In the high-stakes world of neuro-oncology, time is not just a factor—it’s a lifeline. The journey from an initial MRI scan to a definitive brain tumor diagnosis has long been fraught with delays, human error, and the immense cognitive load placed on radiologists who must interpret complex, often subtle, variations in medical imagery. This critical bottleneck in patient care is now being addressed by a groundbreaking fusion of artificial intelligence and cutting-edge transformer models. Introducing BrainDx, a dual-transformer framework that promises to revolutionize how we detect, classify, and segment brain tumors, offering unprecedented speed, accuracy, and reliability for clinicians and patients alike.
This article will delve deep into the architecture, methodology, and real-world performance of the BrainDx system, exploring how it leverages the complementary strengths of the Pyramid Vision Transformer (PVT) and SegFormer to overcome the limitations of traditional methods. We will dissect its technical underpinnings, analyze its impressive results across multiple benchmark datasets, and discuss its potential to transform clinical workflows. For healthcare professionals, researchers, and anyone interested in the future of precision medicine, this is a comprehensive guide to understanding one of the most significant advancements in medical AI today.
The Critical Need for AI in Brain Tumor Diagnosis
The diagnosis of brain tumors remains one of the most challenging tasks in modern medicine. Magnetic Resonance Imaging (MRI) scans, while incredibly detailed, present a formidable challenge due to the inherent complexity of tumor morphology. Tumors can exhibit indistinct boundaries, subtle variations in texture and intensity, and can mimic healthy tissue, making manual interpretation a highly subjective and time-consuming process.
The current diagnostic paradigm is fraught with inefficiencies:
- Time-Consuming Manual Analysis: A single MRI scan can contain hundreds of slices, each requiring meticulous examination by a trained radiologist. This process can take anywhere from 30 minutes to several hours, creating significant backlogs and delaying critical treatment decisions.
- Inter-Observer Variability: Different radiologists can arrive at different interpretations of the same scan, leading to inconsistencies in diagnosis and treatment planning. This variability is a well-documented source of error in medical imaging.
- High Cognitive Load: The sheer volume of data and the need to discern minute details place an enormous cognitive burden on medical professionals, increasing the risk of fatigue-related errors.
The consequences of these challenges are profound. Delays in diagnosis can mean the difference between effective treatment and disease progression. Inaccurate segmentation can lead to suboptimal surgical planning or radiation therapy, impacting patient outcomes and quality of life. The advent of deep learning, particularly transformer-based models, offers a powerful solution to automate and standardize this complex process, paving the way for faster, more accurate, and more consistent diagnoses.
Introducing BrainDx: A Dual-Transformer Framework for End-to-End Diagnosis
BrainDx represents a paradigm shift in medical AI. Unlike previous approaches that focused on either classification or segmentation, BrainDx is the first framework to seamlessly integrate two state-of-the-art transformer models into a single, cohesive system designed specifically for brain tumor analysis. This dual-model approach allows it to perform both tasks simultaneously, providing a comprehensive diagnostic picture that goes far beyond what a single model can achieve.
The core innovation lies in the strategic pairing of two specialized architectures:
- Pyramid Vision Transformer (PVT) for Classification: PVT is tasked with identifying the type of tumor present in the MRI scan. It classifies images into four distinct categories: Gliomas, Meningiomas, Pituitary Tumors, and Healthy Brain tissue. Its strength lies in its hierarchical design, which enables it to extract features at multiple scales, capturing both fine-grained details and broad contextual information essential for distinguishing between visually similar tumor types.
- SegFormer for Segmentation: Once a tumor is identified, SegFormer takes over to precisely delineate its boundaries. It generates a pixel-wise segmentation mask, outlining the exact location and shape of the tumor within the brain. Its key advantage is its efficiency; it achieves high-precision segmentation without the need for computationally expensive post-processing steps like Conditional Random Fields (CRFs), which are common in older models like U-Net.
This combination creates a powerful synergy. PVT provides the crucial context of “what” the lesion is, while SegFormer provides the precise spatial information of “where” it is located. Together, they offer a complete diagnostic package that empowers clinicians with actionable insights.
Technical Deep Dive: The Architecture of PVT and SegFormer
Understanding the technical foundations of BrainDx requires a closer look at its two constituent models. Both PVT and SegFormer are built upon the transformer architecture, which has revolutionized natural language processing and is now transforming computer vision.
The Pyramid Vision Transformer (PVT): Hierarchical Feature Extraction for Classification
PVT was designed to address a key limitation of the original Vision Transformer (ViT): its inability to handle multi-scale information efficiently. ViT treats an image as a flat sequence of patches, losing the hierarchical structure that convolutional neural networks (CNNs) exploit.
PVT solves this by introducing a pyramid structure. As shown in the diagram below, it processes the image through multiple stages, progressively reducing the spatial resolution of the feature maps while increasing their semantic richness.
