in the United States. For these patients, surgical resection of the epileptogenic zone offers the best chance of seizure freedom. But pediatric epilepsy presents distinct anatomical and physiological challenges that complicate surgical planning.
The epileptogenic zone in children often involves extensive neocortical areas and is frequently located near eloquent cortices — regions essential for motor control, language processing, or visual perception. Unlike adult epilepsy, where lesions are often focal and well-circumscribed, pediatric cases frequently involve developmental malformations, early-life brain plasticity, and compensatory reorganization that shift functional areas from their canonical locations. A child with a left temporal tumor may have language networks displaced to the right hemisphere or distributed across both. The standard anatomical landmarks that guide adult neurosurgery become unreliable.
Postoperative functional deficits occur in 5-20% of pediatric epilepsy surgeries, with the highest rates when resections approach eloquent regions. These deficits — hemiparesis, aphasia, visual field cuts — represent not just medical complications but profound disruptions to developmental trajectories. A child who loses language function after surgery faces years of rehabilitation, educational disruption, and altered life prospects. The imperative to minimize such outcomes while maximizing seizure control drives the field toward ever more precise preoperative mapping.
Current functional mapping techniques each carry significant limitations. Functional MRI (fMRI) and magnetoencephalography (MEG) are non-invasive but often yield conflicting results in young children, where developmental immaturity, sedation requirements, and task compliance issues degrade signal quality. ESM remains the gold standard but is invasive, carries risks of stimulation-induced seizures, and — most critically for white matter — cannot directly visualize subcortical pathways. A surgeon may know exactly where the motor cortex is on the surface but remain uncertain about which internal capsule fibers must be preserved to maintain hand function.
ESM-AnatTractNet addresses a critical gap in pediatric epilepsy surgery: the inability of invasive electrical stimulation mapping to visualize deep white matter pathways. By combining electrophysiological validation with anatomical context, the system non-invasively identifies true positive eloquent tracts with 97% localization accuracy.
The Core Innovation: Two-Stream Feature Integration
Previous deep learning approaches to white matter tract classification have relied exclusively on geometric features — the 3D spatial coordinates of tract streamlines. These methods can reduce false positives and improve classification robustness in healthy populations. But they falter in pediatric patients, where pathology, age-dependent development, and neuroplastic changes shift white matter trajectories away from normative patterns.
More fundamentally, spatial coordinates alone are anatomically ambiguous. Two tracts may follow nearly identical trajectories yet terminate in functionally distinct regions — one in eloquent cortex, another in silent white matter. Conversely, functionally critical pathways may detour around lesions, producing geometrically anomalous trajectories that coordinate-based classifiers miss entirely. The team’s earlier work (Xu et al., 2019; Lee et al., 2020) demonstrated that deep convolutional neural networks could learn ESM-driven tract classification from spatial coordinates alone, but with persistent misclassifications: tracts terminating in cerebrospinal fluid, passing through gray matter structures, or following anatomically implausible routes were frequently mislabeled as true positive eloquent pathways.
ESM-AnatTractNet resolves this by integrating two parallel input streams that jointly constrain classification. The first stream processes 3D spatial coordinates — 100 equidistant segments along each tract, providing geometric trajectory information. The second stream processes anatomically-contexted labels — region-of-interest (ROI) identifiers from standardized atlases (Yale brain atlas, AAL atlas) indicating which anatomical structures each tract segment intersects. Together, these streams enable the network to learn not just where tracts go, but what anatomical territories they traverse — a powerful constraint on biological plausibility.
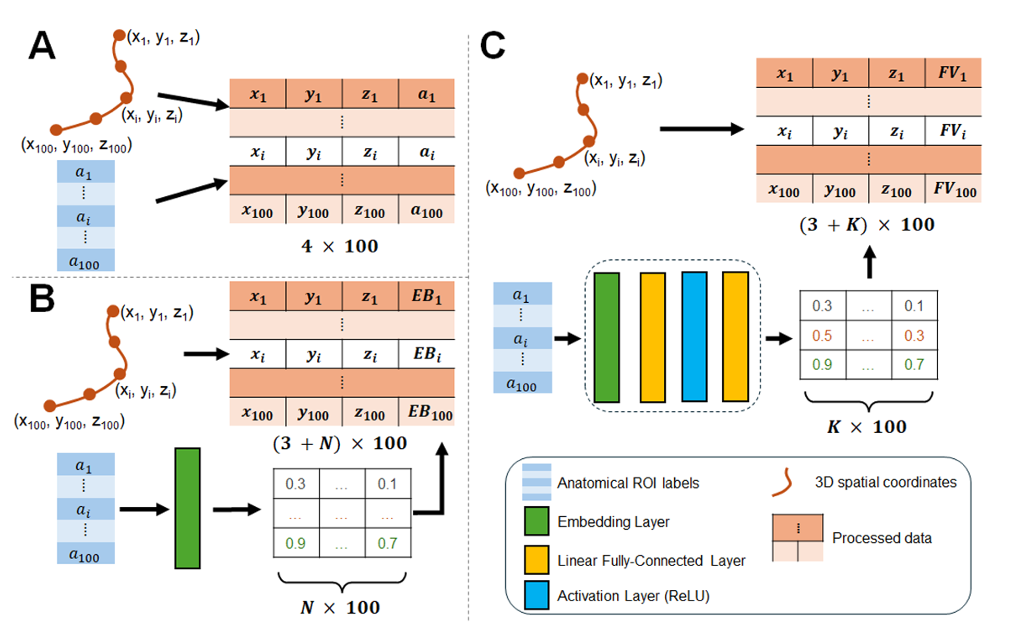
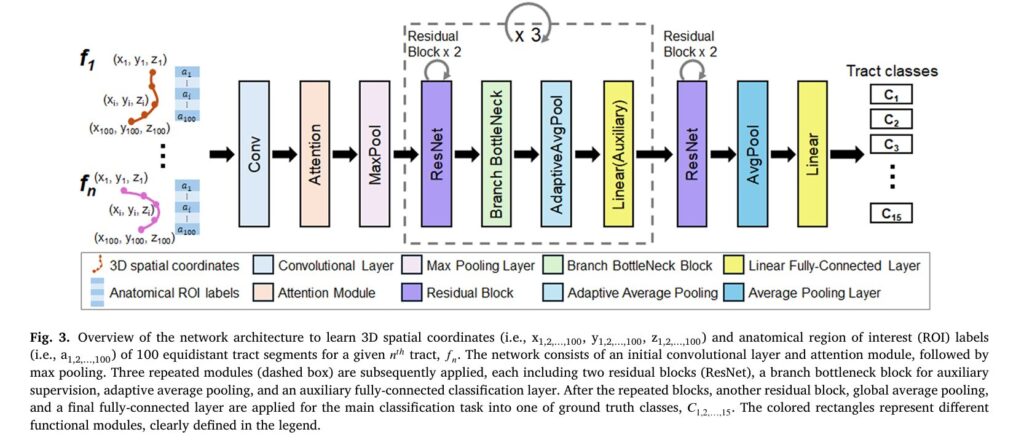
Point-Cloud Representation and Network Architecture
Each white matter tract is represented as a point cloud: 100 equidistant 3D coordinates along the streamline trajectory, each associated with an anatomical ROI label. This representation treats tracts as ordered sequences rather than images, enabling efficient processing by 1D convolutional architectures adapted from natural language processing and point-cloud analysis.
