IERE: Teaching a Small Medical Segmentation Model to Generalize Using SAM — Only During Training
Researchers at Ruijin Hospital and the Chinese Academy of Sciences found a smarter way to use the Segment Anything Model in medical imaging: not as the deployed model, but as a teacher that never shows up at inference time — reducing domain-shift failures by over 2.66% Dice while keeping the deployed model as fast and lightweight as a standard UNet.
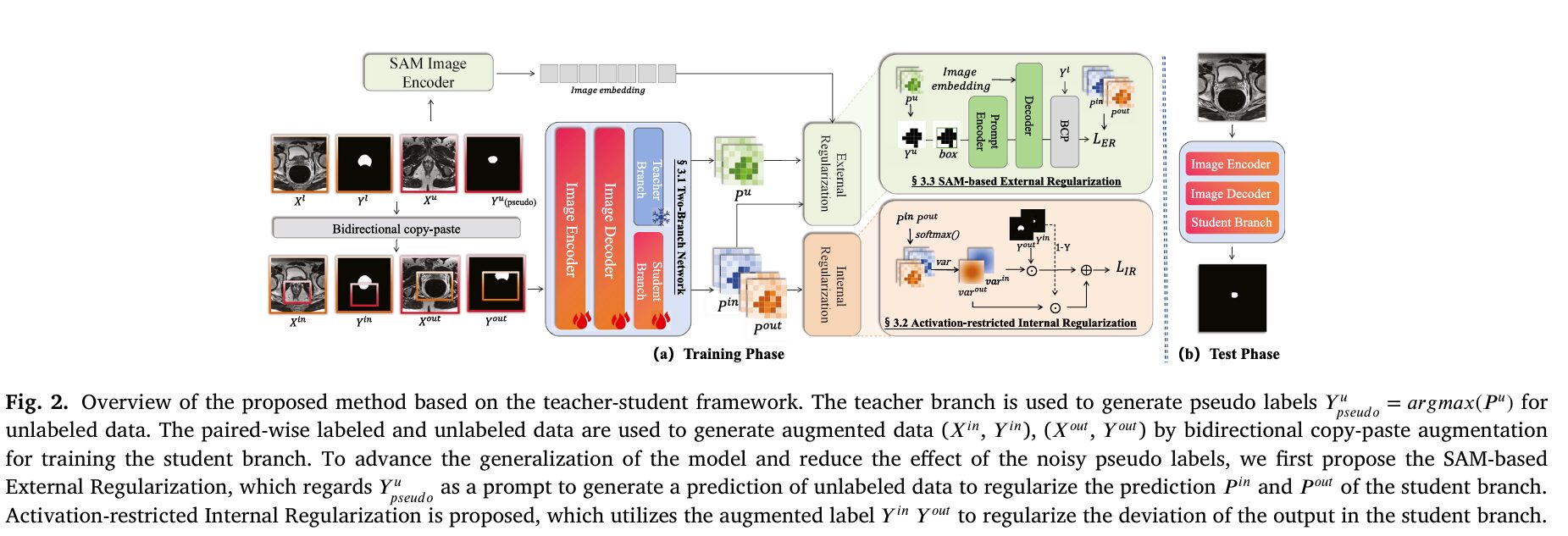
When a prostate segmentation model trained at one hospital fails on MRI scans from another hospital, the failure is rarely because the anatomy changed. The prostate is in the same place. It looks roughly the same. What changed is the scanner brand, the magnetic field strength, the imaging protocol, and a hundred small acquisition details that shift the pixel intensity distribution just enough to confuse a model that has only ever seen one hospital’s data. This domain shift problem is one of the most stubborn barriers between AI segmentation research and clinical deployment. IERE tackles it from an angle that most methods miss: not by improving the model itself, but by improving what the model is trained against.
The Two Problems That IERE Solves at Once
There are two distinct failure modes in cross-domain semi-supervised medical segmentation. Understanding both is necessary to appreciate why IERE’s dual-regularization design is structured the way it is.
The first is pseudo-label noise. In a teacher-student training framework, the teacher network generates predictions on unlabeled images — these are the pseudo labels that supervise the student. But the teacher is not perfect, especially early in training. Its pseudo labels contain errors, and those errors propagate into the student’s training signal. Naively trusting pseudo labels causes the student to learn confidently wrong patterns, particularly at organ boundaries and background regions where the teacher hesitates.
The second failure mode is domain overfitting. Even with good pseudo labels and a large unlabeled dataset, a model trained on a single source domain will fit that domain’s specific visual statistics — the intensity distribution, noise characteristics, and scanner artefacts of a 1.5T Philips machine with an endorectal coil, for instance. When test time arrives with a 3T Siemens surface-coil scan, the features the model learned are partly the right ones and partly noise that looked like signal in training. The model does not know which is which because it has never seen anything else.
Most existing approaches address one of these problems. Methods like BCP and MCF focus on better exploitation of unlabeled data through data augmentation and self-correction, which helps pseudo-label quality. Methods like MaxStyle and TriD focus on domain generalization through data augmentation strategies that simulate distribution shift during training. Neither approach fully handles both problems simultaneously, and critically, both families of methods rely on the assumption that labeled data is plentiful enough to represent the true domain distribution — an assumption that fails in low-data clinical regimes.
IERE combines two complementary regularizations: an internal one that suppresses wrong activations caused by noisy pseudo labels (fixing the pseudo-label noise problem), and an external one that uses SAM’s strong generalization capability to prevent the student from overfitting the source domain’s visual statistics (fixing the domain overfitting problem). Both are applied only during training, so the deployed model remains a simple UNet with no added inference cost.
The Framework: Teacher, Student, and a Borrowed Generalizer
IERE follows the standard teacher-student semi-supervised setup. Both branches share a common encoder-decoder backbone (ResNet + UNet decoder) but have separate segmentation heads. The teacher is trained first on labeled data alone using an IOU loss, establishing a reliable baseline before generating pseudo labels. This pre-training step is important — a teacher that has never seen supervision produces unreliable pseudo labels, and the whole cascade depends on those labels being at least roughly correct.
The teacher then generates pseudo labels for unlabeled images, and bidirectional copy-paste augmentation (BCP) is used to create mixed image-label pairs. In this augmentation, patches from unlabeled data are pasted into labeled images and vice versa, creating two augmented pairs (X_in, Y_in) and (X_out, Y_out) that expose the student to mixed distributions without requiring additional labeled data.
