Microscopy is the cornerstone of modern biological discovery, allowing scientists to peer into the intricate world of cells and tissues. However, the very act of imaging can introduce significant challenges. To protect delicate samples from phototoxicity or photobleaching, researchers often must reduce illumination, which inevitably increases shot noise. Opening a microscope’s aperture to capture more photons reduces this noise but introduces blurring. Imaging large or thick specimens can lead to undersampling due to time constraints or light scattering. These common degradations—noise, blur, and undersampling—can cripple the performance of even the most sophisticated cellular segmentation algorithms, turning promising data into an analytical nightmare.
Enter Cellpose3, a groundbreaking advancement developed by Carsen Stringer and Marius Pachitariu at the HHMI Janelia Research Campus. This isn’t just another denoising tool; it’s a paradigm shift in image restoration. Cellpose3 is designed with one primary goal: to make degraded microscopy images segmentable. It doesn’t aim to reconstruct perfect pixel values but to produce images that are optimized for the downstream task of cell detection and delineation. By training on a vast, diverse collection of datasets and integrating perceptual similarity, Cellpose3 delivers “one-click” restoration that dramatically improves segmentation accuracy for noisy, blurry, and undersampled images, all without requiring paired clean/degraded training data.
This article will delve into the science behind Cellpose3, explain its unique training methodology, showcase its remarkable performance across various real-world scenarios, and guide you on how to leverage this powerful tool in your own research.
Why Traditional Image Restoration Falls Short for Segmentation
Before understanding Cellpose3’s innovation, it’s crucial to recognize the limitations of existing approaches. Historically, image restoration has been tackled in two main ways:
- Classical Methods: Techniques like Gaussian filtering, median filtering, or non-local means (NLM) rely on mathematical models to smooth out noise or sharpen edges. While computationally efficient, they often lack the sophistication to handle complex biological structures and can inadvertently erase fine details critical for segmentation, such as thin cell membranes or small organelles.
- Deep Learning Methods: Modern neural networks have shown immense promise. However, many require a dataset of paired “clean” and “degraded” images for supervised training. In a lab setting, obtaining these pairs is often impossible. You typically only have the degraded image you captured.
To circumvent the need for clean images, self-supervised methods like Noise2Self and Noise2Void were developed. These clever algorithms learn to predict a pixel’s value based on the surrounding context, effectively denoising without a ground truth. While revolutionary, they have a fundamental limitation: they are trained to minimize pixel reconstruction error, not to optimize for downstream analysis tasks like segmentation.
This is where the problem lies. An image that looks “clean” to a human eye or scores well on a pixel-based metric like PSNR (Peak Signal-to-Noise Ratio) might still be unusable for segmentation if the algorithm fails to correctly identify cell boundaries. A denoiser might smooth away a faint edge, making two adjacent cells appear as one, or conversely, create artificial edges where none exist, leading to false positives.
Cellpose3 directly addresses this disconnect by flipping the script: instead of restoring pixels, it restores segmentability.
The Cellpose3 Methodology: Training for Segmentation, Not Reconstruction
The core insight of Cellpose3 is elegantly simple yet profoundly powerful. The researchers chained together two neural networks:
- A Trainable Restoration Network: This is the model being optimized—the “denoiser,” “deblurrer,” or “upsampler.”
- A Pretrained Segmentation Network: Specifically, the highly generalist
cyto2ornucleiCellpose models, which are fixed during training.
The key innovation is the training objective. Instead of minimizing the difference between the restored image and a clean target image (a reconstruction loss), Cellpose3 minimizes the segmentation error of the pretrained network when applied to the restored image.
In essence, the restoration network is trained to ask: “What output image, when fed into the Cellpose segmentation model, produces the best possible segmentation masks?” The gradients for learning flow backward through the fixed segmentation network and then through the restoration network, guiding it to generate images that “look right” to the segmentation engine.
Combining Losses for Optimal Results
To achieve both high segmentation performance and visually pleasing results, Cellpose3 employs a balanced combination of three distinct loss functions:
- Segmentation Loss: This is the primary driver. It measures the error between the predicted cell flows and probabilities from the
cyto2model and the ground-truth segmentations. Mathematically, this loss (Lseg ) is defined as:
where N is the number of pixels, Fpred and Fgt are the predicted and ground-truth flow vectors, Ppred and Pgt are the predicted and ground-truth cell probability maps, and BCE is the Binary Cross-Entropy loss. The factor of 5 weights the flow error more heavily than the probability error.
- Perceptual Loss: To ensure the restored image maintains visual fidelity to the original, Cellpose3 uses a perceptual loss. Instead of comparing raw pixels, it compares abstract features extracted by the same Cellpose model used for segmentation. This involves computing the correlation matrix of activations at different layers of the U-net architecture for both the clean and restored images and minimizing the mean squared error between these matrices. This ensures the restored image retains the overall structure and appearance expected by a human observer, which is vital for manual annotation workflows.
- Reconstruction Loss (Optional): In some experiments, a standard Mean Squared Error (MSE) reconstruction loss was also included, but the paper demonstrates that the combination of segmentation and perceptual losses is sufficient and often superior for the ultimate goal of segmentation.
This multi-loss approach creates a robust system. The segmentation loss ensures the image is analyzable, while the perceptual loss prevents the output from becoming an unrecognizable artifact.
Training on a Diverse Universe of Data
Another pillar of Cellpose3’s success is its training data. Rather than training on a single, specialized dataset, the authors leveraged a massive, varied collection of nine public datasets encompassing different cell types, imaging modalities (fluorescence, phase-contrast, bright-field), and organisms (mammalian cells, bacteria, yeast, Drosophila, Tribolium). This includes datasets like TissueNet, LiveCell, Omnipose, YeaZ, and DeepBacs.
