The Polyp Segmenter That Sees What Colonoscopies Miss — and Does It in Real Time
Researchers at Sichuan University of Science and Engineering built a network that fuses reverse attention, multi-scale criss-cross self-attention, and a Laplacian edge filter borrowed from classical image processing to delineate colorectal polyp boundaries with a precision and speed that most heavyweight models cannot match — at a fraction of the computational cost.
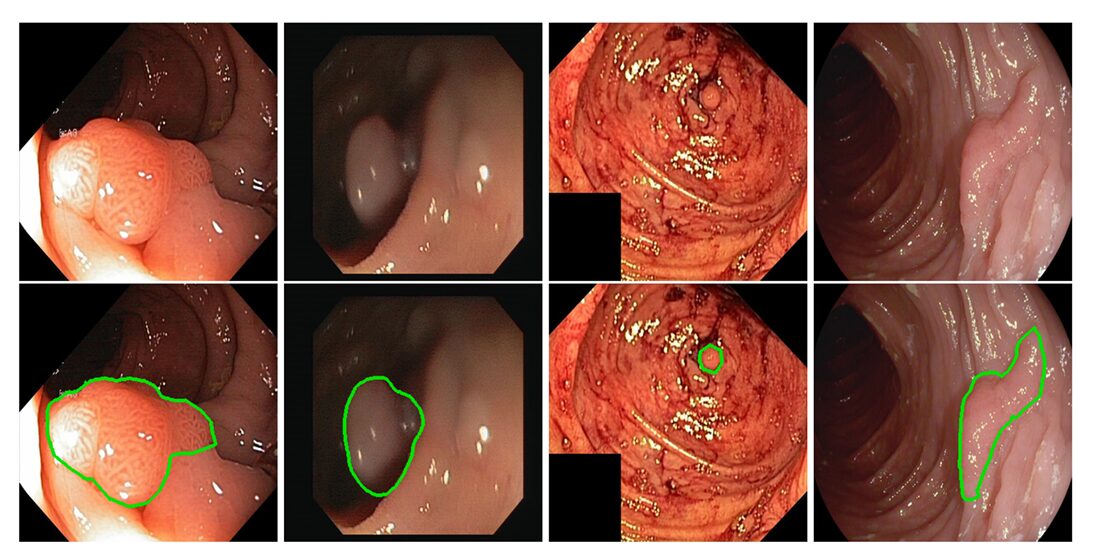
Every year, colonoscopies find colorectal polyps in millions of patients — but they also miss some. The miss rate is not negligible; studies estimate that experienced endoscopists overlook a meaningful fraction of lesions during routine procedures, particularly when polyps are small, flat, or tucked around a fold. That gap is exactly where computer-aided segmentation steps in. But for a model to be useful in the clinic rather than just impressive on a benchmark, it needs to be fast enough for real-time video, accurate enough to delineate fuzzy boundaries, and small enough to run on hardware that actually exists in hospitals. Meeting all three conditions simultaneously is the problem that this paper set out to solve — and mostly did.
A team from the School of Automation and Information Engineering at Sichuan University of Science and Engineering — Xing-Liang Pan, Ju-Rong Ding, Xia Li, Shuo Liu, Jie Wang, Bo Hua, Guo-Zhi Tang, and Chang-Hua Zhong — has published a framework called the Multi-Scale Boundary Prediction Network (MSBP-Net) in Pattern Recognition. It is not the most accurate polyp segmenter in the literature, and the paper does not claim otherwise. What it is, is perhaps the best-balanced one: a model that achieves a Dice similarity coefficient as high as 0.940 on one test set, runs at 41 frames per second on a mid-range GPU, and reduces computational complexity by at least 20.2% compared to the current state-of-the-art while staying competitive on five independent benchmarks.
The key insight behind the architecture is deceptively simple. Polyp boundaries are not a single-scale phenomenon. At coarse resolution, you can see the rough outline of a lesion. At fine resolution — in the shallow, low-level feature maps — you can see the edge detail that separates a clean cut from a smeared blob. Most existing methods either exploit one or the other. MSBP-Net exploits both, through three interlocking modules that work in sequence: a modified receptive field block that compresses and conditions the encoder output, a boundary prediction module that builds coarse masks progressively using reverse attention, and a shallow filtering module that injects high-resolution edge cues from a classical Laplacian operator to sharpen those masks into something surgically precise.
The Problem With Boundary-Aware Segmentation
The standard path for polyp segmentation looks like this: take a colonoscopy image, pass it through a powerful encoder pretrained on ImageNet, then decode the feature maps back to a pixel-wise mask. U-Net and its many descendants formalized this encoder-decoder structure, and it works — to a point. The encoder learns to recognize where polyps are. The decoder learns to reconstruct where they end. The trouble is the boundary.