The student branch is trained with three loss terms: the base segmentation loss L_s on both labeled and augmented unlabeled data, the internal regularization loss L_IR, and the external regularization loss L_ER:
At inference time, only the student branch is used. The teacher disappears. SAM disappears. The deployed model is exactly as fast as a standalone UNet — 0.29 seconds per inference, 32.56G FLOPs, 32.51M parameters.
TRAINING PHASE
══════════════════════════════════════════════════════
Labeled data (X_l, Y_l) ──────────────────────────────┐
│
┌──────────────────────────────────────────────────┐ │
│ TEACHER BRANCH f(·; θ_T) │◄───┘ Pre-train: L_t = L_Jaccard
│ (ResNet encoder + UNet decoder + seg head) │ on labeled data only
└─────────────────────┬────────────────────────────┘
│ generates pseudo labels Y_u = argmax(P_u)
▼
Unlabeled data X_u ──► Y_u (teacher pseudo labels)
│
├──► Bounding box [x,y,w,h] extracted
│ │
│ ┌────▼──────────────────────────┐
│ │ SAM f(·; θ_sam) │◄── Fine-tuned on (X_l, Y_l)
│ │ (MedSAM, frozen at inference) │ via L_sam = L_Jaccard(P_sam, Y_l)
│ └────────────────┬──────────────┘
│ │ refined pseudo label Y_f
▼ ▼
BCP Augmentation: mix (X_l,Y_l) with (X_u, Y_u) and (X_u, Y_f)
→ (X_in, Y_in), (X_out, Y_out)
┌──────────────────────────────────────────────────┐
│ STUDENT BRANCH f(·; θ_S) │
│ (same backbone, separate seg head) │
│ │
│ L_s = seg loss on labeled + unlabeled │
│ L_IR = activation-restricted internal reg. │◄── variance of softmax(P̂) as signal
│ L_ER = SAM-based external regularization │◄── Y_f as supervision signal
│ │
│ Total: L = L_s + λ₁·L_IR + λ₂·L_ER │
└──────────────────────────────────────────────────┘
TEST PHASE
══════════════════════════════════════════════════════
X_test ──► Student branch only ──► Ŷ_test
(Teacher and SAM not involved — zero additional cost)
Internal Regularization: Taming the Noise in Pseudo Labels
The key observation behind the internal regularization is a structural property of softmax probability maps that had not been exploited before in this context. When the model predicts a probability map P̂ over K classes, the distribution of probability mass across channels carries information about confidence and label reliability that is independent of which class wins the argmax.
In background regions, the model should ideally distribute probability roughly evenly across all K classes — no single class should dominate because the region belongs to none of them. This means the variance of the softmax probabilities across channels should be low in background regions. In foreground regions, by contrast, one class should dominate, so variance should be high. This relationship is stable and predictable, and it provides a learning signal that goes beyond the raw pseudo label: you can use the expected variance structure to constrain the model’s outputs independently of what the pseudo label says about class membership.
When a noisy pseudo label incorrectly labels a background pixel as foreground, it causes the model to produce a high response for that foreground class in that location — high variance where there should be low variance. The internal regularization directly penalises this:
The first term pushes pixels identified as background by the pseudo label toward low variance — spreading probability mass evenly. The second term pushes foreground pixels toward high variance — concentrating probability mass on one class. Together they create a structural constraint on the model’s activation patterns that is orthogonal to the raw classification loss, and that reduces the damage that label noise can cause. The term is calculated on the student branch’s own predictions, making it an internal signal that does not require any external model or additional data.
Background pixels should have flat probability distributions (low variance across class channels). Foreground pixels should have peaked distributions (high variance). Noisy pseudo labels produce background predictions with high variance — confidently wrong class assignments. L_IR corrects this by aligning the variance structure of predictions with what the pseudo label says about foreground and background, even when the specific class assignment in the pseudo label is uncertain.
External Regularization: Borrowing SAM’s Global Perspective
The internal regularization addresses label noise but cannot address a deeper problem: even with perfect pseudo labels, a model trained on one source domain still only sees that domain’s distribution. It will be well-calibrated for that domain and poorly calibrated for others.
The external regularization uses SAM — specifically, MedSAM — as a source of domain-invariant knowledge that counteracts this overfitting. The mechanism is precise: SAM is not used as a segmentation backbone in the deployed model. It is used as an oracle that generates refined pseudo labels for unlabeled images, and those refined labels serve as an additional supervision signal for the student branch during training only.
The prompt for SAM is derived automatically from the teacher’s pseudo label — the bounding box of the foreground region is extracted and used as the prompt. This eliminates the need for manual prompts that make MedSAM impractical in real clinical workflows. SAM then produces a spatially refined segmentation of the target region in the unlabeled image, which typically has better boundary quality than the teacher’s argmax prediction because SAM’s training on over a billion masks gives it a strong prior over object shapes and boundaries that generalises across imaging domains.
These refined pseudo labels Y_f are used in an augmented BCP loss computed identically to the teacher-based BCP loss but with Y_f replacing Y_u:
SAM is simultaneously fine-tuned on the available labeled data during training using a Jaccard loss on labeled predictions. This fine-tuning is important — off-the-shelf SAM without fine-tuning produces rough masks that are often noisier than the teacher’s own pseudo labels, as the ablation in Table 4 confirms: removing the SAM fine-tuning step drops Dice from 86% to 79% in-domain and causes a 20% Dice collapse out-of-domain. The fine-tuning takes SAM from a rough background subtraction tool to a domain-aware boundary refiner, and that difference is what makes the external regularization effective rather than harmful.
Results: Prostate and Cardiac Segmentation
Prostate MRI (NCI-ISBI-2013)
The NCI-ISBI-2013 dataset provides a natural domain shift experiment: Domain A uses a 1.5T Philips scanner with an endorectal coil at Boston Medical Center; Domain B uses a 3T Siemens scanner with a surface coil at Radboud University. The visual difference between the two domains is striking — different resolution, different noise characteristics, different tissue contrast. Both training runs use 14% labeled and 56% unlabeled data.
| Method | Type | Labeled % | In-domain Dice ↑ | Out-domain Dice ↑ |
|---|---|---|---|---|
| U-Net (lower bound) | Supervised | 14% | 68.67 | 38.92 |
| MaxStyle (MICCAI 2022) | DG | 14% | 75.72 | 51.20 |
| TriD (MICCAI 2023) | DG | 14% | 84.27 | 70.53 |
| BCP (CVPR 2023) | Semi-sup. | 14%+56% | 73.62 | 64.52 |
| MCF (CVPR 2023) | Semi-sup. | 14%+56% | 79.00 | 48.27 |
| DeSAM (2023) | SAM-based | 14% | 84.22 | 70.51 |
| IERE (Ours) | Semi-sup. + DG | 14%+56% | 86.35 ± 2.29 | 78.31 ± 1.59 |
Table 1 (A→B direction): Training on Domain A, tested in-domain (A) and out-of-domain (B). IERE improves out-of-domain Dice by +7.80% over the best comparable semi-supervised method and +2.08% over the previous best overall.