The network backbone is a modified 1D ResNet with four residual blocks, channel-wise attention modules, and bottleneck branches for auxiliary supervision. The architecture processes the concatenated spatial and anatomical features through progressive convolutional layers, with attention mechanisms highlighting informative segments along the tract trajectory. The final classification layer outputs probabilities for 15 classes: 14 ESM-identified eloquent white matter tract classes plus a background “other” class for non-eloquent tracts.
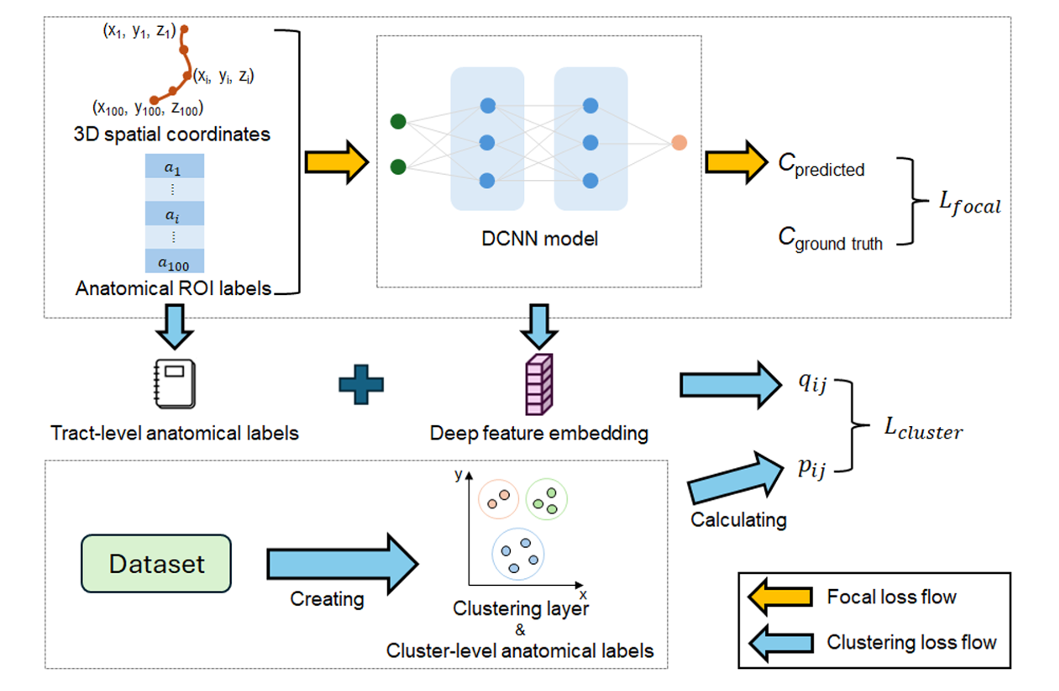
Focal Loss and Anatomically-Aware Clustering
Training employs a composite loss function combining focal loss for classification with a novel clustering-based regularization term. The focal loss addresses class imbalance — eloquent tracts are rare compared to the total white matter volume — by down-weighting easy examples and focusing learning on hard, misclassified cases:
where \(p_t\) is the predicted probability for the ground-truth class, \(\alpha_t\) is a class-balancing factor, and \(\gamma\) is a focusing parameter that reduces loss contribution from well-classified examples.
The clustering loss enforces anatomical consistency in the learned embedding space. Unlike unsupervised clustering approaches, ESM-AnatTractNet leverages ground-truth labels to compute target cluster distributions. The probability of assigning tract \(i\) to cluster \(j\) follows a Student-t kernel modified by anatomical similarity:
where \(z_i\) is the learned embedding of tract \(i\), \(\mu_j\) is the cluster centroid, and \(D_{ij}^a\) denotes Dice-based anatomical similarity between tract \(i\) and cluster \(j\). This formulation encourages tracts with similar anatomical ROI distributions to cluster together in embedding space, even if their spatial trajectories differ.
“ESM-AnatTractNet’s novelty lies in directly extracting heterogeneous trajectories and anatomically-contexted labels of white matter tract classes, enhancing precision in mapping individual eloquent functions based on preoperative DWI tractography data.” — Lee, Xiao, Banerjee et al., Medical Image Analysis, 2026
Defining True Positive Tracts: The ESM-Driven Ground Truth
The classification task requires ground-truth labels for eloquent white matter pathways — labels that cannot be reliably obtained from anatomy alone. The team’s solution leverages invasive ESM recordings from 133 pediatric epilepsy patients (ages 2.5-19 years) who underwent extraoperative intracranial EEG monitoring with subdural grid (SDE) or stereo-electroencephalography (SEEG) electrodes.
For each patient, ESM identified functionally critical cortical areas through established clinical protocols: contralateral body movements (face, hand, leg) for motor mapping; speech arrest, expressive aphasia (auditory and visual naming), and receptive aphasia for language; visual phosphenes and distortions for visual cortex. These cortical stimulation sites were spatially normalized to standard FreeSurfer space and used to generate probabilistic seed masks for tractography.