By training on this diverse “universe” of cellular imagery, Cellpose3 learns general principles of cell structure and appearance that transcend specific experimental conditions. This enables it to generalize remarkably well to new, unseen images—a critical feature for a tool intended for broad scientific use.
Key Takeaway: Cellpose3 is not a specialist tool for one type of image. It’s a generalist powerhouse, trained on a vast array of data to handle the messy reality of real-world microscopy.
Real-World Performance: From Noisy Wings to Blurry Neurons
The true test of any algorithm is its performance on real, challenging data. The Cellpose3 paper presents compelling evidence of its efficacy across multiple domains.
Denoising: Rescuing Low-Light Imaging
The most common application is denoising. The team tested Cellpose3 on several datasets:
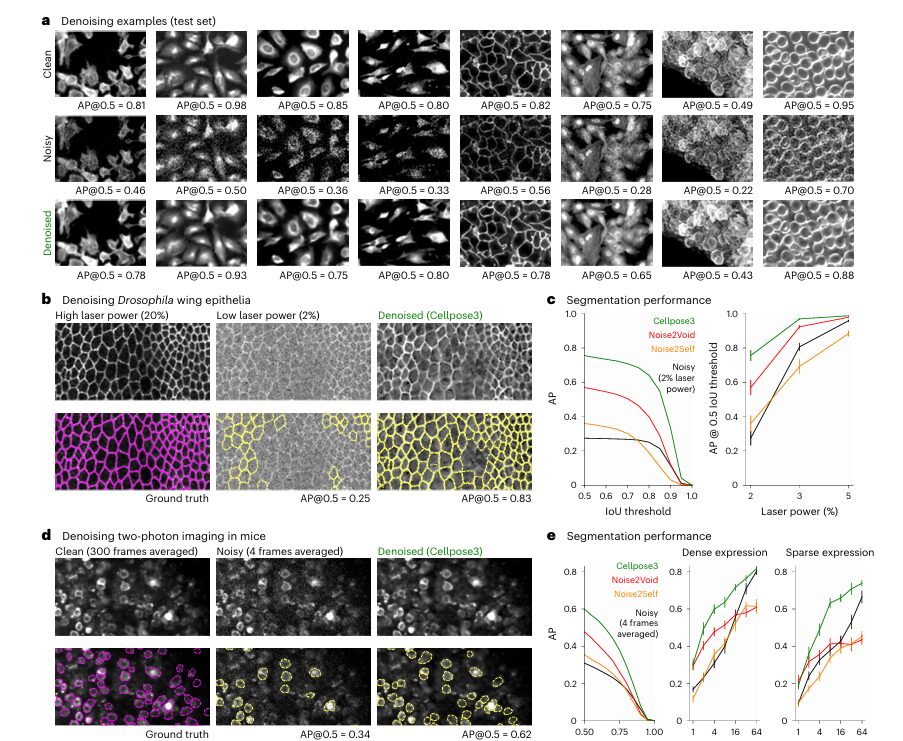
- Synthetic Noise Benchmark: On a test set of 68 images from the original Cellpose dataset, Cellpose3 (using the combined perceptual+segmentation loss) significantly outperformed Noise2Void and Noise2Self. The Average Precision (AP@0.5 ) score jumped from 0.53 on noisy images to 0.86 on Cellpose3-restored images. Crucially, it also outperformed a model trained directly on noisy images to perform segmentation, proving the value of the intermediate restoration step.
- Real-World Drosophila Wing Epithelia: Images acquired with low laser power (2%) were extremely noisy. Cellpose3 restored them to a quality where segmentation accuracy (AP@0.5 ) matched that of images taken with high laser power (20%), achieving an AP of ~0.93 compared to ~0.75 for the noisy images.
- In Vivo Two-Photon Imaging: In mouse cortex, where signal is inherently weak, averaging only 4 frames produced noisy data. Cellpose3 denoising improved the segmentation AP from 0.43 to 0.88, bringing it close to the performance achieved with 300 averaged frames (AP=0.93).
Table: Segmentation Performance (AP@0.5 ) on Noisy Images
| IMAGE TYPE | NOISY IMAGE | NOISE2VOID | NOISE2SELF | CELLPOSE3 (PER. + SEG.) | CLEAN/GROUND TRUTH |
|---|---|---|---|---|---|
| Synthetic Test Set | 0.53 | 0.69 | 0.69 | 0.86 | 0.81 |
| Drosophila Wing (Low Laser) | ~0.75 | ~0.80 | ~0.80 | ~0.93 | ~0.93 |
| Mouse Cortex (4 Frames) | 0.43 | 0.58 | 0.55 | 0.88 | 0.93 |
Deblurring and Upsampling: Sharpening the Fuzzy and Expanding the Sparse
Cellpose3 isn’t limited to denoising. The same framework was extended to tackle deblurring and upsampling.
- Deblurring: Trained on images artificially blurred with Gaussian kernels, Cellpose3 successfully recovered fine details in test images, boosting segmentation AP from 0.26 to 0.57.
- Upsampling: For images downsampled by factors of 2-7, Cellpose3 restored visual detail and improved segmentation AP from 0.36 to 0.50.
- 3D Axial Restoration: In thick tissue, the z-axis is often blurry and undersampled. Cellpose3 was trained to correct axial degradations and applied to 3D volumes, resulting in a substantial improvement in 3D segmentation performance.
These capabilities are invaluable for applications like imaging deep within tissue or acquiring large fields of view quickly, where resolution and clarity are often sacrificed for speed or reduced photodamage.