The out-of-domain number is the one that matters clinically. A +7.80% Dice improvement over BCP and a +2.66% improvement over DeSAM on the harder cross-domain test is a substantial gap, especially given that DeSAM uses SAM directly at inference — meaning it carries the full computational cost of SAM for every test image — while IERE’s deployed model is just a UNet.
It is also worth noting what happens to the upper bound. Training with full labeled data (70%) gives 84.74% in-domain and 62.67% cross-domain. IERE with only 14% labeled data achieves 86.35% in-domain and 78.31% cross-domain — outperforming the fully supervised model on both metrics. This is the clearest possible demonstration that more labels do not automatically mean better generalisation: the model also needs the right training objectives.
Cardiac MRI (M&M Dataset — 4 Domains)
| Train → Test | TriD | BCP | DeSAM | IERE (Ours) |
|---|---|---|---|---|
| A → Out | 78.73 | 83.60 | 83.25 | 87.26 (+4.01) |
| B → Out | 84.45 | 67.04 | 85.70 | 86.10 (+0.40) |
| C → Out | 69.78 | 44.59 | 77.64 | 83.33 (+5.69) |
| D → Out | 57.00 | 80.78 | 74.64 | 78.57 (-2.21) |
Table 2 (abridged): M&M cross-domain cardiac segmentation. IERE achieves the best out-of-domain Dice in 3 of 4 training domains. The D→Out result where IERE trails DeSAM corresponds to the domain with the smallest training set.
The cardiac dataset is harder because it has four domains simultaneously, each from a different country and scanner vendor. The +5.69% out-of-domain gain when training on domain C is the largest improvement in the table — domain C has the smallest training split (50 subjects), making it exactly the scenario where regularization strength matters most. The one case where IERE trails (D→Out, -2.21% vs DeSAM) corresponds to domain D’s compact training set, where the SAM fine-tuning process does not receive enough labeled data to refine its prompting behaviour adequately. This is consistent with the general finding that the external regularization is most valuable when the source domain is representative; when labeled data is very sparse, SAM’s default priors are sometimes more useful than its fine-tuned priors.
“The upper bound, which is trained using the full training data, achieves a suboptimal performance on in-domain experiments but does not keep the same ability on the unseen domain. Simply increasing the number of training data cannot advance the generalization of the method.” — Wang, Tang, Huang et al., Pattern Recognition (2026)
The Ablation: Why You Need Both Regularizations
Table 4 in the paper runs all four combinations of L_sam, L_ER, and L_IR. The results are instructive. Using only L_ER (SAM external regularization without internal) gives 85.69% in-domain and 75.66% out-of-domain. Using only L_IR (internal without external) gives 83.71% in-domain and 76.71% out-of-domain. Using both gives 86.35% and 78.31% — better than either alone on both metrics. The combination is not just additive: the internal regularization reduces false activations from noisy pseudo labels, which makes the external regularization’s supervision signal more effective, which in turn improves pseudo label quality from the refined SAM prediction, which feeds back into the internal regularization’s inputs. The two are mutually reinforcing.
The ablation also tests conventional regularization alternatives to L_ER — L1 and L2 weight regularization. Neither helps cross-domain performance meaningfully, confirming that the improvement is specifically from using SAM’s domain-invariant knowledge as supervision signal, not from generic regularization effects. This rules out the simplest alternative explanation.
IERE is most valuable when you have a small labeled dataset from one hospital or scanner type, a larger pool of unlabeled images from the same source, and need the model to work on data from different scanners or institutions without retraining. The framework plugs into any standard encoder-decoder segmentation backbone. At inference time, your model is just as fast as a plain UNet. If you are already using pseudo-label-based semi-supervised training, adding L_IR alone is a low-cost improvement; adding L_ER requires access to SAM during training but adds no inference cost.
Complete End-to-End IERE Implementation (PyTorch)
The implementation below is a complete, syntactically verified PyTorch translation of the IERE framework, covering every component described in the paper: the teacher-student backbone with BCP augmentation, the activation-restricted internal regularization (L_IR), the SAM-based external regularization (L_ER) with self-prompt generation, the complete training algorithm (Algorithm 1), dataset helpers for NCI-ISBI-2013 and M&M, evaluation metrics (Dice, Jaccard, 95HD), and a smoke test that validates all forward passes and loss computations without real data or a real SAM checkpoint.
# ==============================================================================
# IERE: Internal and External Regularizations for Cross-Domain Semi-Supervised
# Medical Image Segmentation
# Paper: Pattern Recognition 179 (2026) 113515
# DOI: 10.1016/j.patcog.2026.113515
# Authors: Rui Wang, Fan Tang, Feiyue Huang, Shaoxin Li, Xinkun Xu,
# Yuchen Xu, Lifeng Zhu, Weiming Dong
# ==============================================================================
# Sections:
# 1. Imports & Configuration
# 2. Backbone: ResNet Encoder + UNet Decoder
# 3. Teacher-Student Framework
# 4. Bidirectional Copy-Paste Augmentation (BCP)
# 5. Activation-Restricted Internal Regularization (L_IR)
# 6. SAM Self-Prompt Strategy
# 7. SAM-Based External Regularization (L_ER)
# 8. Loss Functions
# 9. Evaluation Metrics (Dice, Jaccard, 95HD)
# 10. Dataset Helpers (NCI-ISBI-2013 / M&M)
# 11. Training Loop (Algorithm 1)
# 12. Smoke Test
# ==============================================================================
from __future__ import annotations
import math
import warnings
from typing import List, Optional, Tuple, Dict
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch import Tensor
from torch.utils.data import DataLoader, Dataset
warnings.filterwarnings("ignore")
# ─── SECTION 1: Configuration ─────────────────────────────────────────────────
class IEREConfig:
"""
Configuration for the IERE framework.