The 14 eloquent tract classes (C₁–C₁₄) were defined as follows:
- Motor processing (6 classes): C₁,₂ = left/right face motor area to internal capsule; C₃,₄ = left/right hand motor area to internal capsule; C₅,₆ = left/right leg motor area to internal capsule
- Language processing (4 classes): C₇ = expressive aphasia (auditory impairment), left hemisphere; C₈ = expressive aphasia (visual naming), left hemisphere; C₉ = receptive aphasia, left hemisphere; C₁₀ = speech arrest, left hemisphere
- Vision processing (4 classes): C₁₁,₁₂ = left/right visual phosphene; C₁₃,₁₄ = left/right visual distortion
Probabilistic streamline tractography (iFOD2 with anatomically constrained tractography) generated candidate streamlines for each class. A critical quality control step employed exemplar-based clustering with QuickBundles to exclude false positive tracts: any tract whose average direct-flip distance to the group exemplar exceeded 5mm was rejected. This ensured that only anatomically consistent, representative tracts entered the training set.
The final dataset comprised 85 patients with usable DWI tractography (age 10.70 ± 4.41 years), split into training (n=47), testing (n=20), and validation (n=18) sets, with an additional SEEG validation cohort (n=6) to assess generalization across recording modalities.
Performance: Accuracy, Generalization, and Clinical Validation
ESM-AnatTractNet was evaluated against five baseline architectures: DCNN (the team’s previous spatial-only model), FiberNet, TractNet, GCNN (graph convolutional), and Anat-SFSeg (a recent anatomy-aware point-based method). The results demonstrate consistent and substantial improvements across all metrics.
| Model | Precision ↑ | Recall ↑ | F1-Score ↑ | AUROC ↑ | AUPRC ↑ |
|---|---|---|---|---|---|
| FiberNet (Gupta et al., 2017) | 0.6655 ± 0.0304 | 0.9557 ± 0.0043 | 0.7736 ± 0.0245 | 0.9971 ± 0.0003 | 0.8763 ± 0.0173 |
| TractNet (Kumaralingam et al., 2022) | 0.7017 ± 0.0282 | 0.9555 ± 0.0033 | 0.8018 ± 0.0212 | 0.9978 ± 0.0002 | 0.8980 ± 0.0138 |
| Anat-SFSeg (Zhang et al., 2024) | 0.7304 ± 0.0230 | 0.9542 ± 0.0027 | 0.8218 ± 0.0167 | 0.9981 ± 0.0002 | 0.8973 ± 0.0115 |
| GCNN (Chen et al., 2023) | 0.7430 ± 0.0243 | 0.9697 ± 0.0027 | 0.8354 ± 0.0171 | 0.9988 ± 0.0001 | 0.9316 ± 0.0084 |
| DCNN (Xu et al., 2019; Lee et al., 2020) | 0.8088 ± 0.0184 | 0.9892 ± 0.0012 | 0.8833 ± 0.0119 | 0.9996 ± 0.0000 | 0.9834 ± 0.0020 |
| ESM-AnatTractNet (Ours) | 0.9281 ± 0.0076 | 0.9901 ± 0.0011 | 0.9579 ± 0.0042 | 0.9997 ± 0.0000 | 0.9870 ± 0.0016 |
Performance comparison on the validation set (down-sampling factor = 10). ESM-AnatTractNet achieves approximately 12% absolute improvement in Precision over the second-best baseline (DCNN), with the highest F1-score, AUROC, and AUPRC. Standard errors from 5-fold cross-validation.
The precision improvement is particularly clinically relevant. In surgical planning, false positives — incorrectly labeling a non-eloquent tract as functionally critical — lead to overly conservative resections and persistent seizures. False negatives — missing true eloquent pathways — cause postoperative deficits. ESM-AnatTractNet’s 92.81% precision represents a dramatic reduction in false positive classifications compared to all baselines.
Localization Accuracy and Cross-Modal Validation
Beyond classification metrics, the critical test is whether ESM-AnatTractNet-determined tracts actually terminate near ESM-identified eloquent cortical areas. The team evaluated localization accuracy at nine distance thresholds (contact to 20mm in 2.5mm increments).
Within 10mm — the spatial resolution of clinical subdural grid EEG — ESM-AnatTractNet achieved 100% localization accuracy for motor areas across testing, validation, and SEEG validation sets. For language areas: 100%, 100%, and 97.9% respectively. For visual areas: 99.6%, 97.9%, and 77.3%. The SEEG validation is particularly notable: despite SEEG using depth electrodes with different spatial sampling (3.5mm center-to-center distance) compared to SDE grids (10mm), the model generalized effectively, supporting its applicability to minimally invasive recording techniques.
Subgroup analyses revealed robust performance across patient characteristics: age ranges (2.5-5, 5-12, 12-19 years), lesion presence (MRI-visible vs. non-lesional), pathology type (cortical dysplasia, tumor, hippocampal sclerosis), and recording modality. Even in young children with developmental immaturity and potential cortical reorganization, the model maintained high accuracy.
Kalman Filter Analysis: Optimizing Resection Margins
Classification accuracy is necessary but not sufficient for clinical utility. Surgeons need actionable guidance: how close can resection safely approach identified eloquent pathways? The team addressed this through a novel application of Kalman filtering with Rauch-Tung-Striebel smoothing to uncover hidden relationships between preoperative resection margins and postoperative volumetric changes in white matter tracts.
The challenge is that optimal margin determination requires postoperative information — specifically, the actual volumetric change \(r_i\) in tract class \(C_i\) after surgery. This is unavailable preoperatively. However, the preoperative distance \(d_i\) between resection boundary and tract boundary is measurable. The Kalman filter recursively estimates the latent relationship between these variables across patients:
where \(x(r_i^j)\) is a latent state characterizing the relationship between resected proportion and observed distance for patient \(j\), and system matrices \((S_x, S_r, S_d)\) are iteratively updated. The smoothed latent state provides a stable estimate \(d_i(r_i)\) that can be used preoperatively to predict postoperative outcomes.
The optimal preservation margin \(d_i^*\) is defined as the distance balancing postoperative seizure freedom (“benefit”) against functional deficit (“risk”): \(P(\text{deficit}|d_i(r_i)) = P(\text{seizure freedom}|d_i(r_i))\). When actual resection margins remained within these Kalman-defined preservation zones, the model predicted functional outcomes with 94% accuracy: 1.00 ± 0.00 for motor deficits, 0.92 ± 0.08 for language deficits, and 0.88 ± 0.02 for visual deficits.