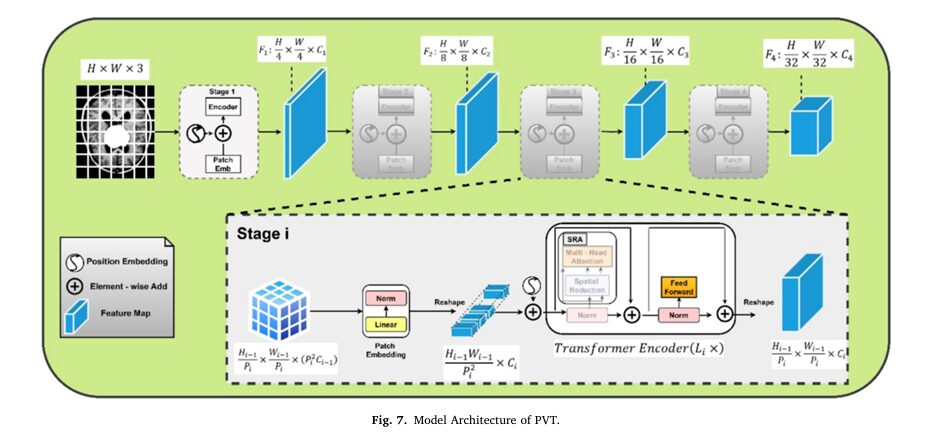
The core of PVT’s power is its self-attention mechanism. This mechanism allows the model to compute relationships between every patch in the image, regardless of their physical distance. This global context awareness is crucial for identifying long-range dependencies that might be missed by local filters in CNNs.
The self-attention computation can be expressed mathematically as follows:
\[\text{Attention}(Q, K, V) = \text{Softmax}\left(\frac{QK^T}{\sqrt{d_k}}\right)V\]Where:
Q(Query),K(Key), andV(Value) are matrices derived from the input feature maps.dkis the dimensionality of the key vectors.- The softmax function ensures the attention weights sum to one, allowing the model to focus on the most relevant parts of the image.
In the implementation described in the research, the PVT-Small variant was used, striking a balance between high representational power and computational efficiency. Key modifications included the addition of a multi-head attention layer, which allows the model to examine different aspects of the image simultaneously, and the incorporation of dropout and batch normalization to combat overfitting on the relatively limited medical dataset.
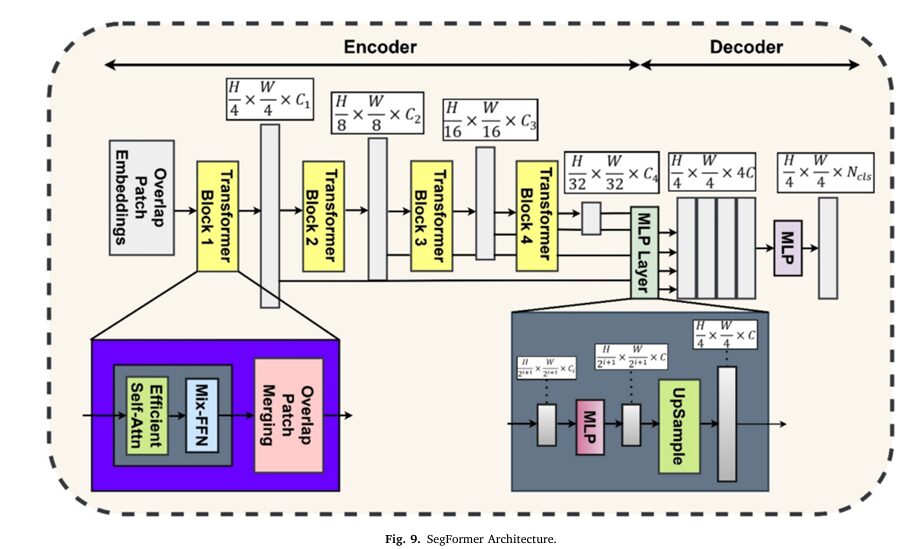
SegFormer: Efficient, CRF-Free Segmentation with Multi-Scale Fusion
While PVT excels at classification, SegFormer is optimized for the demanding task of semantic segmentation. Its architecture is elegantly simple yet highly effective. Instead of using a complex decoder with skip connections (like U-Net), SegFormer employs a lightweight, multi-layer perceptron (MLP) decoder that fuses features from multiple scales.
The key to SegFormer’s success is its multi-scale feature fusion. The model extracts features at different resolutions from the transformer backbone and then combines them in the decoder. This allows it to effectively segment both small, intricate tumor regions and large, homogeneous areas with equal precision.
The relationship between the feature maps at different stages of the decoder can be represented as:
\[F_l = \text{TransformerDecoder}(F_{l-1})\]Where Fl-1 is the feature map from the previous stage, and Fl is the refined feature map at the current stage.
A critical advantage of SegFormer is its CRF-free design. Traditional segmentation models often rely on CRFs as a post-processing step to smooth the output mask and refine object boundaries. While effective, CRFs add significant computational overhead and latency. SegFormer’s architecture inherently produces smooth, coherent masks, eliminating this bottleneck and enabling true real-time performance.