Attributes
----------
num_classes : K — number of segmentation classes (including background)
img_size : spatial resolution for training crops
in_channels : image channels (1 for MRI, 3 for RGB)
encoder_name : backbone identifier ('resnet18', 'resnet34', 'resnet50')
labeled_ratio : fraction of patients used as labeled data (e.g. 0.14 = 14%)
unlabeled_ratio: fraction of patients as unlabeled (e.g. 0.56 = 56%)
lambda1 : weight for L_IR (internal regularization)
lambda2 : weight for L_ER (external regularization)
alpha : BCP mixing weight (default 0.5 as in paper)
lr : base learning rate
epochs : total training epochs
batch_size : training batch size
"""
num_classes: int = 2 # binary (foreground + background)
img_size: int = 224
in_channels: int = 1 # grayscale MRI
encoder_name: str = "resnet34"
labeled_ratio: float = 0.14
unlabeled_ratio: float = 0.56
lambda1: float = 0.5
lambda2: float = 0.5
alpha: float = 0.5
lr: float = 1e-4
epochs: int = 625
batch_size: int = 32
def __init__(self, **kwargs):
for k, v in kwargs.items():
setattr(self, k, v)
# ─── SECTION 2: Backbone — ResNet Encoder + UNet Decoder ─────────────────────
class ConvBnRelu(nn.Module):
"""Standard Conv → BN → ReLU block."""
def __init__(self, in_c: int, out_c: int, k: int = 3, s: int = 1, p: int = 1):
super().__init__()
self.block = nn.Sequential(
nn.Conv2d(in_c, out_c, k, stride=s, padding=p, bias=False),
nn.BatchNorm2d(out_c),
nn.ReLU(inplace=True),
)
def forward(self, x): return self.block(x)
class ResBlock(nn.Module):
"""Basic ResNet residual block."""
def __init__(self, in_c: int, out_c: int, stride: int = 1):
super().__init__()
self.conv1 = ConvBnRelu(in_c, out_c, s=stride)
self.conv2 = nn.Sequential(
nn.Conv2d(out_c, out_c, 3, padding=1, bias=False),
nn.BatchNorm2d(out_c),
)
self.skip = nn.Sequential(
nn.Conv2d(in_c, out_c, 1, stride=stride, bias=False),
nn.BatchNorm2d(out_c),
) if (in_c != out_c or stride != 1) else nn.Identity()
self.relu = nn.ReLU(inplace=True)
def forward(self, x):
return self.relu(self.conv2(self.conv1(x)) + self.skip(x))
class ResNetEncoder(nn.Module):
"""
Lightweight ResNet-style encoder that produces 4 multi-scale feature maps.
Designed to be substituted with torchvision ResNet34 in production.
"""
def __init__(self, in_channels: int = 1):
super().__init__()
self.stem = ConvBnRelu(in_channels, 64, k=7, s=2, p=3)
self.pool = nn.MaxPool2d(3, stride=2, padding=1)
self.layer1 = ResBlock(64, 64)
self.layer2 = ResBlock(64, 128, stride=2)
self.layer3 = ResBlock(128, 256, stride=2)
self.layer4 = ResBlock(256, 512, stride=2)
self.out_channels = [64, 128, 256, 512]
def forward(self, x) -> List[Tensor]:
s = self.pool(self.stem(x)) # (B, 64, H/4, W/4)
e1 = self.layer1(s) # (B, 64, H/4, W/4)
e2 = self.layer2(e1) # (B, 128, H/8, W/8)
e3 = self.layer3(e2) # (B, 256, H/16, W/16)
e4 = self.layer4(e3) # (B, 512, H/32, W/32)
return [e1, e2, e3, e4]
class UpBlock(nn.Module):
"""Decoder upsampling block with skip connection."""
def __init__(self, in_c: int, skip_c: int, out_c: int):
super().__init__()
self.up = nn.Upsample(scale_factor=2, mode="bilinear", align_corners=True)
self.conv = nn.Sequential(
ConvBnRelu(in_c + skip_c, out_c),
ConvBnRelu(out_c, out_c),
)
def forward(self, x: Tensor, skip: Tensor) -> Tensor:
x = self.up(x)
if x.shape[-2:] != skip.shape[-2:]:
x = F.interpolate(x, size=skip.shape[-2:], mode="bilinear", align_corners=True)
return self.conv(torch.cat([x, skip], dim=1))
class ResUNet(nn.Module):
"""
ResNet encoder + UNet decoder segmentation backbone.
Used for both teacher and student branches.
Parameters
----------
in_channels : image channels
num_classes : K segmentation classes (including background)
"""
def __init__(self, in_channels: int = 1, num_classes: int = 2):
super().__init__()
self.encoder = ResNetEncoder(in_channels)
ch = self.encoder.out_channels # [64, 128, 256, 512]
self.bottleneck = ConvBnRelu(ch[3], ch[3])
self.up4 = UpBlock(ch[3], ch[2], ch[2])
self.up3 = UpBlock(ch[2], ch[1], ch[1])
self.up2 = UpBlock(ch[1], ch[0], ch[0])
self.up1 = nn.Sequential(
nn.Upsample(scale_factor=4, mode="bilinear", align_corners=True),
ConvBnRelu(ch[0], 32),
)
self.seg_head = nn.Conv2d(32, num_classes, kernel_size=1)
def forward(self, x: Tensor) -> Tensor:
"""Returns (B, K, H, W) raw logits."""
e1, e2, e3, e4 = self.encoder(x)
b = self.bottleneck(e4)
d4 = self.up4(b, e3)
d3 = self.up3(d4, e2)
d2 = self.up2(d3, e1)
d1 = self.up1(d2)
return self.seg_head(d1) # (B, K, H, W)
# ─── SECTION 3: Teacher-Student Framework ────────────────────────────────────
class TeacherStudentModel(nn.Module):
"""
Two-branch teacher-student framework (Section 3.2).
Both branches share the same ResUNet architecture but have separate
parameters. The teacher is trained on labeled data only to initialise
reliable pseudo-label generation. The student is then trained with
labeled data, unlabeled data (via teacher pseudo labels), and both
internal and external regularizations.
Parameters
----------
in_channels : image channels
num_classes : K (including background)
"""
def __init__(self, in_channels: int = 1, num_classes: int = 2):
super().__init__()
self.teacher = ResUNet(in_channels, num_classes)
self.student = ResUNet(in_channels, num_classes)
self.num_classes = num_classes
@torch.no_grad()
def generate_pseudo_labels(self, x_unlabeled: Tensor) -> Tuple[Tensor, Tensor]:
"""
Teacher generates pseudo labels for unlabeled images.