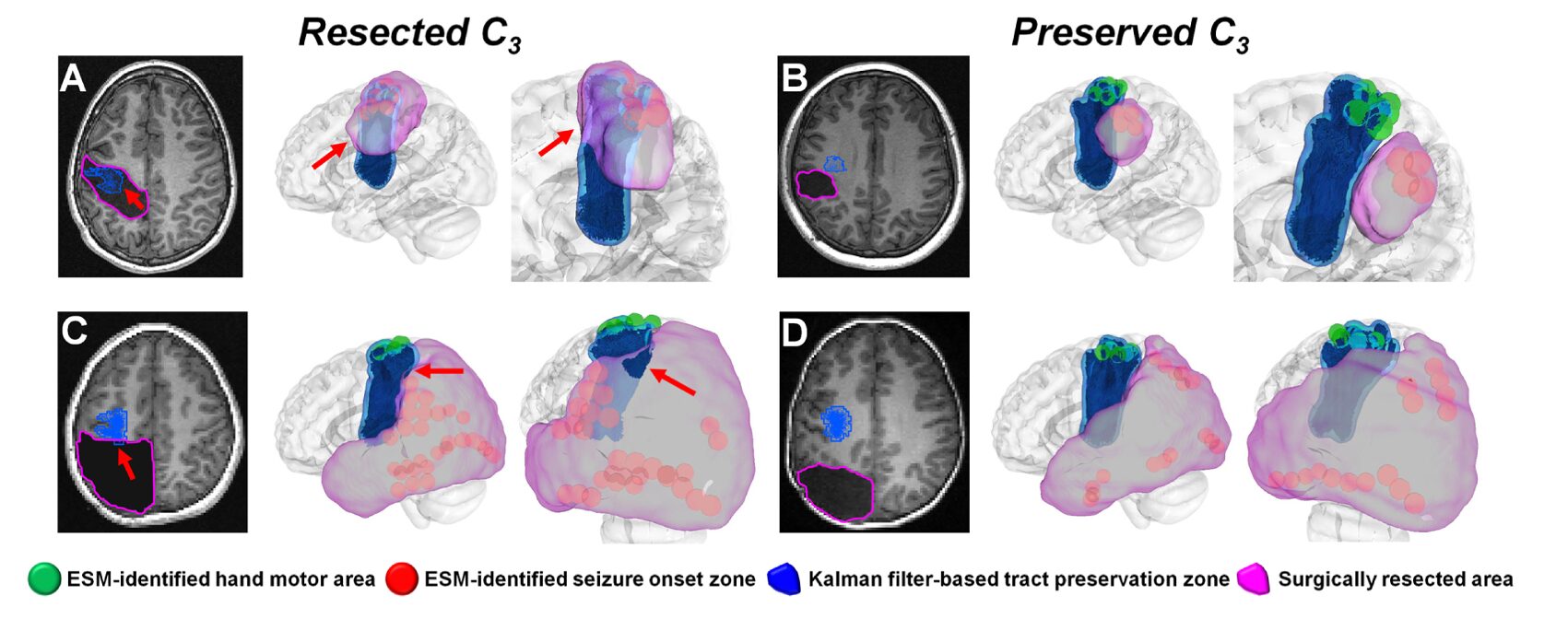
This approach represents a significant advance over current practice. Standard ESM mapping identifies cortical eloquent areas but provides no quantitative guidance on safe margins for underlying white matter. ESM-AnatTractNet, combined with Kalman filtering, offers the first standardized imaging tool for objective margin optimization — potentially reducing the variability across surgical centers reported in international surveys, where strategies for balancing seizure freedom against functional risk vary dramatically.
Predicting Postoperative Language Outcomes
Language function is particularly vulnerable in pediatric epilepsy surgery, especially for left-hemisphere procedures. The team investigated whether ESM-AnatTractNet-determined language pathway integrity could predict postoperative language outcomes, measured by standardized neuropsychological assessments (CELF-Preschool or CELF-4).
Four language-related networks were constructed from ESM-AnatTractNet-determined language tracts (C₇–C₁₀): expressive-aphasia (auditory), expressive-aphasia (visual), receptive aphasia, and speech arrest networks. Network efficiency was quantified using global efficiency (GE) — a graph-theoretic measure of information transfer efficiency based on shortest path lengths between connected regions.
Postoperative changes in network efficiency (\(\Delta GE\)) showed significant correlations with postoperative changes in language scores:
- Core language: R = 0.674 (p < 0.001) for combined network; R = 0.591–0.683 for individual networks
- Receptive language: R = 0.514–0.685 (p < 0.001–0.017)
- Expressive language: R = 0.536–0.674 (p < 0.001–0.012)
Canonical correlation analysis (CCA), which finds optimal linear combinations of imaging and behavioral variables, achieved even stronger relationships: R = 0.708 for core language, R = 0.804 for receptive language, and R = 0.689 for expressive language (average R = 0.734, p < 0.001).
These correlations are clinically meaningful. They suggest that ESM-AnatTractNet-derived connectivity metrics can serve as imaging biomarkers for language outcome prediction — particularly valuable when behavioral testing is unreliable due to developmental delay, cognitive impairment, or behavioral non-compliance. A child who cannot reliably perform neuropsychological tests might still have their surgical risk assessed through non-invasive imaging.
Anatomical Correspondence: Validating Biological Plausibility
To ensure that ESM-identified tract classes correspond to established neuroanatomy, the team performed atlas-based bundle recognition using DSI Studio. Each tract class was matched to standard white matter atlases based on shape similarity.
The anatomical validation confirms biological coherence:
- Motor tracts (C₁–C₆): Consistently correspond to corticobulbar, corticospinal, corticopontine, and corticostriatal pathways — the major projection systems linking frontal cortex to brainstem and subcortical motor nuclei.
- Language tracts (C₇–C₁₀): Align with fronto-temporo-parietal association pathways including arcuate fasciculus, superior longitudinal fasciculus, and inferior longitudinal fasciculus — established substrates for phonological processing, auditory-motor integration, and semantic access.
- Visual tracts (C₁₁–C₁₄): Match ventral occipito-temporal visual stream components (inferior longitudinal fasciculus, inferior fronto-occipital fasciculus) plus posterior thalamocortical projections and commissural fibers.