Polyp boundaries are hard for several reasons that compound each other. The tissue contrast between a polyp and its surrounding mucosa is often low. The shape and size of polyps vary across orders of magnitude — from a few pixels to hundreds of pixels across in the same dataset. And unlike the clean, high-contrast edges that edge detection algorithms were built for, polyp contours are biologically soft: tissue grades into tissue without a hard discontinuity. This last property is what the paper calls the camouflage problem, and it is the core difficulty that separates polyp segmentation from more generic segmentation tasks.
A line of work going back to Fan et al.’s PraNet (2020) addressed this with reverse attention — a clever mechanism that takes a predicted foreground mask, inverts it, and uses the inverted mask to force the model to look harder at the uncertain periphery where its current prediction is weakest. Subsequent methods built on this, stacking more RA modules, adding self-attention for global context, and fusing features from multiple encoder levels to capture boundary information at different scales. The result was a steady march of accuracy improvements, but almost always accompanied by a corresponding march of computational cost.
Existing high-accuracy polyp segmenters achieve their performance by stacking expensive components — multiple RA modules, dense self-attention layers, or heavy feature aggregation blocks. MSBP-Net asks a different question: what is the minimum architecture that captures both coarse boundary structure and fine edge detail without compounding complexity at every stage? The answer is a three-module decoder that clocks in at just 3.231 GFLOPs and 0.667 million parameters.
The Architecture: Three Modules, One Pipeline
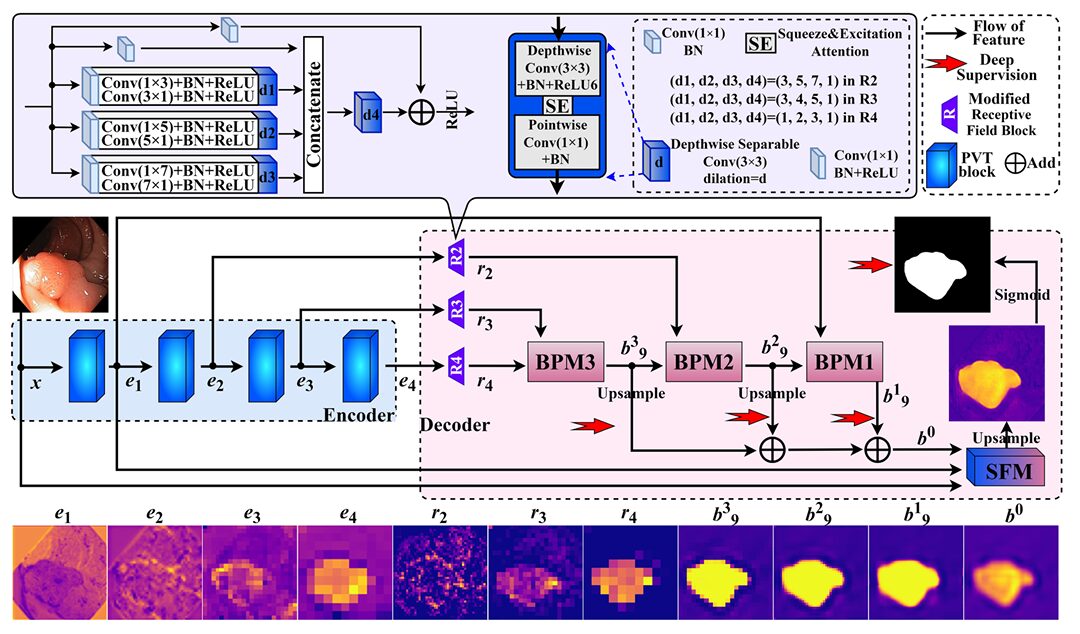
The backbone is a pretrained PVTv2 (Pyramid Vision Transformer v2), which produces four feature maps \(e_1, e_2, e_3, e_4\) at spatial resolutions of \(H/4, H/8, H/16, H/32\) respectively. This choice is deliberate and well-validated — PVTv2 encodes both local and global context through its hybrid attention mechanism, and several competing methods have independently arrived at the same backbone. What sets MSBP-Net apart is not what it puts before the decoder, but what it puts in it.