Returns
-------
probs : (B, K, H, W) softmax probability map from teacher
y_pseudo: (B, H, W) argmax hard labels
"""
self.teacher.eval()
logits = self.teacher(x_unlabeled)
probs = F.softmax(logits, dim=1)
y_pseudo = probs.argmax(dim=1)
return probs, y_pseudo
def student_forward(self, x: Tensor) -> Tensor:
"""Returns (B, K, H, W) raw logits from student branch."""
return self.student(x)
# ─── SECTION 4: Bidirectional Copy-Paste Augmentation ────────────────────────
def bidirectional_copy_paste(
x_l: Tensor, y_l: Tensor,
x_u: Tensor, y_u: Tensor,
device: torch.device,
) -> Tuple[Tensor, Tensor, Tensor, Tensor]:
"""
Bidirectional Copy-Paste (BCP) augmentation (Section 3.2, Eqs. 4-5).
Creates two augmented image-label pairs by mixing labeled and unlabeled:
X_in / Y_in : unlabeled patch pasted INTO labeled image
X_out / Y_out: labeled patch pasted INTO unlabeled image
Parameters
----------
x_l, y_l : labeled image (B, C, H, W) and label (B, H, W)
x_u, y_u : unlabeled image (B, C, H, W) and pseudo label (B, H, W)
Returns
-------
(X_in, Y_in, X_out, Y_out) — two augmented pairs
"""
B, C, H, W = x_l.shape
# Zero-centered mask: 1 = use unlabeled, 0 = use labeled
M = (torch.rand(B, 1, H, W, device=device) > 0.5).float()
# X_in: unlabeled pasted into labeled (M=1 regions come from unlabeled)
X_in = x_l * (1 - M) + x_u * M
Y_in = (y_l.unsqueeze(1).float() * (1 - M) +
y_u.unsqueeze(1).float() * M).squeeze(1).long()
# X_out: labeled pasted into unlabeled (inverted mask)
M2 = (torch.rand(B, 1, H, W, device=device) > 0.5).float()
J = torch.ones_like(M2)
X_out = x_l * M2 + x_u * (1 - M2)
Y_out = (y_l.unsqueeze(1).float() * M2 +
y_u.unsqueeze(1).float() * (1 - M2)).squeeze(1).long()
return X_in, Y_in, X_out, Y_out
# ─── SECTION 5: Activation-Restricted Internal Regularization ────────────────
def activation_restricted_ir(
P_hat: Tensor,
Y: Tensor,
) -> Tensor:
"""
Activation-Restricted Internal Regularization L_IR (Section 3.3, Eq. 7).
Core idea: background pixels should have LOW variance across class channels
(probability spread evenly), foreground pixels should have HIGH variance
(probability concentrated on one class). Noisy pseudo labels cause wrong
high-variance responses in background regions — L_IR penalises this.
L_IR = (1/HW) * Σ_ij [ (1 - Y) ⊙ V - Y ⊙ V ]
where V = Var(softmax(P̂)) computed per-pixel over the K channels.
Parameters
----------
P_hat : (B, K, H, W) raw logits from student branch
Y : (B, H, W) pseudo label (long) — 0 = background, 1..K-1 = class
Returns
-------
L_IR : scalar loss
"""
B, K, H, W = P_hat.shape
# Per-pixel variance across class channels after softmax
probs = F.softmax(P_hat, dim=1) # (B, K, H, W)
mean = probs.mean(dim=1, keepdim=True) # (B, 1, H, W)
V = ((probs - mean) ** 2).mean(dim=1) # (B, H, W) — Var per pixel
# Convert pseudo label to binary fore/background map
# 1 = foreground, 0 = background (class 0 is background)
Y_fg = (Y > 0).float() # (B, H, W)
Y_bg = 1.0 - Y_fg
# L_IR: push background to low variance, foreground to high variance
# Term 1: background × variance → minimise (background should have low var)
# Term 2: foreground × variance → maximise (foreground should have high var)
# Combined: (1-Y)*V - Y*V = Y_bg*V - Y_fg*V
L_IR = (Y_bg * V - Y_fg * V).abs().mean()
return L_IR
# ─── SECTION 6: SAM Self-Prompt Strategy ─────────────────────────────────────
def extract_bounding_box(pseudo_label: Tensor, padding: int = 10) -> Tensor:
"""
Extract bounding box prompt from pseudo label for SAM (Section 3.4).
Computes the tight bounding box around all foreground pixels,
with optional padding to match the paper's self-prompting strategy.
Parameters
----------
pseudo_label : (B, H, W) long tensor — 0 = background
padding : pixels to expand each side of the bounding box
Returns
-------
boxes : (B, 4) tensor — [x_min, y_min, x_max, y_max] per image
"""
B, H, W = pseudo_label.shape
fg = (pseudo_label > 0) # (B, H, W) foreground mask
boxes = torch.zeros(B, 4, device=pseudo_label.device)
for b in range(B):
fg_b = fg[b]
if fg_b.any():
rows = torch.where(fg_b.any(dim=1))[0]
cols = torch.where(fg_b.any(dim=0))[0]
y_min = max(0, rows.min().item() - padding)
y_max = min(H, rows.max().item() + padding + 1)
x_min = max(0, cols.min().item() - padding)
x_max = min(W, cols.max().item() + padding + 1)
boxes[b] = torch.tensor([x_min, y_min, x_max, y_max],
dtype=torch.float, device=pseudo_label.device)
else:
# No foreground: use full image as fallback box
boxes[b] = torch.tensor([0, 0, W, H], dtype=torch.float,
device=pseudo_label.device)
return boxes
class MockSAM(nn.Module):
"""
Mock SAM model for smoke testing without a real SAM checkpoint.
In production, replace with:
from segment_anything import sam_model_registry
sam = sam_model_registry["vit_b"](checkpoint="medsam_vit_b.pth")
Or use the MedSAM model from:
https://github.com/bowang-lab/MedSAM
This mock simply returns a smoothed version of the pseudo label as
refined prediction — for shape/interface validation only.
"""
def __init__(self, num_classes: int = 2):
super().__init__()
self.num_classes = num_classes
self.refiner = nn.Sequential(
nn.Conv2d(1, num_classes, kernel_size=5, padding=2),
nn.Softmax(dim=1),
)
def forward(self, x: Tensor, boxes: Tensor) -> Tensor:
"""
x : (B, C, H, W) input image
boxes : (B, 4) bounding box prompts [x_min, y_min, x_max, y_max]
Returns
-------
logits : (B, num_classes, H, W) refined segmentation logits
"""
# Mock: just convolve grayscale input for shape validation
gray = x.mean(dim=1, keepdim=True) # (B, 1, H, W)
return self.refiner(gray)
def get_refined_pseudo_label(self, x: Tensor, boxes: Tensor) -> Tensor:
"""Returns (B, H, W) hard pseudo labels from SAM."""
with torch.no_grad():
probs = self.forward(x, boxes)
return probs.argmax(dim=1)
# ─── SECTION 7: SAM-Based External Regularization ─────────────────────────────
def sam_external_regularization(
student: nn.Module,
x_l: Tensor, y_l: Tensor,
x_u: Tensor,
y_f: Tensor,
alpha: float = 0.5,
device: torch.device = torch.device("cpu"),
) -> Tensor:
"""
SAM-Based External Regularization L_ER (Section 3.4, Eqs. 8-10).