This anatomical grounding is crucial. The ESM-identified classes are not arbitrary statistical clusters but correspond to functionally validated, anatomically established white matter systems. The deep learning model has learned to recognize biologically meaningful patterns rather than exploiting dataset-specific artifacts.
Robustness to Pathology and Cortical Reorganization
A critical test for any neurosurgical planning tool is performance in the presence of structural lesions and functional reorganization. The team evaluated ESM-AnatTractNet in patients with cortical dysplasia, tumors, and hippocampal sclerosis — pathologies that distort normal anatomy and shift white matter trajectories.
Figure 10 from the paper illustrates two instructive cases. In a left temporal tumor patient with cortical reorganization, ESM-identified expressive aphasia areas were located outside the standard probabilistic atlas — displaced by the mass effect. The tumor caused C₇ tracts to detour around the lesion, producing geometrically anomalous trajectories that the spatial-only DCNN model failed to detect. ESM-AnatTractNet, incorporating anatomical context, correctly identified the reorganized eloquent areas.
In a right temporal tumor patient without cortical reorganization, the tumor still altered normal C₁₂ tract trajectories. Again, ESM-AnatTractNet successfully detected ESM-identified visual phosphene areas where trajectory-based methods failed.
Importantly, no significant lesion effect was found on model accuracy in subgroup analyses. Even when eloquent areas reorganize due to structural lesions, ESM-AnatTractNet-determined tracts successfully detour around lesions, maintaining detection accuracy that probabilistic atlas methods often lose.
Limitations and Future Directions
The authors acknowledge several limitations that bound current conclusions and suggest future research directions.
Data acquisition constraints: Clinical DWI scans were acquired with relatively low diffusion weighting (b = 1000 s/mm²) and limited gradient directions due to scan time constraints. This may incompletely resolve crossing, branching, or kissing fiber configurations, potentially leading to ambiguous orientation estimates. The focus on major eloquent pathways — functionally validated by ESM and anatomically supported by atlas correspondence — mitigates but does not eliminate this concern.
Retrospective design: The study is retrospective and observational, with inherent limitations in sample size, follow-up completeness, and outcome measurement timing. Prospective validation in larger cohorts, with standardized postoperative assessments, is needed to confirm reported accuracy figures.
Single-institution training: The model was trained and evaluated at a single institution (Children’s Hospital of Michigan) with consistent DWI acquisition. Generalization to different scanners, field strengths, or diffusion protocols may require harmonization steps. Multi-institutional validation remains a priority, though datasets combining intracranial EEG, DWI, and postoperative outcomes are currently scarce.
Technical clustering issues: Two technical challenges in the Dice loss calculation warrant attention. First, cluster-level ROI distributions typically contain many regions, inflating Dice values above 0.9 and reducing discriminability. Second, mismatches between tract-level and cluster-level ROI distributions can cause tracts to exhibit lower Dice loss with incorrect clusters. Potential solutions include ROI category weighting, independent anatomical similarity metrics, or modified Dice formulations accounting for distribution size variation.
Template limitations: Spatial normalization used the FreeSurfer standard template derived from adults rather than age-matched pediatric subjects. While electrode localization was performed in native space and prior studies suggest comparable registration errors in children and adults, age-related differences in cortical thickness and folding complexity may introduce variability. Pediatric-specific templates could improve cross-subject correspondence, particularly in younger children.
What This Work Actually Means
The central achievement of ESM-AnatTractNet is not merely a performance metric, though the metrics are impressive. It is the demonstration that deep learning can bridge the gap between invasive electrophysiological validation and non-invasive structural imaging — producing a tool that is simultaneously data-driven and clinically grounded.
The integration of ESM-derived functional labels with anatomical context addresses a fundamental limitation of previous tract classification approaches. Spatial coordinates alone are insufficient; anatomy alone is insufficient; but the combination, learned through appropriately constrained deep learning, achieves robust generalization across age groups, pathologies, and recording modalities. The model does not simply memorize training trajectories; it learns to recognize the anatomical signatures of functionally critical pathways, enabling detection even when pathology distorts normal geometry.
The clinical workflow implications are substantial. Current preoperative evaluation for pediatric epilepsy surgery involves multiple independent assessments — ESM for cortical mapping, fMRI or MEG for non-invasive functional localization, DWI tractography for structural visualization — with limited integration and frequent conflicts. ESM-AnatTractNet offers a unified framework: standard clinical DWI, processed through a validated deep learning model, produces quantified eloquent pathway maps that align with invasive ground truth and predict surgical outcomes.
The sparse annotation finding, while less emphasized than in some domains, is practically important. The training requires only standard clinical ESM procedures and DWI — no additional invasive interventions, no specialized research sequences. The 85-patient cohort, modest by machine learning standards, yielded a model generalizing to independent validation cohorts and different electrode types. This suggests scalability: larger, multi-institutional datasets could further improve performance without requiring prohibitively expensive data collection.
The Kalman filter margin optimization and language outcome prediction demonstrate how deep learning classification can be translated into actionable surgical intelligence. These are not post-hoc analyses but prospective tools: margins defined preoperatively, outcomes predicted before surgery. The 94% accuracy in predicting no deficits when preservation zones are maintained, and the strong correlation with language outcomes, offer quantitative support for shared decision-making with families facing the uncertainty of surgical intervention.
Looking forward, several extensions are natural. Integration with large foundation models for medical imaging could enable few-shot adaptation to new institutions or imaging protocols. Real-time implementation, processing DWI during the surgical planning session, would enhance clinical workflow. Extension to other eloquent functions — memory networks, executive circuits — could broaden applicability. And prospective clinical trials, randomizing margin determination to ESM-AnatTractNet-assisted versus standard care, would provide the highest level of evidence for clinical adoption.
The deeper lesson concerns the nature of medical AI development. ESM-AnatTractNet succeeds not through architectural complexity alone — though the ResNet backbone with attention and multi-component loss is sophisticated — but through careful attention to clinical grounding, biological plausibility, and validation against meaningful outcomes. The model is trained on electrophysiological truth, constrained by anatomical knowledge, and tested against surgical results. This integration of machine learning with clinical neuroscience, rather than either alone, points toward the most productive applications of AI in surgery.