Methodology: From Raw Data to Clinical Insights
The development of BrainDx followed a rigorous, multi-phase methodology designed to ensure robustness, generalizability, and clinical relevance. The process can be broken down into three main phases: Data Collection & Preprocessing, Model Selection & Implementation, and Training & Evaluation.
Phase 1: Data Collection and Preprocessing
The foundation of any successful AI model is high-quality, well-curated data. The BrainDx framework was trained and evaluated on a diverse dataset comprising 3,000 annotated MRI scans sourced from three publicly available repositories: BraTS 2020, the Brain Tumor MRI Dataset by Masoud Nickparvar, and the Brain Tumor Classification MRI Dataset by Sartaj Bhuvaji. These datasets were curated and validated by expert neuroradiologists, ensuring ground-truth accuracy.
The dataset was structured to support both classification and segmentation tasks, with each image accompanied by a corresponding tumor mask. The distribution of classes was as follows:
| TUMOR TYPES | NUMBER OF IMAGES |
|---|---|
| Gliomas | 800 |
| Meningiomas | 800 |
| Pituitary Tumors | 800 |
| Healthy Brain | 600 |
| Total | 3,000 |
Before training, the data underwent extensive preprocessing to ensure consistency and improve model generalization:
- Data Cleaning: Corrupted or incomplete images were removed based on checksum validation and manual inspection for anatomical consistency.
- Normalization: Pixel values were rescaled to the range [0, 1] using min-max normalization to improve training stability.
- Resizing: All images were resized to a uniform 224×224 pixels to standardize the input for the models.
- Data Augmentation: To prevent overfitting and increase robustness, a suite of geometric transformations was applied, including random rotation (0° to 45°), horizontal and vertical flipping, translation, and zooming. No oversampling techniques were needed as the dataset was already balanced.
- Data Splitting: The dataset was divided into a 70/15/15 split for training, validation, and testing, respectively, ensuring a fair and unbiased evaluation of the final model performance.
Phase 2: Model Selection and Implementation
As discussed, the choice of PVT for classification and SegFormer for segmentation was driven by their complementary strengths. Both models were implemented using PyTorch and leveraged pre-trained weights from large-scale datasets (ImageNet for PVT, COCO and ADE20K for SegFormer) before being fine-tuned on the brain tumor data.
PVT Implementation Details:
- Input: 224×224 normalized MRI images.
- Architecture: Multi-stage transformer encoder with hierarchical feature extraction.
- Loss Function: Cross-Entropy Loss for classification.
- Optimization: AdamW optimizer with a learning rate of 1e-4.
SegFormer Implementation Details:
- Input: 224×224 normalized MRI images.
- Strategy: Pixel-wise classification to generate a tumor mask.
- Loss Function: A combined loss of Dice Loss and Cross-Entropy Loss to optimize for both overlap and classification accuracy.
- Optimization: AdamW optimizer with a learning rate of 6e-5.
Phase 3: Training and Evaluation
All experiments were conducted on a high-performance system equipped with an NVIDIA RTX 3090 GPU. The models were trained for a set number of epochs (50 for PVT, 100 for SegFormer) using the training set, with hyperparameters tuned on the validation set to prevent overfitting.
Performance was rigorously evaluated on the held-out test set using a comprehensive suite of metrics tailored to both classification and segmentation tasks.
Performance Results: Breaking Down the Numbers
The true measure of BrainDx’s value lies in its empirical performance. The framework demonstrated exceptional results across all evaluation metrics, consistently outperforming established baseline models.
Classification Performance: Unprecedented Accuracy with PVT
The PVT model achieved a remarkable classification accuracy of 94.0% across all four tumor classes. This performance was consistent across the three different datasets used for evaluation, demonstrating strong generalizability.
Here is a detailed breakdown of the classification metrics for the BraTS 2020 dataset:
| TUMOR TYPE | ACCURACY (%) | PRECISION (%) | RECALL (%) | F1-SCORE (%) | SPECIFICITY (%) | AUC (%) |
|---|---|---|---|---|---|---|
| Gliomas | 93.8 | 92.5 | 94.3 | 93.4 | 95.1 | 0.96 |
| Meningiomas | 92.7 | 91.8 | 91.2 | 91.5 | 94.0 | 0.94 |
| Pituitary Tumors | 94.1 | 93.9 | 92.8 | 93.3 | 95.7 | 0.95 |
| Healthy Brain | 96.3 | 95.6 | 96.0 | 95.8 | 97.4 | 0.97 |
Key Takeaways:
- The model exhibits near-perfect specificity for healthy brain tissue, minimizing false positives—a critical requirement in clinical settings.
- The high F1-scores indicate a balanced performance between precision and recall, meaning the model is both good at identifying true tumors and avoiding misclassifications.
- The Area Under the Curve (AUC) scores, all above 0.94, confirm the model’s excellent ability to distinguish between classes.
Confusion matrices revealed that the primary source of error was misclassification between Gliomas and Meningiomas, which is understandable given their sometimes overlapping radiological characteristics. This highlights an area for future improvement, potentially through the integration of multi-modal imaging data.