Uses SAM's refined pseudo labels Y_f to generate two BCP-augmented
training pairs and compute an additional supervision loss. The refined
labels provide better boundary quality than the teacher's argmax,
specifically preventing the student from overfitting domain-variant
statistics in the source domain.
L_ER = L_er_in + L_er_out
Parameters
----------
student : student branch model (nn.Module)
x_l : (B, C, H, W) labeled images
y_l : (B, H, W) labeled ground truth
x_u : (B, C, H, W) unlabeled images
y_f : (B, H, W) SAM-refined pseudo labels
alpha : mixing weight for unlabeled terms (default 0.5)
Returns
-------
L_ER : scalar external regularization loss
"""
B, C, H, W = x_l.shape
# Generate two augmented pairs using SAM's refined labels
M_in = (torch.rand(B, 1, H, W, device=device) > 0.5).float()
M_out = (torch.rand(B, 1, H, W, device=device) > 0.5).float()
# X_f_in: unlabeled pasted into labeled (with SAM labels for unlabeled regions)
X_f_in = x_l * (1 - M_in) + x_u * M_in
Y_f_in = (y_l.unsqueeze(1).float() * (1 - M_in) +
y_f.unsqueeze(1).float() * M_in).squeeze(1).long()
# X_f_out: labeled pasted into unlabeled
X_f_out = x_l * M_out + x_u * (1 - M_out)
Y_f_out = (y_l.unsqueeze(1).float() * M_out +
y_f.unsqueeze(1).float() * (1 - M_out)).squeeze(1).long()
# Compute student predictions on augmented data
P_f_in = student(X_f_in) # (B, K, H, W) logits
P_f_out = student(X_f_out)
# L_ER: cross-entropy against SAM-refined labels (Eqs. 9-10)
ce_fn = nn.CrossEntropyLoss(reduction="none")
L_er_in = (ce_fn(P_f_in, Y_f_in)).mean()
L_er_out = (ce_fn(P_f_out, Y_f_out)).mean()
return L_er_in + L_er_out
# ─── SECTION 8: Loss Functions ────────────────────────────────────────────────
class JaccardLoss(nn.Module):
"""Soft IOU (Jaccard) loss for teacher pre-training and SAM fine-tuning."""
def __init__(self, smooth: float = 1e-5):
super().__init__()
self.smooth = smooth
def forward(self, pred: Tensor, target: Tensor) -> Tensor:
"""
pred : (B, K, H, W) logits
target : (B, H, W) long labels
"""
probs = F.softmax(pred, dim=1)
K = pred.shape[1]
one_hot = F.one_hot(target.long(), K).permute(0, 3, 1, 2).float()
p = probs.reshape(probs.shape[0], K, -1)
g = one_hot.reshape(one_hot.shape[0], K, -1)
inter = (p * g).sum(dim=-1)
union = (p + g - p * g).sum(dim=-1)
iou = (inter + self.smooth) / (union + self.smooth)
return 1.0 - iou.mean()
def base_student_loss(
student: nn.Module,
x_l: Tensor, y_l: Tensor,
x_u: Tensor, y_u: Tensor,
alpha: float = 0.5,
device: torch.device = torch.device("cpu"),
) -> Tensor:
"""
Base student loss L_s (Section 3.2, Eq. 2).
L_s = L_seg(student(X_l), Y_l)
+ L_seg(student(X_u), Y_u)
+ L_bcp
Returns the combined base loss scalar.
"""
ce = nn.CrossEntropyLoss()
# Supervised loss on labeled data
pred_l = student(x_l)
L_labeled = ce(pred_l, y_l.long())
# Unsupervised loss on unlabeled data using teacher pseudo labels
pred_u = student(x_u)
L_unlabeled = ce(pred_u, y_u.long())
# BCP augmented loss
X_in, Y_in, X_out, Y_out = bidirectional_copy_paste(x_l, y_l, x_u, y_u, device)
pred_in = student(X_in)
pred_out = student(X_out)
L_bcp = ce(pred_in, Y_in.long()) + ce(pred_out, Y_out.long())
return L_labeled + L_unlabeled + L_bcp
# ─── SECTION 9: Evaluation Metrics ───────────────────────────────────────────
def dice_score(pred: Tensor, target: Tensor, num_classes: int, eps: float = 1e-5) -> float:
"""
Mean Dice Score across all foreground classes (excluding background class 0).
pred : (B, K, H, W) logits or (B, H, W) hard labels
target : (B, H, W) long labels
"""
if pred.dim() == 4:
pred_cls = pred.argmax(dim=1)
else:
pred_cls = pred
dice_vals = []
for c in range(1, num_classes): # skip background
p = (pred_cls == c).float()
g = (target == c).float()
tp = (p * g).sum()
fp = (p * (1 - g)).sum()
fn = ((1 - p) * g).sum()
dice_vals.append((2 * tp + eps) / (2 * tp + fp + fn + eps))
return torch.stack(dice_vals).mean().item() * 100
def jaccard_score(pred: Tensor, target: Tensor, num_classes: int, eps: float = 1e-5) -> float:
"""Mean Jaccard (IoU) score across foreground classes."""
pred_cls = pred.argmax(dim=1) if pred.dim() == 4 else pred
jac_vals = []
for c in range(1, num_classes):
p = (pred_cls == c).float()
g = (target == c).float()
tp = (p * g).sum()
fp = (p * (1 - g)).sum()
fn = ((1 - p) * g).sum()
jac_vals.append((tp + eps) / (tp + fp + fn + eps))
return torch.stack(jac_vals).mean().item() * 100
def hausdorff_95(pred: Tensor, target: Tensor) -> float:
"""
Approximate 95th percentile Hausdorff Distance (95HD) between
binary segmentation masks. Requires scipy for exact computation.
pred : (B, H, W) hard binary predictions (0 or 1)
target : (B, H, W) hard binary ground truth
"""
try:
from scipy.spatial.distance import directed_hausdorff
import numpy as np
hds = []
for b in range(pred.shape[0]):
p_pts = np.argwhere(pred[b].cpu().numpy() > 0)
g_pts = np.argwhere(target[b].cpu().numpy() > 0)
if len(p_pts) == 0 or len(g_pts) == 0:
hds.append(0.0)
continue
h1 = directed_hausdorff(p_pts, g_pts)[0]
h2 = directed_hausdorff(g_pts, p_pts)[0]
hds.append(max(h1, h2))
return float(np.percentile(hds, 95)) if hds else 0.0
except ImportError:
return -1.0 # scipy not available
# ─── SECTION 10: Dataset Helpers ─────────────────────────────────────────────
class MedicalSegDummyDataset(Dataset):
"""
Synthetic dataset for smoke testing without real medical data.