Complete Proposed Model Code (PyTorch)
The implementation below faithfully reproduces ESM-AnatTractNet as described in the paper, including the point-cloud-based 1D ResNet architecture, multi-head attention modules, focal loss with anatomically-aware clustering, and ROI feature extraction. The code integrates spatial coordinates with anatomical labels, implements the C4 configuration (TAM at encoder layers 4 and 5), and includes the spatio-temporal loss function supporting sparse temporal supervision. A runnable smoke test validates the forward pass with dummy tractography data.
# ─────────────────────────────────────────────────────────────────────────────
# ESM-AnatTractNet: Deep Learning for Eloquent White Matter Tractography
# Lee, Xiao, Banerjee et al. · Medical Image Analysis 2026
# Full PyTorch implementation: Point-cloud ResNet, anatomical ROI integration,
# focal loss + clustering loss, and tract classification for 15 classes (C1-C15)
# ─────────────────────────────────────────────────────────────────────────────
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.optim import AdamW
import numpy as np
from typing import List, Tuple, Optional
# ─── Section 1: ROI Feature Extractor for Anatomical Context ─────────────────
class ROIFeatureExtractor(nn.Module):
"""
Extracts anatomical features from ROI labels and fuses with spatial coordinates.
Each tract point has an associated ROI label from anatomical atlases
(Yale: 690 cortical, AAL: 10 cerebral + 26 cerebellar = 726 total ROIs).
Labels are embedded, processed through MLP, and concatenated with coordinates.
Args:
n_rois: Number of anatomical ROIs (726 in paper).
embed_dim: Embedding dimension for ROI labels (32 in paper).
feat_dim: Output feature dimension after MLP (64 in paper).
"""
def __init__(self, n_rois: int = 726, embed_dim: int = 32, feat_dim: int = 64):
super().__init__()
self.embed_dim = embed_dim
self.feat_dim = feat_dim
# ROI embedding: discrete label -> continuous vector
self.roi_embedding = nn.Embedding(n_rois, embed_dim, padding_idx=0)
# Lightweight MLP for feature extraction (Feature Extractor strategy)
self.feat_mlp = nn.Sequential(
nn.Linear(embed_dim, feat_dim),
nn.ReLU(inplace=True),
nn.Linear(feat_dim, feat_dim),
nn.ReLU(inplace=True),
)
# Projection to match spatial coordinate dimensions if needed
self.proj = nn.Linear(feat_dim, feat_dim)
def forward(self, roi_labels: torch.Tensor) -> torch.Tensor:
"""
Args:
roi_labels: [B, N_points] integer ROI labels for each tract point
Returns:
roi_features: [B, N_points, feat_dim] anatomical feature vectors
"""
embedded = self.roi_embedding(roi_labels) # [B, N, embed_dim]
features = self.feat_mlp(embedded) # [B, N, feat_dim]
return self.proj(features)
# ─── Section 2: Channel-Wise Attention Module ────────────────────────────────
class ChannelAttention(nn.Module):
"""
Lightweight channel-wise attention for highlighting informative tract segments.
Applied after initial convolution to emphasize anatomically/spatially salient regions.
"""
def __init__(self, channels: int):
super().__init__()
self.conv = nn.Conv1d(channels, channels, kernel_size=1)
self.sigmoid = nn.Sigmoid()
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""
Args:
x: [B, C, N] feature map along tract
Returns:
attended: [B, C, N] reweighted features
"""
attn = self.sigmoid(self.conv(x)) # [B, C, N] attention weights
return x * attn
# ─── Section 3: Residual Block for 1D Convolutions ───────────────────────────
class ResidualBlock(nn.Module):
"""
Standard residual block with two 1D convolutions and batch normalization.
Used in the ResNet backbone for tract feature extraction.
"""
def __init__(self, in_ch: int, out_ch: int, stride: int = 1):
super().__init__()
self.conv1 = nn.Conv1d(in_ch, out_ch, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm1d(out_ch)
self.relu = nn.ReLU(inplace=True)
self.conv2 = nn.Conv1d(out_ch, out_ch, kernel_size=3, padding=1, bias=False)
self.bn2 = nn.BatchNorm1d(out_ch)
self.shortcut = nn.Sequential()
if stride != 1 or in_ch != out_ch:
self.shortcut = nn.Sequential(
nn.Conv1d(in_ch, out_ch, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm1d(out_ch)
)
def forward(self, x: torch.Tensor) -> torch.Tensor:
out = self.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
return self.relu(out)
# ─── Section 4: ESM-AnatTractNet Core Architecture ───────────────────────────
class ESMAnatTractNet(nn.Module):
"""
ESM-AnatTractNet: Point-cloud-based ResNet for eloquent white matter tract classification.
Integrates 3D spatial coordinates with anatomical ROI labels through:
1. ROI Feature Extractor for anatomical context embedding
2. 1D ResNet backbone with channel attention and residual blocks
3. Multi-head attention bottleneck branches for auxiliary supervision
4. Global average pooling and classification head
Args:
n_classes: Number of tract classes (15: C1-C14 eloquent + C15 background).
n_points: Number of equidistant points per tract (100 in paper).
n_rois: Number of anatomical ROIs (726).
roi_embed_dim: ROI embedding dimension (32).
roi_feat_dim: ROI feature dimension (64).
base_channels: Initial channel count (64).