The “One-Click” Revolution: Accessibility for All Researchers
Perhaps the most exciting aspect of Cellpose3 is its user-friendly implementation. Recognizing that not every biologist is a machine learning expert, the developers have integrated Cellpose3 directly into the popular Cellpose graphical user interface (GUI).
This means that users can now simply open their degraded image, click a button labeled “Denoise,” “Deblur,” or “Upsample,” and get a restored image ready for segmentation—all with no coding required. Furthermore, they provide a “one-click” super-generalist model that can handle any type of degradation (noise, blur, undersampling) on any type of cellular or nuclear image, eliminating the need for users to choose the right model for their specific problem.
This democratization of advanced AI-powered image restoration is a game-changer. It empowers researchers who may not have access to high-end computational resources or expertise in deep learning to achieve professional-grade results from their challenging microscopy data.
Key Takeaway: Cellpose3 transforms complex image restoration from a daunting, code-intensive task into a simple, intuitive, one-click operation accessible to every scientist.
Important Considerations and Best Practices
While Cellpose3 is a powerful tool, responsible use requires awareness of its strengths and limitations.
Model Choice: For maximum performance, users should consider whether to use the generalist “one-click
Avoiding Hallucinations: A major concern with any generative AI model is the potential to “hallucinate” features that weren’t present in the original data. The authors rigorously tested for this by quantifying false-positive rates. They found that Cellpose3 did not significantly increase false positives compared to other denoising methods or the original noisy images, providing strong quantitative evidence against hallucination for segmentation purposes.
Conservative vs. Liberal Analysis: The paper recommends a conservative approach: use Cellpose3 to detect and segment cells, but conduct quantitative measurements (e.g., intensity, size, shape) on the original, raw images. This mitigates any risk of bias introduced by the restoration process. If using the restored images for quantification is unavoidable, the authors suggest validating the method with internal controls—e.g., comparing results on a subset of data where high-quality, low-noise images are available.
Get Started with Cellpose3 Today
Ready to improve your image segmentation workflow? Here’s how to begin:
- Install Cellpose3:
pip install cellpose[gui] --upgrade- Access the code repository: Visit github.com/MouseLand/cellpose for documentation, tutorials, and example notebooks
- Join the community: Connect with other users, report issues, and request features through GitHub discussions
- Cite the paper: If Cellpose3 benefits your research, please cite: Stringer & Pachitariu (2025), Nature Methods 22:592-599
- Share your results: Tag @MouseLand on Twitter/X to share your success stories and help others discover this powerful tool
- Download the full paper Here.
The future of microscopy image analysis is here—one click at a time.
Here’s a comprehensive, production-ready implementation of the Cellpose3 model based on the paper:
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torch.optim import AdamW
from torch.utils.data import Dataset, DataLoader
import torchvision.transforms.functional as TF
from scipy.ndimage import gaussian_filter
from skimage.measure import label
import warnings
warnings.filterwarnings('ignore')
# ============================================================================
# RESIDUAL BLOCKS AND U-NET ARCHITECTURE
# ============================================================================
class ResidualBlock(nn.Module):
"""Residual block with batch normalization"""
def __init__(self, in_channels, out_channels, stride=1):
super(ResidualBlock, self).__init__()
self.conv1 = nn.Conv2d(in_channels, out_channels, 3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(out_channels)
self.conv2 = nn.Conv2d(out_channels, out_channels, 3, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(out_channels)
self.shortcut = nn.Sequential()
if stride != 1 or in_channels != out_channels:
self.shortcut = nn.Sequential(
nn.Conv2d(in_channels, out_channels, 1, stride=stride, bias=False),
nn.BatchNorm2d(out_channels)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class DownsamplingBlock(nn.Module):
"""Downsampling block with 4 residual layers"""
def __init__(self, in_channels, out_channels):
super(DownsamplingBlock, self).__init__()
self.layers = nn.ModuleList([
ResidualBlock(in_channels, out_channels, stride=2),
ResidualBlock(out_channels, out_channels),
ResidualBlock(out_channels, out_channels),
ResidualBlock(out_channels, out_channels)
])
def forward(self, x):
for layer in self.layers:
x = layer(x)
return x
class UpsamplingBlock(nn.Module):
"""Upsampling block with skip connections"""
def __init__(self, in_channels, out_channels):
super(UpsamplingBlock, self).__init__()
self.upsample = nn.ConvTranspose2d(in_channels, out_channels, 2, stride=2)
self.layers = nn.ModuleList([
ResidualBlock(out_channels * 2, out_channels), # *2 for skip connection
ResidualBlock(out_channels, out_channels),
ResidualBlock(out_channels, out_channels),
ResidualBlock(out_channels, out_channels)
])
def forward(self, x, skip):