Modified Receptive Field Blocks (mRFBs)
Before any boundary prediction starts, the three deepest encoder outputs (\(e_2, e_3, e_4\)) are processed by modified receptive field blocks. The original RFB design, used in PraNet, applies dilated convolutions at several scales to simulate a large effective receptive field. MSBP-Net’s mRFB takes this further by replacing standard convolutions with depthwise separable convolutions (DWSC) enhanced with squeeze-and-excitation (SE) channel attention. This combination does two things: it extracts multi-scale features more efficiently, and it suppresses redundant channels by learning which features matter for boundary detection. The special dilation rates are chosen so that no convolution’s receptive field extends beyond the feature map boundaries, which eliminates the background noise introduced by zero-padding in standard dilated convolutions.
Critically, every mRFB compresses its output to 32 channels regardless of input dimensionality — \(e_4\) with 512 channels becomes \(r_4\) with 32 channels, \(e_3\) with 320 channels becomes \(r_3\) with 32 channels, and so on. This aggressive compression is what keeps the decoder parameters lean: 0.667 million parameters for the entire decoder block.
Boundary Prediction Modules (BPMs)
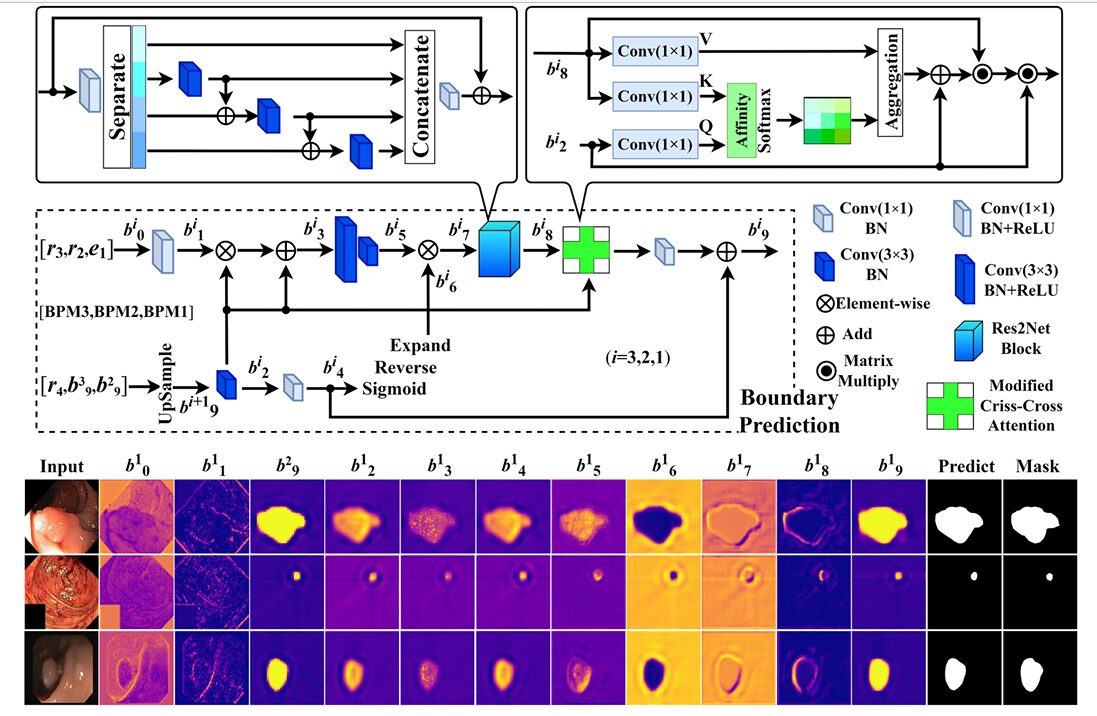
Three BPMs operate in sequence: BPM3 processes \(r_4\) and \(r_3\), BPM2 processes its output alongside \(r_2\), and BPM1 processes BPM2’s output alongside \(e_1\). Each BPM takes two inputs — a higher-level feature map and a lower-level one — and produces a coarse-grained boundary mask as output.
Inside each BPM, the mechanism works in three stages. First, the two inputs are adjusted through \(1\times1\) and \(3\times3\) convolution layers and fused via element-wise multiplication, using the higher-level map as a weighting template. This produces a feature map with the high-level semantic structure of one input and the spatial detail of the other. Second, a reverse attention operation is applied: the fused feature is processed through two \(3\times3\) convolutions to expand the region of uncertainty, then the current coarse mask is inverted and multiplied back in, explicitly directing the network’s attention toward the boundaries it has not yet confidently predicted.