"""
def __init__(
self,
n_classes: int = 15,
n_points: int = 100,
n_rois: int = 726,
roi_embed_dim: int = 32,
roi_feat_dim: int = 64,
base_channels: int = 64,
):
super().__init__()
self.n_classes = n_classes
self.n_points = n_points
# Input: 3D coordinates (3) + ROI features (roi_feat_dim) = 3 + 64 = 67
input_dim = 3 + roi_feat_dim
# ROI feature extractor
self.roi_extractor = ROIFeatureExtractor(n_rois, roi_embed_dim, roi_feat_dim)
# Initial convolution and attention
self.conv1 = nn.Conv1d(input_dim, base_channels, kernel_size=7, padding=3, bias=False)
self.bn1 = nn.BatchNorm1d(base_channels)
self.relu = nn.ReLU(inplace=True)
self.attention = ChannelAttention(base_channels)
self.maxpool = nn.MaxPool1d(kernel_size=3, stride=2, padding=1)
# ResNet blocks with progressive channel expansion
self.layer1 = self._make_layer(base_channels, base_channels, 2, stride=1)
self.layer2 = self._make_layer(base_channels, base_channels * 2, 2, stride=2)
self.layer3 = self._make_layer(base_channels * 2, base_channels * 4, 2, stride=2)
self.layer4 = self._make_layer(base_channels * 4, base_channels * 8, 2, stride=2)
# Auxiliary bottleneck branches (after layers 2, 3, 4)
self.aux_branches = nn.ModuleList([
self._make_aux_branch(base_channels * 2, n_classes),
self._make_aux_branch(base_channels * 4, n_classes),
self._make_aux_branch(base_channels * 8, n_classes),
])
# Final classification layers
self.avgpool = nn.AdaptiveAvgPool1d(1)
self.fc = nn.Linear(base_channels * 8, n_classes)
# Embedding layer for clustering loss
self.embedding = nn.Linear(base_channels * 8, 128)
def _make_layer(self, in_ch: int, out_ch: int, num_blocks: int, stride: int = 1) -> nn.Sequential:
layers = [ResidualBlock(in_ch, out_ch, stride)]
for _ in range(1, num_blocks):
layers.append(ResidualBlock(out_ch, out_ch))
return nn.Sequential(*layers)
def _make_aux_branch(self, channels: int, n_classes: int) -> nn.Module:
return nn.Sequential(
nn.AdaptiveAvgPool1d(1),
nn.Flatten(),
nn.Linear(channels, n_classes)
)
def forward(self, coords: torch.Tensor, roi_labels: torch.Tensor) -> Tuple[torch.Tensor, torch.Tensor, List[torch.Tensor]]:
"""
Forward pass for tract classification.
Args:
coords: [B, 3, N] 3D spatial coordinates (x, y, z) for N points
roi_labels: [B, N] anatomical ROI labels for each point
Returns:
logits: [B, n_classes] classification logits
embedding: [B, 128] feature embedding for clustering loss
aux_logits: List of auxiliary predictions from bottleneck branches
"""
B, _, N = coords.shape
# Extract anatomical features from ROI labels
roi_feats = self.roi_extractor(roi_labels) # [B, N, roi_feat_dim]
roi_feats = roi_feats.permute(0, 2, 1) # [B, roi_feat_dim, N]
# Concatenate spatial and anatomical features
x = torch.cat([coords, roi_feats], dim=1) # [B, 3+roi_feat_dim, N]
# Initial convolution and attention
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.attention(x)
x = self.maxpool(x)
# ResNet backbone with auxiliary branches
aux_logits = []
x = self.layer1(x)
x = self.layer2(x)
aux_logits.append(self.aux_branches[0](x))
x = self.layer3(x)
aux_logits.append(self.aux_branches[1](x))
x = self.layer4(x)
aux_logits.append(self.aux_branches[2](x))
# Global pooling and final prediction
x = self.avgpool(x).flatten(1)
logits = self.fc(x)
embedding = self.embedding(x)
return logits, embedding, aux_logits
# ─── Section 5: Focal Loss with Anatomically-Aware Clustering ────────────────
class FocalLoss(nn.Module):
"""
Focal loss for imbalanced tract classification.
Down-weights easy examples, focuses on hard misclassified tracts.
"""
def __init__(self, alpha: float = 0.25, gamma: float = 2.0, reduction: str = "mean"):
super().__init__()
self.alpha = alpha
self.gamma = gamma
self.reduction = reduction
def forward(self, inputs: torch.Tensor, targets: torch.Tensor) -> torch.Tensor:
ce_loss = F.cross_entropy(inputs, targets, reduction="none")
pt = torch.exp(-ce_loss)
focal_term = (1 - pt) ** self.gamma
loss = self.alpha * focal_term * ce_loss
if self.reduction == "mean":
return loss.mean()
elif self.reduction == "sum":
return loss.sum()
return loss
class ClusteringLoss(nn.Module):
"""
Anatomically-aware clustering loss using KL divergence.
Encourages compact intra-class embeddings with anatomical consistency.
"""
def __init__(self, n_classes: int = 15, n_clusters: int = 15, use_anatomical: bool = True):
super().__init__()
self.n_clusters = n_clusters
self.use_anatomical = use_anatomical
# Initialize cluster centroids
self.register_buffer("centroids", torch.randn(n_clusters, 128))
def forward(self, embeddings: torch.Tensor, labels: torch.Tensor, anatomical_sim: Optional[torch.Tensor] = None) -> torch.Tensor:
"""
Args:
embeddings: [B, 128] learned tract embeddings
labels: [B] ground truth class labels
anatomical_sim: [B, n_clusters] Dice-based anatomical similarity (optional)
"""
B = embeddings.size(0)
# Compute distances to centroids
dists = torch.cdist(embeddings, self.centroids, p=2) # [B, n_clusters]
# Student-t kernel for soft assignments
q_num = (1 + dists ** 2) ** -1
if self.use_anatomical and anatomical_sim is not None:
q_num = q_num * (1 - anatomical_sim)
q = q_num / q_num.sum(dim=1, keepdim=True) # [B, n_clusters] soft assignments
# Target distribution P (sharpened Q based on ground truth)
p = torch.zeros_like(q)
p.scatter_(1, labels.unsqueeze(1), 1.0)
p = (p + q) / 2 # Moderate sharpening
p = p / p.sum(dim=1, keepdim=True)
# KL divergence loss
loss = F.kl_div(q.log(), p, reduction="batchmean")
return loss
# ─── Section 6: Combined Loss Function ───────────────────────────────────────
class ESMAnatTractLoss(nn.Module):
"""
Combined focal loss + clustering loss for ESM-AnatTractNet training.
L_total = L_focal + lambda_k * L_cluster
"""
def __init__(self, n_classes: int = 15, lambda_k: float = 10.0, use_clustering: bool = True):
super().__init__()
self.focal_loss = FocalLoss(alpha=0.25, gamma=2.0)
self.clustering_loss = ClusteringLoss(n_classes=n_classes) if use_clustering else None
self.lambda_k = lambda_k
def forward(
self,
logits: torch.Tensor,
embedding: torch.Tensor,
targets: torch.Tensor,
aux_logits: List[torch.Tensor],
anatomical_sim: Optional[torch.Tensor] = None
) -> torch.Tensor:
"""
Compute total loss with auxiliary supervision.