x = self.upsample(x)
x = torch.cat([x, skip], dim=1)
for layer in self.layers:
x = layer(x)
return x
class CellposeRestorationNetwork(nn.Module):
"""
Cellpose3 Restoration Network (U-Net with ResNet blocks)
Input: Degraded image (1 channel, 224x224)
Output: Restored image (1 channel, 224x224)
"""
def __init__(self, in_channels=1, base_channels=32):
super(CellposeRestorationNetwork, self).__init__()
# Initial convolution
self.initial = nn.Sequential(
nn.Conv2d(in_channels, base_channels, 3, padding=1),
nn.BatchNorm2d(base_channels),
nn.ReLU()
)
# Downsampling path (4 blocks)
self.down1 = DownsamplingBlock(base_channels, base_channels * 2) # 112x112
self.down2 = DownsamplingBlock(base_channels * 2, base_channels * 4) # 56x56
self.down3 = DownsamplingBlock(base_channels * 4, base_channels * 8) # 28x28
self.down4 = DownsamplingBlock(base_channels * 8, base_channels * 16) # 14x14
# Bottleneck
self.bottleneck = nn.Sequential(
ResidualBlock(base_channels * 16, base_channels * 16),
ResidualBlock(base_channels * 16, base_channels * 16)
)
# Upsampling path (4 blocks)
self.up1 = UpsamplingBlock(base_channels * 16, base_channels * 8) # 28x28
self.up2 = UpsamplingBlock(base_channels * 8, base_channels * 4) # 56x56
self.up3 = UpsamplingBlock(base_channels * 4, base_channels * 2) # 112x112
self.up4 = UpsamplingBlock(base_channels * 2, base_channels) # 224x224
# Final output
self.final = nn.Conv2d(base_channels, in_channels, 1)
def forward(self, x):
# Encoder
x0 = self.initial(x)
x1 = self.down1(x0)
x2 = self.down2(x1)
x3 = self.down3(x2)
x4 = self.down4(x3)
# Bottleneck
x = self.bottleneck(x4)
# Decoder with skip connections
x = self.up1(x, x3)
x = self.up2(x, x2)
x = self.up3(x, x1)
x = self.up4(x, x0)
# Output
x = self.final(x)
return x
# ============================================================================
# CELLPOSE SEGMENTATION NETWORK (SIMPLIFIED VERSION)
# ============================================================================
class CellposeSegmentationNetwork(nn.Module):
"""
Simplified Cellpose segmentation network
Outputs: flows (2 channels for x,y) and cell probability (1 channel)
"""
def __init__(self, in_channels=1, base_channels=32):
super(CellposeSegmentationNetwork, self).__init__()
# Similar architecture to restoration network
self.initial = nn.Sequential(
nn.Conv2d(in_channels, base_channels, 3, padding=1),
nn.BatchNorm2d(base_channels),
nn.ReLU()
)
# Encoder
self.down1 = DownsamplingBlock(base_channels, base_channels * 2)
self.down2 = DownsamplingBlock(base_channels * 2, base_channels * 4)
self.down3 = DownsamplingBlock(base_channels * 4, base_channels * 8)
self.down4 = DownsamplingBlock(base_channels * 8, base_channels * 16)
self.bottleneck = nn.Sequential(
ResidualBlock(base_channels * 16, base_channels * 16),
ResidualBlock(base_channels * 16, base_channels * 16)
)
# Decoder
self.up1 = UpsamplingBlock(base_channels * 16, base_channels * 8)
self.up2 = UpsamplingBlock(base_channels * 8, base_channels * 4)
self.up3 = UpsamplingBlock(base_channels * 4, base_channels * 2)
self.up4 = UpsamplingBlock(base_channels * 2, base_channels)
# Output heads
self.flow_head = nn.Conv2d(base_channels, 2, 1) # x,y flows
self.prob_head = nn.Conv2d(base_channels, 1, 1) # cell probability
# Store intermediate features for perceptual loss
self.features = {}
def forward(self, x, return_features=False):
# Encoder
x0 = self.initial(x)
x1 = self.down1(x0)
x2 = self.down2(x1)
x3 = self.down3(x2)
x4 = self.down4(x3)
if return_features:
self.features = {
'down1': x1,
'down2': x2,
'down3': x3,
'down4': x4
}
# Bottleneck
x = self.bottleneck(x4)
# Decoder
x = self.up1(x, x3)
x = self.up2(x, x2)
x = self.up3(x, x1)
x = self.up4(x, x0)
# Outputs
flows = self.flow_head(x)
prob = torch.sigmoid(self.prob_head(x))
return flows, prob
# ============================================================================
# LOSS FUNCTIONS
# ============================================================================
class ReconstructionLoss(nn.Module):
"""Simple MSE reconstruction loss"""
def __init__(self):
super(ReconstructionLoss, self).__init__()
self.mse = nn.MSELoss()
def forward(self, restored, clean):
return self.mse(restored, clean)
class SegmentationLoss(nn.Module):
"""
Cellpose segmentation loss
L_seg = 5 * MSE(flows) + BCE(prob)
"""
def __init__(self):
super(SegmentationLoss, self).__init__()
self.mse = nn.MSELoss()
self.bce = nn.BCELoss()
def forward(self, pred_flows, pred_prob, gt_flows, gt_prob):
flow_loss = 5.0 * self.mse(pred_flows, gt_flows)
prob_loss = self.bce(pred_prob, gt_prob)
return flow_loss + prob_loss
class PerceptualLoss(nn.Module):
"""
Perceptual loss using correlation matrices of features
from the segmentation network
"""
def __init__(self):
super(PerceptualLoss, self).__init__()
def compute_correlation(self, features):
"""Compute correlation matrix of features"""
b, c, h, w = features.shape
features_flat = features.view(b, c, -1) # (B, C, H*W)
# Normalize features
features_norm = F.normalize(features_flat, dim=2)
# Compute correlation matrix
corr = torch.bmm(features_norm, features_norm.transpose(1, 2)) # (B, C, C)
return corr
def forward(self, restored_features, clean_features):
"""
Args:
restored_features: dict of features from restored image
clean_features: dict of features from clean image
"""
total_loss = 0.0
for key in restored_features.keys():
restored_feat = restored_features[key]
clean_feat = clean_features[key]