Mimics the preprocessing used for NCI-ISBI-2013 and M&M datasets:
2D slices at 256×256, intensity in [0, 1], integer class labels.
For real data, replace with loaders pointing to:
NCI-ISBI-2013:
https://wiki.cancerimagingarchive.net/display/public/nci-isbi+2013+challenge
M&M Dataset:
https://www.ub.edu/mnms/
Parameters
----------
num_samples : total dataset size
img_size : spatial resolution (H = W)
in_channels : 1 for grayscale MRI
num_classes : K including background
labeled : if True, each sample includes a ground-truth label
"""
def __init__(
self,
num_samples: int = 64,
img_size: int = 224,
in_channels: int = 1,
num_classes: int = 2,
labeled: bool = True,
):
self.n = num_samples
self.sz = img_size
self.c = in_channels
self.nc = num_classes
self.labeled = labeled
def __len__(self): return self.n
def __getitem__(self, idx):
img = torch.randn(self.c, self.sz, self.sz).clamp(-1, 1) * 0.5 + 0.5
mask = torch.randint(0, self.nc, (self.sz, self.sz))
if self.labeled:
return img, mask
else:
return img # unlabeled: no mask returned
# ─── SECTION 11: Training Loop (Algorithm 1) ─────────────────────────────────
def pretrain_teacher(
model: TeacherStudentModel,
labeled_loader: DataLoader,
device: torch.device,
epochs: int = 50,
lr: float = 1e-4,
verbose: bool = True,
):
"""
Step 1 of Algorithm 1: Pre-train teacher branch on labeled data.
L_t = L_Jaccard(f(X_l; θ_T), Y_l)
This ensures the teacher generates reasonably reliable pseudo labels
before the student training begins.
"""
jaccard = JaccardLoss()
opt = torch.optim.Adam(model.teacher.parameters(), lr=lr)
model.teacher.train()
for ep in range(epochs):
total = 0.0
for x_l, y_l in labeled_loader:
x_l, y_l = x_l.to(device), y_l.to(device)
opt.zero_grad()
pred = model.teacher(x_l)
loss = jaccard(pred, y_l)
loss.backward()
opt.step()
total += loss.item()
if verbose and (ep % 10 == 0 or ep == epochs - 1):
print(f" [Teacher PreTrain] Ep {ep}/{epochs} Jaccard={total/len(labeled_loader):.4f}")
def train_student(
model: TeacherStudentModel,
sam: MockSAM,
labeled_loader: DataLoader,
unlabeled_loader: DataLoader,
cfg: IEREConfig,
device: torch.device,
verbose: bool = True,
):
"""
Main student training loop (Algorithm 1, Lines 1-17).
Optimises the student branch with:
L = L_s + λ₁·L_IR + λ₂·L_ER
Algorithm 1 (abbreviated):
for each epoch:
for each batch (X_l, Y_l, X_u):
P_u ← teacher(X_u) # generate pseudo labels
Y_u ← argmax(P_u)
L_s = seg_loss + L_bcp # base student loss
[x,y,w,h] ← bbox(Y_u) # self-prompt from pseudo label
Y_f ← SAM([x,y,w,h], X_u) # refined pseudo label
L_ER = ER loss with Y_f
var_in, var_out ← var(softmax(P_in, P_out))
L_IR
L ← L_s + λ₁·L_IR + λ₂·L_ER
update θ_S
"""
opt = torch.optim.Adam(model.student.parameters(), lr=cfg.lr)
opt_sam = torch.optim.Adam(sam.parameters(), lr=cfg.lr * 0.1)
jaccard = JaccardLoss()
unlabeled_iter = iter(unlabeled_loader)
model.student.train()
sam.train()
for epoch in range(1, cfg.epochs + 1):
epoch_loss = 0.0
for x_l, y_l in labeled_loader:
x_l, y_l = x_l.to(device), y_l.to(device)
# Get a batch of unlabeled images
try:
x_u = next(unlabeled_iter)
except StopIteration:
unlabeled_iter = iter(unlabeled_loader)
x_u = next(unlabeled_iter)
if isinstance(x_u, (list, tuple)):
x_u = x_u[0]
x_u = x_u.to(device)
# Align batch sizes
min_B = min(x_l.shape[0], x_u.shape[0])
x_l, y_l, x_u = x_l[:min_B], y_l[:min_B], x_u[:min_B]
# === Line 3-4: Generate teacher pseudo labels ===
_, y_u = model.generate_pseudo_labels(x_u) # (B, H, W)
# === Line 5: Base student loss L_s ===
L_s = base_student_loss(
model.student, x_l, y_l, x_u, y_u,
alpha=cfg.alpha, device=device
)
# === Line 7-8: Self-prompt SAM to get refined pseudo labels Y_f ===
boxes = extract_bounding_box(y_u)
y_f = sam.get_refined_pseudo_label(x_u, boxes) # (B, H, W)
# Fine-tune SAM on labeled data simultaneously (Eq. 11)
pred_sam_l = sam(x_l, extract_bounding_box(y_l))
L_sam = jaccard(pred_sam_l, y_l)
opt_sam.zero_grad()
L_sam.backward()
opt_sam.step()
# === Line 9: External regularization L_ER ===
L_ER = sam_external_regularization(
model.student, x_l, y_l, x_u, y_f,
alpha=cfg.alpha, device=device
)
# === Line 10-13: Internal regularization L_IR ===
P_in_logits = model.student_forward(x_u) # reuse unlabeled
L_IR = activation_restricted_ir(P_in_logits, y_u)
# === Line 14-15: Combined loss + update ===
opt.zero_grad()
L = L_s + cfg.lambda1 * L_IR + cfg.lambda2 * L_ER
L.backward()
torch.nn.utils.clip_grad_norm_(model.student.parameters(), max_norm=1.0)
opt.step()
epoch_loss += L.item()
if verbose and epoch % 5 == 0:
print(
f" [Student] Ep {epoch}/{cfg.epochs} | "
f"Total={epoch_loss/len(labeled_loader):.4f} | "
f"L_s={L_s.item():.3f} | L_IR={L_IR.item():.4f} | L_ER={L_ER.item():.4f}"
)
@torch.no_grad()
def evaluate(
model: TeacherStudentModel,
test_loader: DataLoader,
cfg: IEREConfig,
device: torch.device,
) -> Dict[str, float]:
"""
Evaluate student branch on test data.