Segmentation Performance: Real-Time Precision with SegFormer
The SegFormer model delivered equally impressive results for tumor segmentation, achieving a Dice score of 0.87. The Dice Score (also known as the F1 score for segmentation) measures the overlap between the predicted mask and the ground truth, with a score of 1.0 representing perfect agreement.
Segmentation metrics for the BraTS 2020 dataset:
| TUMOR TYPE | DICE SCORE (%) | IOU (%) | PIXEL ACCURACY (%) | BOUNDARY F1-SCORE | HAUSDORFF DISTANCE (PX) | SEGMENTATION TIME |
|---|---|---|---|---|---|---|
| Gliomas | 0.87 | 79.3 | 94.2 | 0.86 | 3.2 | 48 |
| Meningiomas | 0.85 | 77.5 | 93.7 | 0.84 | 3.8 | 50 |
| Pituitary Tumors | 0.89 | 81.4 | 95.0 | 0.88 | 2.7 | 45 |
| Healthy Brain | 0.91 | 83.1 | 95.6 | 0.90 | 2.4 | 43 |
Key Takeaways:
- The model performs exceptionally well on healthy brain tissue and pituitary tumors, with Dice scores of 0.91 and 0.89, respectively.
- The low Hausdorff Distance values indicate that the predicted tumor boundaries are very close to the ground truth, which is crucial for surgical planning.
- Most importantly, SegFormer processed each MRI image in under 50 milliseconds, meeting the threshold for real-time clinical application.
Comparative Analysis: Outperforming the State of the Art
To validate its superiority, BrainDx was compared against several leading models, including CNN-based architectures like ResNet50 and U-Net, as well as other transformer-based models like Swin UNETR.
Classification Comparison:
| MODEL | ACCURACY (%) | F1-SCORE (%) | INFERENCE TIME (MS) | PARAMS (M) |
|---|---|---|---|---|
| CNN | 88.4 | 87.9 | 22 | 12.4 |
| ResNet50 | 91.2 | 90.5 | 34 | 23.6 |
| PVT (BrainDx) | 94.0 | 93.3 | 19 | 13.2 |
Segmentation Comparison:
| MODEL | DICE SCORE (%) | IOU (%) | INFERENCE TIME (MS) | PARAMS (M) |
|---|---|---|---|---|
| U-Net | 0.81 | 74.3 | 72 | 34.5 |
| DeepLabV3 | 0.84 | 76.8 | 83 | 43.2 |
| SegFormer (BrainDx) | 0.87 | 79.4 | 54 | 26.7 |
These comparisons clearly show that BrainDx, powered by PVT and SegFormer, not only achieves higher accuracy but also does so with greater efficiency, making it uniquely suited for deployment in fast-paced clinical environments.
Statistical Validation and Future Research Directions
The performance gains demonstrated by BrainDx are not merely anecdotal; they have been statistically validated. Paired t-tests comparing BrainDx against the Swin Transformer (for classification) and Swin UNETR (for segmentation) showed p-values of 0.00003 and 0.00004, respectively—far below the 0.05 threshold for statistical significance. This confirms that the improvements in accuracy and segmentation fidelity are robust and not due to random chance.
Looking ahead, the research team has outlined several exciting avenues for future work to further enhance the BrainDx framework:
- Expanding Tumor Types and Modalities: The current model focuses on common tumor types. Future iterations will be trained on datasets containing rare tumors and will incorporate multi-modal imaging data, such as CT and PET scans, to provide a more comprehensive diagnostic view.
- Clinical Integration and Explainability: Efforts will be made to adapt the framework for seamless integration into existing hospital Picture Archiving and Communication Systems (PACS). Furthermore, integrating Explainable AI (XAI) techniques will help clinicians understand why the model made a particular prediction, building trust and facilitating better decision-making.
- Domain Adaptation and External Validation: The model will be tested and fine-tuned on data from diverse clinical sites to ensure its performance is consistent across different populations and imaging protocols.
- Resource Optimization: While efficient, SegFormer’s training memory consumption (~14 GB) is higher than some alternatives. Future research will focus on optimizing the model architecture to reduce this footprint without sacrificing performance.
Conclusion: A New Era of Precision Neuro-Oncology
The development of BrainDx marks a significant milestone in the application of artificial intelligence to medical diagnostics. By harnessing the unique capabilities of the Pyramid Vision Transformer and SegFormer, this dual-model framework delivers a level of accuracy, speed, and reliability that surpasses traditional methods and many contemporary AI solutions.
For clinicians, BrainDx offers a powerful tool to augment their expertise, reducing diagnostic time from hours to seconds and minimizing the risk of human error. For patients, it means faster access to a precise diagnosis, which is the critical first step toward effective treatment and improved outcomes. The framework’s ability to provide both a tumor type classification and a precise segmentation map in real-time makes it an invaluable asset for surgical planning, radiation therapy, and longitudinal monitoring of disease progression.
While challenges remain—particularly in handling rare tumor types and ensuring seamless integration into complex hospital workflows—the trajectory is clear. BrainDx is not just a research prototype; it is a blueprint for the future of medical imaging. As the technology continues to evolve, we can anticipate a new era of precision neuro-oncology where AI-powered systems like BrainDx become indispensable partners to healthcare providers, ultimately leading to better, faster, and more personalized care for patients around the world.