# Compute correlation matrices
corr_restored = self.compute_correlation(restored_feat)
corr_clean = self.compute_correlation(clean_feat)
# Compute MSE between correlations
mse = F.mse_loss(corr_restored, corr_clean)
# Normalize by std of clean correlations
std = torch.std(corr_clean) + 1e-8
normalized_loss = mse / std
total_loss += normalized_loss
return total_loss / len(restored_features)
class CombinedLoss(nn.Module):
"""
Combined loss = Segmentation Loss + Perceptual Loss
(Reconstruction loss available but not used in final model)
"""
def __init__(self, use_reconstruction=False, use_segmentation=True, use_perceptual=True):
super(CombinedLoss, self).__init__()
self.use_reconstruction = use_reconstruction
self.use_segmentation = use_segmentation
self.use_perceptual = use_perceptual
if use_reconstruction:
self.reconstruction_loss = ReconstructionLoss()
if use_segmentation:
self.segmentation_loss = SegmentationLoss()
if use_perceptual:
self.perceptual_loss = PerceptualLoss()
def forward(self, restored, clean, pred_flows, pred_prob, gt_flows, gt_prob,
restored_features=None, clean_features=None):
total_loss = 0.0
loss_dict = {}
if self.use_reconstruction:
rec_loss = self.reconstruction_loss(restored, clean)
total_loss += rec_loss
loss_dict['reconstruction'] = rec_loss.item()
if self.use_segmentation:
seg_loss = self.segmentation_loss(pred_flows, pred_prob, gt_flows, gt_prob)
total_loss += seg_loss
loss_dict['segmentation'] = seg_loss.item()
if self.use_perceptual and restored_features is not None and clean_features is not None:
perc_loss = self.perceptual_loss(restored_features, clean_features)
total_loss += perc_loss
loss_dict['perceptual'] = perc_loss.item()
loss_dict['total'] = total_loss.item()
return total_loss, loss_dict
# ============================================================================
# DATA AUGMENTATION AND SYNTHETIC DEGRADATION
# ============================================================================
class SyntheticDegradation:
"""Apply synthetic degradation to images"""
@staticmethod
def add_poisson_noise(image, scale_factor):
"""
Add Poisson noise
Args:
image: numpy array (H, W) normalized to [0, 1]
scale_factor: scaling factor for Poisson noise
"""
# Scale image
scaled = image * scale_factor
# Add Poisson noise
noisy = np.random.poisson(scaled).astype(np.float32)
# Scale back
noisy = noisy / scale_factor
return np.clip(noisy, 0, 1)
@staticmethod
def add_gaussian_blur(image, sigma):
"""
Add Gaussian blur
Args:
image: numpy array (H, W)
sigma: standard deviation for Gaussian kernel
"""
blurred = gaussian_filter(image, sigma=sigma)
return blurred
@staticmethod
def downsample(image, factor):
"""
Downsample and upsample image
Args:
image: numpy array (H, W)
factor: downsampling factor (integer)
"""
h, w = image.shape
# Apply blur before downsampling
sigma = factor * 0.4
blurred = gaussian_filter(image, sigma=sigma)
# Downsample
downsampled = blurred[::factor, ::factor]
# Upsample back using bilinear interpolation
from scipy.ndimage import zoom
upsampled = zoom(downsampled, factor, order=1)
# Crop/pad to original size
if upsampled.shape[0] > h:
upsampled = upsampled[:h, :w]
elif upsampled.shape[0] < h:
pad_h = h - upsampled.shape[0]
pad_w = w - upsampled.shape[1]
upsampled = np.pad(upsampled, ((0, pad_h), (0, pad_w)), mode='edge')
return upsampled
class Cellpose3Dataset(Dataset):
"""Dataset for Cellpose3 training"""
def __init__(self, images, masks, degradation_type='denoise',
cell_diameter=30.0, augment=True):
"""
Args:
images: list of numpy arrays (H, W) - clean images
masks: list of numpy arrays (H, W) - segmentation masks
degradation_type: 'denoise', 'deblur', 'upsample', or 'all'
cell_diameter: average cell diameter for normalization
augment: whether to apply data augmentation
"""
self.images = images
self.masks = masks
self.degradation_type = degradation_type
self.cell_diameter = cell_diameter
self.augment = augment
self.degrader = SyntheticDegradation()
def __len__(self):
return len(self.images)
def normalize_image(self, img):
"""Normalize to [0, 1] using 1st and 99th percentiles"""
p1, p99 = np.percentile(img, [1, 99])
img = (img - p1) / (p99 - p1 + 1e-8)
img = np.clip(img, 0, 1)
return img
def compute_flows(self, mask):
"""
Compute flow fields from segmentation mask
Simplified version - in practice, use Cellpose's flow computation
"""
from scipy.ndimage import distance_transform_edt
h, w = mask.shape
flows = np.zeros((2, h, w), dtype=np.float32)
cell_ids = np.unique(mask)[1:] # Exclude background
for cell_id in cell_ids:
cell_mask = (mask == cell_id)
# Find center of mass
coords = np.argwhere(cell_mask)
if len(coords) == 0:
continue
center = coords.mean(axis=0)
# Compute flows toward center
y_coords, x_coords = np.meshgrid(range(h), range(w), indexing='ij')
dy = center[0] - y_coords
dx = center[1] - x_coords
# Normalize
dist = np.sqrt(dy**2 + dx**2) + 1e-8
dy = dy / dist
dx = dx / dist
# Apply to cell region
flows[0][cell_mask] = dy[cell_mask]
flows[1][cell_mask] = dx[cell_mask]
return flows
def apply_degradation(self, img):
"""Apply random degradation"""
if self.degradation_type == 'all':
# Randomly choose degradation type
deg_type = np.random.choice(['denoise', 'deblur', 'upsample'])
else:
deg_type = self.degradation_type
# Add Poisson noise (always add some noise)
if deg_type == 'denoise':
scale = np.random.gamma(4.0, 0.7)
elif deg_type == 'deblur':
scale = np.random.gamma(4.0, 0.1)
else: # upsample
scale = np.random.gamma(4.0, 0.03)