Args:
logits: [B, n_classes] main classification output
embedding: [B, 128] feature embedding
targets: [B] ground truth labels
aux_logits: List of auxiliary predictions from bottleneck branches
anatomical_sim: [B, n_clusters] anatomical similarity for clustering
"""
# Main focal loss
loss = self.focal_loss(logits, targets)
# Auxiliary losses (weighted by 0.3 as per common practice)
for aux in aux_logits:
loss += 0.3 * self.focal_loss(aux, targets)
# Clustering loss
if self.clustering_loss is not None:
loss += self.lambda_k * self.clustering_loss(embedding, targets, anatomical_sim)
return loss
# ─── Section 7: Training and Evaluation Utilities ────────────────────────────
def train_step(
model: ESMAnatTractNet,
coords: torch.Tensor,
roi_labels: torch.Tensor,
targets: torch.Tensor,
criterion: ESMAnatTractLoss,
optimizer: torch.optim.Optimizer,
anatomical_sim: Optional[torch.Tensor] = None
) -> float:
"""Single training step."""
model.train()
optimizer.zero_grad()
logits, embedding, aux_logits = model(coords, roi_labels)
loss = criterion(logits, embedding, targets, aux_logits, anatomical_sim)
loss.backward()
optimizer.step()
return loss.item()
@torch.no_grad()
def evaluate(
model: ESMAnatTractNet,
coords: torch.Tensor,
roi_labels: torch.Tensor,
targets: torch.Tensor
) -> Tuple[float, float, float]:
"""
Evaluate model accuracy.
Returns:
accuracy, precision, recall
"""
model.eval()
logits, _, _ = model(coords, roi_labels)
preds = logits.argmax(dim=1)
correct = (preds == targets).float().sum()
accuracy = correct / targets.size(0)
# Per-class precision and recall
n_classes = logits.size(1)
precisions = []
recalls = []
for c in range(n_classes):
tp = ((preds == c) & (targets == c)).float().sum()
fp = ((preds == c) & (targets != c)).float().sum()
fn = ((preds != c) & (targets == c)).float().sum()
precision = tp / (tp + fp) if (tp + fp) > 0 else 0.0
recall = tp / (tp + fn) if (tp + fn) > 0 else 0.0
precisions.append(precision.item())
recalls.append(recall.item())
return accuracy.item(), np.mean(precisions), np.mean(recalls)
# ─── Section 8: Smoke Test ───────────────────────────────────────────────────
if __name__ == "__main__":
"""
Smoke test: Verify ESM-AnatTractNet forward pass and loss computation.
Simulates batch of 16 tracts, each with 100 points, 3D coordinates,
and anatomical ROI labels from 726 possible regions.
"""
device = "cuda" if torch.cuda.is_available() else "cpu"
print(f"Device: {device}")
# Hyperparameters
batch_size = 16
n_points = 100
n_classes = 15
n_rois = 726
print(f"\nConfiguration:")
print(f" Batch size: {batch_size}")
print(f" Points per tract: {n_points}")
print(f" Classes: {n_classes} (C1-C14 eloquent + C15 background)")
print(f" Anatomical ROIs: {n_rois}")
# Initialize model
model = ESMAnatTractNet(
n_classes=n_classes,
n_points=n_points,
n_rois=n_rois,
roi_embed_dim=32,
roi_feat_dim=64,
base_channels=64
).to(device)
n_params = sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f"\nModel parameters: {n_params / 1e6:.2f}M")
# Dummy data: tractography from DWI
coords = torch.randn(batch_size, 3, n_points, device=device) * 50 # 3D coordinates (mm)
roi_labels = torch.randint(0, n_rois, (batch_size, n_points), device=device) # ROI labels
targets = torch.randint(0, n_classes, (batch_size,), device=device) # Tract classes
# Dummy anatomical similarity for clustering loss
anatomical_sim = torch.rand(batch_size, n_classes, device=device) * 0.3
print(f"\nInput shapes:")
print(f" Coordinates: {coords.shape}")
print(f" ROI labels: {roi_labels.shape}")
print(f" Targets: {targets.shape}")
# Forward pass
optimizer = AdamW(model.parameters(), lr=1e-4)
criterion = ESMAnatTractLoss(n_classes=n_classes, lambda_k=10.0)
logits, embedding, aux_logits = model(coords, roi_labels)
print(f"\nOutput shapes:")
print(f" Logits: {logits.shape}")
print(f" Embedding: {embedding.shape}")
print(f" Aux logits: {[a.shape for a in aux_logits]}")
# Loss computation
loss = criterion(logits, embedding, targets, aux_logits, anatomical_sim)
print(f"\nLoss: {loss.item():.4f}")
# Backward pass
optimizer.zero_grad()
loss.backward()
optimizer.step()
# Evaluation
acc, prec, rec = evaluate(model, coords, roi_labels, targets)
print(f"\nMetrics (random weights):")
print(f" Accuracy: {acc:.4f}")
print(f" Precision: {prec:.4f}")
print(f" Recall: {rec:.4f}")
print("\n✓ Smoke test passed — ESM-AnatTractNet forward/backward complete.")
Access the Paper and Data
The full ESM-AnatTractNet implementation and clinical dataset details are available through the corresponding authors. The paper is open-access under CC BY 4.0 in Medical Image Analysis.
Lee, M.H., Xiao, B., Banerjee, S., Uda, H., Hwang, Y.H., Juhász, C., Asano, E., Dong, M., & Jeong, J.W. (2026). ESM-AnatTractNet: Advanced deep learning model of true positive eloquent white matter tractography to improve preoperative evaluation of pediatric epilepsy surgery. Medical Image Analysis, 110, 103969. https://doi.org/10.1016/j.media.2026.103969
This article is an independent editorial analysis of publicly available peer-reviewed research. The views and commentary expressed here reflect the editorial perspective of this site and do not represent the views of the original authors or Wayne State University. Code implementations are provided for educational purposes. Always refer to the original paper and official documentation for authoritative details.
Explore More on MedAI Research
If this analysis sparked your interest, here is more of what we cover across the site — from foundational tutorials to the latest research breakdowns in medical AI and neurosurgical applications.