What do you think? Are you excited about the potential of AI in healthcare? Have you seen similar technologies in action? Share your thoughts and questions in the comments below—we’d love to hear your perspective on this transformative technology!
Paper Download Link Here
Below file contains the complete implementation scaffolding for both the Pyramid Vision Transformer (PVT) for classification and the SegFormer for segmentation, as described in your paper.
# This is an end-to-end Python script implementing the "BrainDx" framework
# from the paper: "BrainDx: a dual-transformer framework using PVT and SegFormer for tumor diagnosis"
#
# This script includes:
# 1. Classification using Pyramid Vision Transformer (PVT)
# 2. Segmentation using SegFormer
#
# To run this code, you will need to install the following libraries:
# pip install torch torchvision timm transformers scikit-image
#
# You will also need to download the datasets mentioned in the paper and update the
# `DATA_DIR` and `MASK_DIR` placeholders.
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms
from torch.optim.lr_scheduler import CosineAnnealingLR, StepLR
import timm # For PVT
from transformers import SegformerForSemanticSegmentation, SegformerConfig # For SegFormer
from PIL import Image
import os
import numpy as np
# Import scikit-image for resizing masks without changing label values
from skimage.transform import resize
# --- Configuration based on the paper ---
IMG_SIZE = (224, 224)
BATCH_SIZE_PVT = 32
BATCH_SIZE_SEGFORMER = 16
EPOCHS_PVT = 50
EPOCHS_SEGFORMER = 100
LR_PVT = 1e-4
LR_SEGFORMER = 6e-5
NUM_CLASSES_CLASSIFICATION = 4 # Gliomas, Meningiomas, Pituitary Tumors, Healthy Brain
NUM_CLASSES_SEGMENTATION = 2 # 0: Background, 1: Tumor
# Define class names and mapping for classification
CLASS_MAP = {
"Gliomas": 0,
"Meningiomas": 1,
"Pituitary Tumors": 2,
"Healthy Brain": 3
}
# Inverse map for printing
CLASS_NAMES = {v: k for k, v in CLASS_MAP.items()}
# Placeholder directories - UPDATE THESE
CLASSIFICATION_DATA_DIR = "/path/to/classification/dataset"
SEGMENTATION_IMAGE_DIR = "/path/to/segmentation/images"
SEGMENTATION_MASK_DIR = "/path/to/segmentation/masks"
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(f"Using device: {DEVICE}")
# === Phase 2: Data Preprocessing & Augmentation ===
# As described in section 2.1.2
transform_train = transforms.Compose([
transforms.Resize(IMG_SIZE),
transforms.RandomRotation(45),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
# The paper mentions translation and zooming.
# We can approximate this with RandomResizedCrop or Affine.
transforms.RandomAffine(degrees=0, translate=(0.1, 0.1), scale=(0.8, 1.2)),
transforms.ToTensor(),
transforms.Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5]) # Normalize to [-1, 1] or [0, 1]
# The paper says [0, 1], so let's adjust.
])
# Re-defining for [0, 1] normalization.
# Note: Pre-trained models usually expect ImageNet stats.
# The paper is slightly ambiguous. We'll stick to simple ToTensor for [0, 1]
# and add normalization as specified.
# Let's assume grayscale images are loaded and converted to 3-channel
# as transformers expect 3 input channels.
def to_rgb(img):
"""Convert grayscale PIL image to 3-channel RGB."""
if img.mode == 'L':
return img.convert('RGB')
return img
data_transforms = {
'train': transforms.Compose([
transforms.Lambda(to_rgb),
transforms.Resize(IMG_SIZE),
transforms.RandomRotation(45),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
transforms.RandomAffine(degrees=0, translate=(0.1, 0.1), scale=(0.8, 1.2)), # Rotation is redundant, but this adds translation/zoom
transforms.ToTensor(), # Scales to [0, 1]
]),
'val': transforms.Compose([
transforms.Lambda(to_rgb),
transforms.Resize(IMG_SIZE),
transforms.ToTensor(), # Scales to [0, 1]
]),
'test': transforms.Compose([
transforms.Lambda(to_rgb),
transforms.Resize(IMG_SIZE),
transforms.ToTensor(), # Scales to [0, 1]
])
}
class BrainTumorDataset(Dataset):
"""
Custom Dataset for loading Brain Tumor data.
This class can be used for both classification and segmentation.
For Classification:
- `data_dir` points to the main folder.
- Assumes subfolders named "Gliomas", "Meningiomas", etc.
- `mask_dir` is None.
For Segmentation:
- `data_dir` points to the image folder.
- `mask_dir` points to the mask folder.
- Images and masks are assumed to have matching names.