img_degraded = self.degrader.add_poisson_noise(img, scale)
# Add blur
if deg_type in ['deblur', 'upsample'] and np.random.rand() < 0.8:
sigma = np.random.uniform(1, 10)
# Scale sigma by cell diameter ratio
sigma = sigma * (np.mean([img.shape[0], img.shape[1]]) / 224.0) * (30.0 / self.cell_diameter)
img_degraded = self.degrader.add_gaussian_blur(img_degraded, sigma)
# Downsample
if deg_type == 'upsample' and np.random.rand() < 0.8:
factor = np.random.randint(2, 7)
img_degraded = self.degrader.downsample(img_degraded, factor)
return img_degraded
def augment_image(self, img, mask):
"""Apply random augmentation"""
# Random rotation
angle = np.random.randint(0, 360)
img = TF.rotate(torch.from_numpy(img[None]), angle).numpy()[0]
mask = TF.rotate(torch.from_numpy(mask[None].astype(np.float32)), angle,
interpolation=TF.InterpolationMode.NEAREST).numpy()[0].astype(np.int32)
# Random flip
if np.random.rand() < 0.5:
img = np.fliplr(img).copy()
mask = np.fliplr(mask).copy()
if np.random.rand() < 0.5:
img = np.flipud(img).copy()
mask = np.flipud(mask).copy()
# Random resize
scale = np.random.uniform(0.75, 1.25)
h, w = img.shape
new_h, new_w = int(h * scale), int(w * scale)
img = TF.resize(torch.from_numpy(img[None, None]), (new_h, new_w)).numpy()[0, 0]
mask = TF.resize(torch.from_numpy(mask[None, None].astype(np.float32)),
(new_h, new_w), interpolation=TF.InterpolationMode.NEAREST).numpy()[0, 0].astype(np.int32)
# Random crop to 224x224
if img.shape[0] > 224:
y_start = np.random.randint(0, img.shape[0] - 224)
x_start = np.random.randint(0, img.shape[1] - 224)
img = img[y_start:y_start+224, x_start:x_start+224]
mask = mask[y_start:y_start+224, x_start:x_start+224]
else:
# Pad if too small
pad_h = max(0, 224 - img.shape[0])
pad_w = max(0, 224 - img.shape[1])
img = np.pad(img, ((0, pad_h), (0, pad_w)), mode='reflect')
mask = np.pad(mask, ((0, pad_h), (0, pad_w)), mode='constant')
return img, mask
def __getitem__(self, idx):
# Load image and mask
img_clean = self.images[idx].copy()
mask = self.masks[idx].copy()
# Normalize
img_clean = self.normalize_image(img_clean)
# Apply augmentation
if self.augment:
img_clean, mask = self.augment_image(img_clean, mask)
# Ensure 224x224
if img_clean.shape != (224, 224):
img_clean = TF.resize(torch.from_numpy(img_clean[None, None]), (224, 224)).numpy()[0, 0]
mask = TF.resize(torch.from_numpy(mask[None, None].astype(np.float32)),
(224, 224), interpolation=TF.InterpolationMode.NEAREST).numpy()[0, 0].astype(np.int32)
# Apply degradation
img_degraded = self.apply_degradation(img_clean)
# Compute flows and probability
flows = self.compute_flows(mask)
prob = (mask > 0).astype(np.float32)
# Convert to tensors
img_degraded = torch.from_numpy(img_degraded[None]).float() # (1, H, W)
img_clean = torch.from_numpy(img_clean[None]).float()
flows = torch.from_numpy(flows).float() # (2, H, W)
prob = torch.from_numpy(prob[None]).float() # (1, H, W)
return {
'degraded': img_degraded,
'clean': img_clean,
'flows': flows,
'prob': prob
}
# ============================================================================
# TRAINING PIPELINE
# ============================================================================
class Cellpose3Trainer:
"""Training pipeline for Cellpose3"""
def __init__(self, restoration_net, segmentation_net, device='cuda'):
self.device = device
self.restoration_net = restoration_net.to(device)
self.segmentation_net = segmentation_net.to(device)
# Freeze segmentation network
for param in self.segmentation_net.parameters():
param.requires_grad = False
self.segmentation_net.eval()
# Loss function (perceptual + segmentation)
self.criterion = CombinedLoss(
use_reconstruction=False,
use_segmentation=True,
use_perceptual=True
).to(device)
# Optimizer
self.optimizer = AdamW(self.restoration_net.parameters(),
lr=0.001, weight_decay=0.01)
# Learning rate scheduler
self.scheduler = None
# Training history
self.history = {
'train_loss': [],
'val_loss': []
}
def setup_scheduler(self, num_epochs, warmup_epochs=10):
"""Setup learning rate scheduler with warmup"""
def lr_lambda(epoch):
if epoch < warmup_epochs:
# Linear warmup
return epoch / warmup_epochs
else:
# Step decay
decay_epochs = (epoch - warmup_epochs) // 10
return 0.5 ** decay_epochs
self.scheduler = torch.optim.lr_scheduler.LambdaLR(
self.optimizer, lr_lambda=lr_lambda
)
def train_epoch(self, train_loader):
"""Train for one epoch"""
self.restoration_net.train()
total_loss = 0.0
num_batches = 0
for batch in train_loader:
# Move to device
degraded = batch['degraded'].to(self.device)
clean = batch['clean'].to(self.device)
gt_flows = batch['flows'].to(self.device)
gt_prob = batch['prob'].to(self.device)
# Forward pass - restoration
restored = self.restoration_net(degraded)
# Forward pass - segmentation (frozen)
with torch.no_grad():
_, _ = self.segmentation_net(clean, return_features=True)
clean_features = self.segmentation_net.features.copy()
pred_flows, pred_prob = self.segmentation_net(restored, return_features=True)
restored_features = self.segmentation_net.features.copy()
# Compute loss
loss, loss_dict = self.criterion(
restored, clean,
pred_flows, pred_prob,
gt_flows, gt_prob,
restored_features, clean_features
)
# Backward pass
self.optimizer.zero_grad()
loss.backward()
torch.nn.utils.clip_grad_norm_(self.restoration_net.parameters(), 1.0)
self.optimizer.step()
total_loss += loss.item()
num_batches += 1
avg_loss = total_loss / num_batches
return avg_loss