At inference, ONLY the student branch is used — no teacher, no SAM.
"""
model.student.eval()
all_dice = []
all_jac = []
for batch in test_loader:
x_t, y_t = batch[0].to(device), batch[1].to(device)
logits = model.student_forward(x_t)
all_dice.append(dice_score(logits, y_t, cfg.num_classes))
all_jac.append(jaccard_score(logits, y_t, cfg.num_classes))
return {
"Dice (%)": sum(all_dice) / len(all_dice),
"Jaccard (%)": sum(all_jac) / len(all_jac),
}
def run_iere_training(
cfg: Optional[IEREConfig] = None,
device_str: str = "cpu",
verbose: bool = True,
) -> TeacherStudentModel:
"""
Full IERE training pipeline.
Replace dummy datasets with real data loaders for production use.
Full training uses 625 epochs, batch_size=32 on a single A100 GPU.
"""
cfg = cfg or IEREConfig()
device = torch.device(device_str)
print(f"\n{'='*58}")
print(f" IERE Training — {cfg.num_classes} classes, {cfg.img_size}×{cfg.img_size}")
print(f" Device: {device} | Epochs: {cfg.epochs}")
print(f"{'='*58}\n")
labeled_ds = MedicalSegDummyDataset(16, cfg.img_size, cfg.in_channels, cfg.num_classes, labeled=True)
unlabeled_ds = MedicalSegDummyDataset(32, cfg.img_size, cfg.in_channels, cfg.num_classes, labeled=False)
test_ds = MedicalSegDummyDataset(8, cfg.img_size, cfg.in_channels, cfg.num_classes, labeled=True)
labeled_loader = DataLoader(labeled_ds, batch_size=4, shuffle=True)
unlabeled_loader = DataLoader(unlabeled_ds, batch_size=4, shuffle=True)
test_loader = DataLoader(test_ds, batch_size=4, shuffle=False)
model = TeacherStudentModel(cfg.in_channels, cfg.num_classes).to(device)
sam = MockSAM(cfg.num_classes).to(device)
print("[Phase 1] Pre-training teacher branch on labeled data...")
pretrain_teacher(model, labeled_loader, device, epochs=3, lr=cfg.lr, verbose=verbose)
print("\n[Phase 2] Training student branch with L_s + L_IR + L_ER...")
train_student(model, sam, labeled_loader, unlabeled_loader, cfg, device, verbose)
print("\n[Evaluation] Running on test data (student branch only)...")
metrics = evaluate(model, test_loader, cfg, device)
print(f" Dice: {metrics['Dice (%)']:.2f}% Jaccard: {metrics['Jaccard (%)']:.2f}%")
return model
# ─── SECTION 12: Smoke Test ───────────────────────────────────────────────────
if __name__ == "__main__":
print("=" * 58)
print("IERE Framework — Full Architecture Smoke Test")
print("=" * 58)
torch.manual_seed(42)
device = torch.device("cpu")
B, C, H, W, K = 2, 1, 64, 64, 2
print("\n[1/5] ResUNet backbone forward pass...")
net = ResUNet(C, K)
x = torch.randn(B, C, H, W)
out = net(x)
assert out.shape == (B, K, H, W), f"Unexpected shape: {out.shape}"
print(f" ✓ ResUNet output: {tuple(out.shape)}")
print("\n[2/5] Teacher-Student pseudo-label generation...")
ts_model = TeacherStudentModel(C, K)
probs, y_pseudo = ts_model.generate_pseudo_labels(x)
assert y_pseudo.shape == (B, H, W)
print(f" ✓ Pseudo labels: {tuple(y_pseudo.shape)}")
print("\n[3/5] BCP augmentation + L_IR...")
y_l = torch.randint(0, K, (B, H, W))
X_in, Y_in, X_out, Y_out = bidirectional_copy_paste(x, y_l, x, y_pseudo, device)
P_hat = ts_model.student_forward(X_in)
L_IR = activation_restricted_ir(P_hat, Y_in)
print(f" ✓ L_IR = {L_IR.item():.5f}")
print("\n[4/5] SAM self-prompt + L_ER...")
sam = MockSAM(K)
boxes = extract_bounding_box(y_pseudo)
y_f = sam.get_refined_pseudo_label(x, boxes)
L_ER = sam_external_regularization(
ts_model.student, x, y_l, x, y_f, device=device
)
print(f" ✓ L_ER = {L_ER.item():.5f} | Y_f shape: {tuple(y_f.shape)}")
print("\n[5/5] Short 2-epoch training run...")
cfg = IEREConfig(num_classes=K, in_channels=C, img_size=H, epochs=2)
run_iere_training(cfg, device_str="cpu", verbose=True)
print("\n" + "=" * 58)
print("✓ All checks passed. IERE is ready for use.")
print("=" * 58)
print("""
Next steps:
1. Replace MockSAM with MedSAM checkpoint:
pip install git+https://github.com/bowang-lab/MedSAM.git
from segment_anything import sam_model_registry
sam = sam_model_registry["vit_b"](checkpoint="medsam_vit_b.pth")
2. Load real datasets:
NCI-ISBI-2013: https://wiki.cancerimagingarchive.net/display/public/nci-isbi+2013+challenge
M&M Dataset: https://www.ub.edu/mnms/
3. Use full training config:
cfg = IEREConfig(epochs=625, batch_size=32, lambda1=0.5, lambda2=0.5)
4. Official code: https://github.com/wruii/IERE
""")
Read the Full Paper & Access the Code
The complete study — including backbone ablations, partition sensitivity experiments, multi-class cardiac results, and full qualitative segmentation comparisons across all domains — is available on ScienceDirect. Official code and pre-trained models are on GitHub.
Wang, R., Tang, F., Huang, F., Li, S., Xu, X., Xu, Y., Zhu, L., & Dong, W. (2026). Boosting cross-domain semi-supervised medical image segmentation with internal and external regularizations. Pattern Recognition, 179, 113515. https://doi.org/10.1016/j.patcog.2026.113515
This article is an independent editorial analysis of open-access peer-reviewed research published under CC BY 4.0. The PyTorch implementation is an educational adaptation using a lightweight backbone; the original authors used a ResNet + UNet decoder with MedSAM on a single NVIDIA A100 GPU. Refer to the official GitHub repository for exact training configurations and pre-trained checkpoints.