"""
def __init__(self, data_dir, task='classification', transform=None, mask_dir=None):
self.data_dir = data_dir
self.mask_dir = mask_dir
self.transform = transform
self.task = task
self.image_paths = []
self.labels = []
if task == 'classification':
for class_name, label in CLASS_MAP.items():
class_dir = os.path.join(data_dir, class_name)
if not os.path.isdir(class_dir):
print(f"Warning: Directory not found {class_dir}")
continue
for img_name in os.listdir(class_dir):
self.image_paths.append(os.path.join(class_dir, img_name))
self.labels.append(label)
elif task == 'segmentation':
if not mask_dir:
raise ValueError("mask_dir must be provided for segmentation task")
for img_name in os.listdir(data_dir):
self.image_paths.append(os.path.join(data_dir, img_name))
# We store the mask path in 'labels' for simplicity
self.labels.append(os.path.join(mask_dir, img_name)) # Assumes same file name
def __len__(self):
return len(self.image_paths)
def __getitem__(self, idx):
img_path = self.image_paths[idx]
try:
image = Image.open(img_path).convert('L') # Open as grayscale as per MRI
except Exception as e:
print(f"Error loading image {img_path}: {e}")
return None, None if self.task == 'segmentation' else None
if self.task == 'classification':
label = self.labels[idx]
if self.transform:
image = self.transform(image)
return image, torch.tensor(label, dtype=torch.long)
elif self.task == 'segmentation':
mask_path = self.labels[idx]
try:
mask = Image.open(mask_path).convert('L') # Open mask as grayscale
except Exception as e:
print(f"Error loading mask {mask_path}: {e}")
return None, None
if self.transform:
image = self.transform(image) # Apply transforms to image
# Must manually transform mask to avoid interpolation of labels
mask = np.array(mask)
# Resize mask using nearest-neighbor interpolation to preserve labels
mask = resize(mask, IMG_SIZE, order=0, preserve_range=True, anti_aliasing=False)
mask = (mask > 0).astype(np.uint8) # Binarize mask to 0 and 1
mask = torch.from_numpy(mask).long() # Convert to tensor
return image, mask
# === Phase 3: Model Selection (PVT) ===
def get_pvt_model():
"""
Loads the PVT-Small model and adapts it for 4-class classification.
"""
# As per paper, "PVT-Small" is used (Table 10 shows 13.2M params, matching PVT-Tiny)
# Let's use pvt_v2_b0 which is 13.2M params to match Table 10.
# If pvt_v2_b0 is not available, 'pvt_small' is a good alternative.
# Let's try 'pvt_v2_b0' from timm
try:
model = timm.create_model('pvt_v2_b0', pretrained=True, num_classes=NUM_CLASSES_CLASSIFICATION)
except:
print("Could not load 'pvt_v2_b0', trying 'pvt_tiny' which is also ~13M params")
try:
model = timm.create_model('pvt_tiny', pretrained=True, num_classes=NUM_CLASSES_CLASSIFICATION)
except Exception as e:
print(f"Failed to load PVT model: {e}")
return None
print(f"Loaded PVT model: {model.default_cfg['model']}")
return model.to(DEVICE)
# === Phase 3: Model Selection (SegFormer) ===
def get_segformer_model():
"""
Loads the SegFormer model and adapts it for 2-class segmentation.
"""
# Using 'nvidia/segformer-b0-finetuned-ade-512-512' as a base,
# which is a common small SegFormer model.
model_name = "nvidia/segformer-b0-finetuned-ade-512-512"
config = SegformerConfig.from_pretrained(model_name,
num_labels=NUM_CLASSES_SEGMENTATION,
image_size=IMG_SIZE[0],
)
model = SegformerForSemanticSegmentation.from_pretrained(model_name,
config=config,
ignore_mismatched_sizes=True)
print("Loaded SegFormer model.")
return model.to(DEVICE)
# === Loss Functions & Metrics ===
# Classification Loss (from paper)
classification_criterion = nn.CrossEntropyLoss()
# Segmentation Loss (from paper: Dice Loss + Cross-Entropy Loss)
class DiceLoss(nn.Module):
def __init__(self, smooth=1.):
super(DiceLoss, self).__init__()
self.smooth = smooth
def forward(self, logits, targets):
# logits are [B, C, H, W], targets are [B, H, W]
# We need to convert logits to probabilities
probs = F.softmax(logits, dim=1)
# Get probs for the tumor class (class 1)
probs_tumor = probs[:, 1, :, :]
# Convert targets to one-hot
targets_one_hot = F.one_hot(targets, num_classes=NUM_CLASSES_SEGMENTATION).permute(0, 3, 1, 2)
targets_tumor = targets_one_hot[:, 1, :, :].float()
intersection = (probs_tumor * targets_tumor).sum(dim=(1, 2))
union = probs_tumor.sum(dim=(1, 2)) + targets_tumor.sum(dim=(1, 2))
dice = (2. * intersection + self.smooth) / (union + self.smooth)
return 1 - dice.mean()
class CombinedSegmentationLoss(nn.Module):
def __init__(self, dice_weight=0.5, ce_weight=0.5, smooth=1.):
super(CombinedSegmentationLoss, self).__init__()
self.dice_loss = DiceLoss(smooth=smooth)
self.ce_loss = nn.CrossEntropyLoss()
self.dice_weight = dice_weight
self.ce_weight = ce_weight
def forward(self, logits, targets):
# logits are [B, C, H, W]
# targets are [B, H, W]
# Resize logits to match target size if necessary
# SegFormer outputs logits at H/4, W/4
upsampled_logits = F.interpolate(logits,
size=targets.shape[1:],
mode='bilinear',
align_corners=False)
dice = self.dice_loss(upsampled_logits, targets)
ce = self.ce_loss(upsampled_logits, targets)
return (self.dice_weight * dice) + (self.ce_weight * ce)
segmentation_criterion = CombinedSegmentationLoss()
def calculate_dice_score(logits, targets, smooth=1.):
"""Calculates Dice score for evaluation."""