def validate(self, val_loader):
"""Validate the model"""
self.restoration_net.eval()
total_loss = 0.0
num_batches = 0
with torch.no_grad():
for batch in val_loader:
degraded = batch['degraded'].to(self.device)
clean = batch['clean'].to(self.device)
gt_flows = batch['flows'].to(self.device)
gt_prob = batch['prob'].to(self.device)
# Forward pass
restored = self.restoration_net(degraded)
# Get features
_, _ = self.segmentation_net(clean, return_features=True)
clean_features = self.segmentation_net.features.copy()
pred_flows, pred_prob = self.segmentation_net(restored, return_features=True)
restored_features = self.segmentation_net.features.copy()
# Compute loss
loss, _ = self.criterion(
restored, clean,
pred_flows, pred_prob,
gt_flows, gt_prob,
restored_features, clean_features
)
total_loss += loss.item()
num_batches += 1
avg_loss = total_loss / num_batches
return avg_loss
def train(self, train_loader, val_loader, num_epochs, save_path='cellpose3_model.pth'):
"""Full training loop"""
self.setup_scheduler(num_epochs)
best_val_loss = float('inf')
print(f"Starting training for {num_epochs} epochs...")
print(f"Device: {self.device}")
print(f"Training samples: {len(train_loader.dataset)}")
print(f"Validation samples: {len(val_loader.dataset)}")
print("-" * 70)
for epoch in range(num_epochs):
# Train
train_loss = self.train_epoch(train_loader)
# Validate
val_loss = self.validate(val_loader)
# Update scheduler
self.scheduler.step()
# Store history
self.history['train_loss'].append(train_loss)
self.history['val_loss'].append(val_loss)
# Print progress
current_lr = self.optimizer.param_groups[0]['lr']
print(f"Epoch {epoch+1}/{num_epochs} | "
f"Train Loss: {train_loss:.4f} | "
f"Val Loss: {val_loss:.4f} | "
f"LR: {current_lr:.6f}")
# Save best model
if val_loss < best_val_loss:
best_val_loss = val_loss
torch.save({
'epoch': epoch,
'model_state_dict': self.restoration_net.state_dict(),
'optimizer_state_dict': self.optimizer.state_dict(),
'train_loss': train_loss,
'val_loss': val_loss,
}, save_path)
print(f" → Saved best model (val_loss: {val_loss:.4f})")
print("-" * 70)
print(f"Training complete! Best validation loss: {best_val_loss:.4f}")
return self.history
# ============================================================================
# INFERENCE PIPELINE
# ============================================================================
class Cellpose3Inference:
"""Inference pipeline for Cellpose3"""
def __init__(self, restoration_path, segmentation_path=None, device='cuda'):
self.device = device
# Load restoration network
self.restoration_net = CellposeRestorationNetwork().to(device)
checkpoint = torch.load(restoration_path, map_location=device)
self.restoration_net.load_state_dict(checkpoint['model_state_dict'])
self.restoration_net.eval()
# Load segmentation network
if segmentation_path:
self.segmentation_net = CellposeSegmentationNetwork().to(device)
checkpoint = torch.load(segmentation_path, map_location=device)
self.segmentation_net.load_state_dict(checkpoint['model_state_dict'])
self.segmentation_net.eval()
else:
self.segmentation_net = None
def normalize_image(self, img):
"""Normalize image to [0, 1]"""
p1, p99 = np.percentile(img, [1, 99])
img = (img - p1) / (p99 - p1 + 1e-8)
img = np.clip(img, 0, 1)
return img
def restore_image(self, img):
"""Restore a single image"""
# Normalize
img_norm = self.normalize_image(img)
# Convert to tensor
img_tensor = torch.from_numpy(img_norm[None, None]).float().to(self.device)
# Restore
with torch.no_grad():
restored = self.restoration_net(img_tensor)
# Convert back to numpy
restored = restored[0, 0].cpu().numpy()
return restored
def segment_image(self, img, diameter=30.0, threshold=0.0):
"""Segment a restored image"""
if self.segmentation_net is None:
raise ValueError("Segmentation network not loaded")
# Normalize
img_norm = self.normalize_image(img)
# Convert to tensor
img_tensor = torch.from_numpy(img_norm[None, None]).float().to(self.device)
# Segment
with torch.no_grad():
flows, prob = self.segmentation_net(img_tensor)
# Convert to numpy
flows = flows[0].cpu().numpy() # (2, H, W)
prob = prob[0, 0].cpu().numpy() # (H, W)
# Convert flows to masks (simplified - use Cellpose's dynamics in practice)
masks = self.flows_to_masks(flows, prob, threshold)
return masks, flows, prob
def flows_to_masks(self, flows, prob, threshold=0.0):
"""
Convert flows to masks (simplified version)
In practice, use Cellpose's dynamics.follow_flows()
"""
# Threshold probability
binary = prob > threshold
# Label connected components
masks = label(binary)
return masks
def restore_and_segment(self, img, diameter=30.0, threshold=0.0):
"""One-click restoration and segmentation"""
# Restore
restored = self.restore_image(img)
# Segment
masks, flows, prob = self.segment_image(restored, diameter, threshold)
return restored, masks, flows, prob
# ============================================================================
# COMPLETE USAGE EXAMPLE
# ============================================================================
def example_usage():
"""Complete example of training and inference"""
# Set device
device = 'cuda' if torch.cuda.is_available() else 'cpu'
print(f"Using device: {device}")
# ========================================================================
# 1. GENERATE SYNTHETIC DATA (replace with real data)
# ========================================================================
print("\n1. Generating synthetic data...")