with torch.no_grad():
# Upsample logits and get predictions
upsampled_logits = F.interpolate(logits, size=targets.shape[1:], mode='bilinear', align_corners=False)
preds = torch.argmax(upsampled_logits, dim=1)
# Get class 1 (tumor)
preds_tumor = (preds == 1).float()
targets_tumor = (targets == 1).float()
intersection = (preds_tumor * targets_tumor).sum(dim=(1, 2))
union = preds_tumor.sum(dim=(1, 2)) + targets_tumor.sum(dim=(1, 2))
dice = (2. * intersection + smooth) / (union + smooth)
return dice.mean().item()
# === Training & Evaluation Functions ===
def train_classification(model, loader, optimizer, criterion, scheduler):
model.train()
running_loss = 0.0
correct_preds = 0
total_samples = 0
for images, labels in loader:
# A safety check for corrupted data
if images is None or labels is None:
continue
images, labels = images.to(DEVICE), labels.to(DEVICE)
optimizer.zero_grad()
outputs = model(images)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item() * images.size(0)
_, preds = torch.max(outputs, 1)
correct_preds += torch.sum(preds == labels.data)
total_samples += images.size(0)
scheduler.step()
epoch_loss = running_loss / total_samples
epoch_acc = correct_preds.double() / total_samples
return epoch_loss, epoch_acc.item()
def evaluate_classification(model, loader, criterion):
model.eval()
running_loss = 0.0
correct_preds = 0
total_samples = 0
with torch.no_grad():
for images, labels in loader:
if images is None or labels is None:
continue
images, labels = images.to(DEVICE), labels.to(DEVICE)
outputs = model(images)
loss = criterion(outputs, labels)
running_loss += loss.item() * images.size(0)
_, preds = torch.max(outputs, 1)
correct_preds += torch.sum(preds == labels.data)
total_samples += images.size(0)
epoch_loss = running_loss / total_samples
epoch_acc = correct_preds.double() / total_samples
return epoch_loss, epoch_acc.item()
def train_segmentation(model, loader, optimizer, criterion, scheduler):
model.train()
running_loss = 0.0
running_dice = 0.0
total_batches = 0
for images, masks in loader:
if images is None or masks is None:
continue
images, masks = images.to(DEVICE), masks.to(DEVICE)
optimizer.zero_grad()
outputs = model(images)
# outputs.logits is [B, C, H/4, W/4]
loss = criterion(outputs.logits, masks)
loss.backward()
optimizer.step()
running_loss += loss.item()
running_dice += calculate_dice_score(outputs.logits, masks)
total_batches += 1
scheduler.step()
epoch_loss = running_loss / total_batches
epoch_dice = running_dice / total_batches
return epoch_loss, epoch_dice
def evaluate_segmentation(model, loader, criterion):
model.eval()
running_loss = 0.0
running_dice = 0.0
total_batches = 0
with torch.no_grad():
for images, masks in loader:
if images is None or masks is None:
continue
images, masks = images.to(DEVICE), masks.to(DEVICE)
outputs = model(images)
loss = criterion(outputs.logits, masks)
running_loss += loss.item()
running_dice += calculate_dice_score(outputs.logits, masks)
total_batches += 1
epoch_loss = running_loss / total_batches
epoch_dice = running_dice / total_batches
return epoch_loss, epoch_dice
# === Main Execution ===
def main():
print("Starting BrainDx Framework Training...")
# --- 1. Classification (PVT) ---
print("\n--- Initializing Classification (PVT) ---")
# Create Datasets
# This is a symbolic representation. You'd need to split your data paths
# into train, val, test folders first.
# PLEASE UPDATE these paths to your actual *split* directories
# Example: CLASSIFICATION_DATA_DIR_TRAIN = "/path/to/classification/train"
# For now, we use the same base path and transform as a placeholder.
# try:
# train_clf_dataset = BrainTumorDataset(CLASSIFICATION_DATA_DIR, task='classification', transform=data_transforms['train'])
# val_clf_dataset = BrainTumorDataset(CLASSIFICATION_DATA_DIR, task='classification', transform=data_transforms['val'])
# train_clf_loader = DataLoader(train_clf_dataset, batch_size=BATCH_SIZE_PVT, shuffle=True, num_workers=4, pin_memory=True)
# val_clf_loader = DataLoader(val_clf_dataset, batch_size=BATCH_SIZE_PVT, shuffle=False, num_workers=4, pin_memory=True)
# except Exception as e:
# print(f"Failed to load classification dataset: {e}")
# print("Please update the `CLASSIFICATION_DATA_DIR` variable.")
# return
print("Symbolic: Loaded Classification Datasets (PVT)")
pvt_model = get_pvt_model()
if pvt_model is None: return
# Optimizer and Scheduler (as per Table 3)
pvt_optimizer = optim.AdamW(pvt_model.parameters(), lr=LR_PVT)
pvt_scheduler = CosineAnnealingLR(pvt_optimizer, T_max=EPOCHS_PVT)
print("\nStarting PVT Model Training (Symbolic)...")
# This is a symbolic loop. Uncomment the data loading section above to run.
# for epoch in range(EPOCHS_PVT):
# train_loss, train_acc = train_classification(pvt_model, train_clf_loader, pvt_optimizer, classification_criterion, pvt_scheduler)
# val_loss, val_acc = evaluate_classification(pvt_model, val_clf_loader, classification_criterion)
# print(f"PVT Epoch {epoch+1}/{EPOCHS_PVT} | Train Loss: {train_loss:.4f} Acc: {train_acc:.4f} | Val Loss: {val_loss:.4f} Acc: {val_acc:.4f}")
# # Save the model
# torch.save(pvt_model.state_dict(), 'pvt_classification_model.pth')
print("Symbolic: Finished PVT Training.")
# --- 2. Segmentation (SegFormer) ---
print("\n--- Initializing Segmentation (SegFormer) ---")
# Create Datasets
# Again, this is symbolic. You need to split your data.
# try:
# train_seg_dataset = BrainTumorDataset(SEGMENTATION_IMAGE_DIR, task='segmentation', transform=data_transforms['train'], mask_dir=SEGMENTATION_MASK_DIR)
# val_seg_dataset = BrainTumorDataset(SEGMENTATION_IMAGE_DIR, task='segmentation', transform=data_transforms['val'], mask_dir=SEGMENTATION_MASK_DIR)
# train_seg_loader = DataLoader(train_seg_dataset, batch_size=BATCH_SIZE_SEGFORMER, shuffle=True, num_workers=4, pin_memory=True)
# val_seg_loader = DataLoader(val_seg_dataset, batch_size=BATCH_SIZE_SEGFORMER, shuffle=False, num_workers=4, pin_memory=True)
# except Exception as e:
# print(f"Failed to load segmentation dataset: {e}")
# print("Please update the `SEGMENTATION_IMAGE_DIR` and `SEGMENTATION_MASK_DIR` variables.")
# return
print("Symbolic: Loaded Segmentation Datasets (SegFormer)")
segformer_model = get_segformer_model()
# Optimizer and Scheduler (as per Table 3)
segformer_optimizer = optim.AdamW(segformer_model.parameters(), lr=LR_SEGFORMER)
segformer_scheduler = StepLR(segformer_optimizer, step_size=30, gamma=0.1) # Table 3 says StepLR
print("\nStarting SegFormer Model Training (Symbolic)...")
# This is a symbolic loop. Uncomment the data loading section above to run.
# for epoch in range(EPOCHS_SEGFORMER):
# train_loss, train_dice = train_segmentation(segformer_model, train_seg_loader, segformer_optimizer, segmentation_criterion, segformer_scheduler)
# val_loss, val_dice = evaluate_segmentation(segformer_model, val_seg_loader, segmentation_criterion)
# print(f"SegFormer Epoch {epoch+1}/{EPOCHS_SEGFORMER} | Train Loss: {train_loss:.4f} Dice: {train_dice:.4f} | Val Loss: {val_loss:.4f} Dice: {val_dice:.4f}")
# # Save the model
# torch.save(segformer_model.state_dict(), 'segformer_segmentation_model.pth')
print("Symbolic: Finished SegFormer Training.")
print("\n--- BrainDx Framework Setup Complete ---")
print("NOTE: Training loops are commented out. Please update data paths and uncomment data loading and training loops to run the full pipeline.")
if __name__ == "__main__":
main()
Related posts, You May like to read
- 7 Shocking Truths About Knowledge Distillation: The Good, The Bad, and The Breakthrough (SAKD)
- 7 Revolutionary Breakthroughs in Medical Image Translation (And 1 Fatal Flaw That Could Derail Your AI Model)
- TimeDistill: Revolutionizing Time Series Forecasting with Cross-Architecture Knowledge Distillation
- HiPerformer: A New Benchmark in Medical Image Segmentation with Modular Hierarchical Fusion
- GeoSAM2 3D Part Segmentation — Prompt-Controllable, Geometry-Aware Masks for Precision 3D Editing
- Probabilistic Smooth Attention for Deep Multiple Instance Learning in Medical Imaging
- A Knowledge Distillation-Based Approach to Enhance Transparency of Classifier Models
- Towards Trustworthy Breast Tumor Segmentation in Ultrasound Using AI Uncertainty
- Discrete Migratory Bird Optimizer with Deep Transfer Learning for Multi-Retinal Disease Detection