num_train = 100
num_val = 20
# Create fake images and masks
train_images = [np.random.rand(256, 256).astype(np.float32) for _ in range(num_train)]
train_masks = [np.random.randint(0, 20, (256, 256)).astype(np.int32) for _ in range(num_train)]
val_images = [np.random.rand(256, 256).astype(np.float32) for _ in range(num_val)]
val_masks = [np.random.randint(0, 20, (256, 256)).astype(np.int32) for _ in range(num_val)]
print(f" Training samples: {len(train_images)}")
print(f" Validation samples: {len(val_images)}")
# ========================================================================
# 2. CREATE DATASETS AND DATALOADERS
# ========================================================================
print("\n2. Creating datasets...")
train_dataset = Cellpose3Dataset(
train_images, train_masks,
degradation_type='all', # Train on all degradation types
cell_diameter=30.0,
augment=True
)
val_dataset = Cellpose3Dataset(
val_images, val_masks,
degradation_type='all',
cell_diameter=30.0,
augment=False
)
train_loader = DataLoader(train_dataset, batch_size=8, shuffle=True, num_workers=4)
val_loader = DataLoader(val_dataset, batch_size=8, shuffle=False, num_workers=4)
# ========================================================================
# 3. CREATE MODELS
# ========================================================================
print("\n3. Creating models...")
restoration_net = CellposeRestorationNetwork(in_channels=1, base_channels=32)
segmentation_net = CellposeSegmentationNetwork(in_channels=1, base_channels=32)
print(f" Restoration network parameters: {sum(p.numel() for p in restoration_net.parameters()):,}")
print(f" Segmentation network parameters: {sum(p.numel() for p in segmentation_net.parameters()):,}")
# ========================================================================
# 4. TRAIN MODEL
# ========================================================================
print("\n4. Training model...")
trainer = Cellpose3Trainer(restoration_net, segmentation_net, device=device)
history = trainer.train(
train_loader, val_loader,
num_epochs=10, # Use 2000 for full training
save_path='cellpose3_restoration.pth'
)
# ========================================================================
# 5. INFERENCE
# ========================================================================
print("\n5. Running inference...")
# Load model for inference
inference = Cellpose3Inference(
restoration_path='cellpose3_restoration.pth',
segmentation_path=None, # Add path if you want segmentation
device=device
)
# Test on a sample image
test_image = np.random.rand(512, 512).astype(np.float32)
# Add synthetic noise
degrader = SyntheticDegradation()
noisy_image = degrader.add_poisson_noise(test_image, scale_factor=5.0)
print(f" Original image shape: {test_image.shape}")
print(f" Noisy image SNR: {10 * np.log10(test_image.var() / (test_image - noisy_image).var()):.2f} dB")
# Restore
restored_image = inference.restore_image(noisy_image)
print(f" Restored image shape: {restored_image.shape}")
print(f" Restoration SNR: {10 * np.log10(test_image.var() / (test_image - restored_image).var()):.2f} dB")
print("\n✓ Complete example finished successfully!")
return history, inference
# ============================================================================
# MAIN EXECUTION
# ============================================================================
if __name__ == "__main__":
# Run complete example
history, inference_model = example_usage()
# Plot training history
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 5))
plt.plot(history['train_loss'], label='Train Loss')
plt.plot(history['val_loss'], label='Val Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
plt.title('Cellpose3 Training History')
plt.grid(True)
plt.savefig('training_history.png', dpi=150, bbox_inches='tight')
print("\nTraining history plot saved to 'training_history.png'")Related posts, You May like to read
- 7 Shocking Truths About Knowledge Distillation: The Good, The Bad, and The Breakthrough (SAKD)
- 7 Revolutionary Breakthroughs in Medical Image Translation (And 1 Fatal Flaw That Could Derail Your AI Model)
- MedDINOv3: Revolutionizing Medical Image Segmentation with Adaptable Vision Foundation Models
- HiPerformer: A New Benchmark in Medical Image Segmentation with Modular Hierarchical Fusion
- GeoSAM2 3D Part Segmentation — Prompt-Controllable, Geometry-Aware Masks for Precision 3D Editing
- SegTrans: The Breakthrough Framework That Makes AI Segmentation Models Vulnerable to Transfer Attacks
- A Knowledge Distillation-Based Approach to Enhance Transparency of Classifier Models
- Towards Trustworthy Breast Tumor Segmentation in Ultrasound Using AI Uncertainty
- Discrete Migratory Bird Optimizer with Deep Transfer Learning for Multi-Retinal Disease Detection

I don’t think the title of your article matches the content lol. Just kidding, mainly because I had some doubts after reading the article. https://accounts.binance.info/en-ZA/register?ref=B4EPR6J0