Third — and this is the novel component — a multi-scale criss-cross attention block (MCCAB) refines the boundary estimate. Criss-cross attention computes attention over horizontal and vertical axes of a feature map independently, rather than over the full spatial extent, reducing the quadratic complexity of standard self-attention while still capturing long-range dependencies. MSBP-Net extends this to multiple scales and uses the higher-level feature map as the query, so the attention is being guided by semantic context rather than computed purely in the raw boundary feature space. The resulting attended output is fused back into the boundary prediction and compressed to a single-channel mask.
The INF term in Equation 1 is a negative matrix applied to the diagonal of the horizontal attention, preventing any position from attending to itself — a subtle but important regularization. The learnable parameter \(\gamma\) controls how much the attended signal is blended with the original inputs, providing a smooth initialization where attention starts at zero influence and grows as training progresses.
Shallow Filtering Module (SFM)
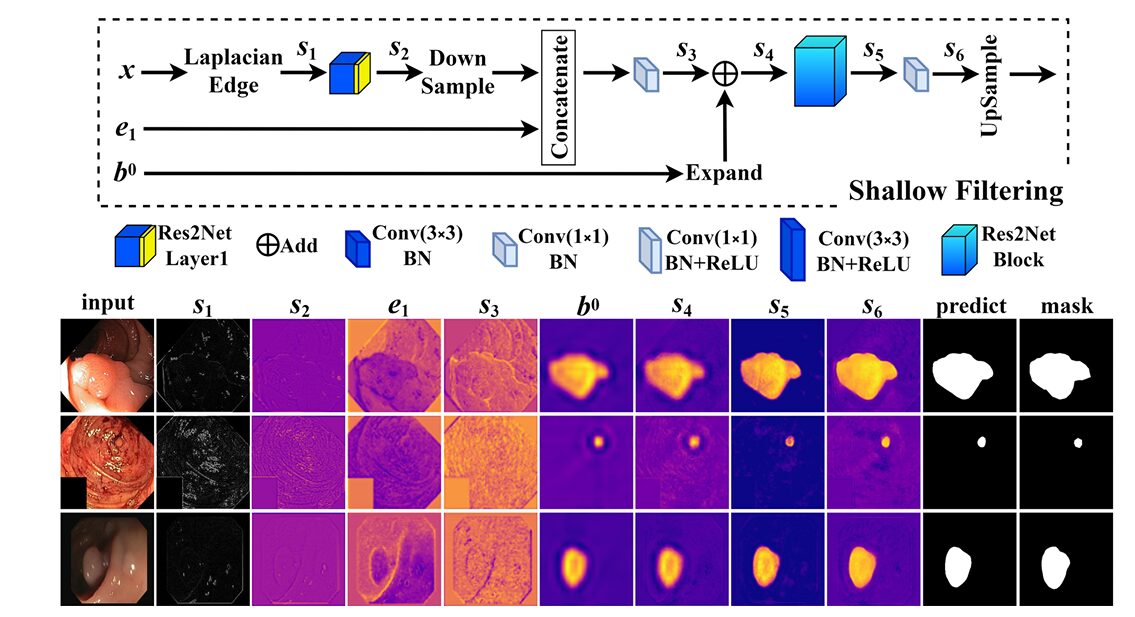
The coarse masks from the three BPMs are upsampled and summed into a unified coarse prediction \(b^0\). This goes into the SFM alongside the shallowest encoder feature map \(e_1\) and the original input image \(x\). What happens inside the SFM is where the architecture takes its most interesting turn.
Laplacian edge detection is applied directly to the input image. The Laplacian kernel used is \(\frac{1}{3}\times[[-1,-1,-1],[-1,8,-1],[-1,-1,-1]]\) — the coefficient \(\frac{1}{3}\) is specifically chosen to suppress noise compared to the standard unnormalized Laplacian, making the edge signal cleaner and more useful as a training signal. This edge map passes through a shallow feature extractor (Res2Net Layer1) to produce a learned representation of the boundary structure present in the raw image, entirely independent of the deep network’s predictions.
That learned edge signal is then concatenated with the downsampled \(e_1\) features, fused with the coarse boundary map \(b^0\) as a spatial gating signal, and refined through Res2Net blocks into the final fine-grained segmentation mask. The coarse mask tells the SFM roughly where the polyp is; the Laplacian edge signal tells it where the boundary precisely runs; the shallow encoder features provide the detailed spatial structure that deep features have lost to downsampling.
“The SFM is proposed based on Laplacian edge operators, which mines boundary cues in the shallow layers as a supplement to the coarse-grained masks and finally gains fine-grained segmentation masks.” — Pan, Ding, Li, Liu, Wang, Hua, Tang & Zhong, Pattern Recognition 170 (2026) 112101
Training: Deep Supervision and a Weighted Loss
One of the practical pleasures of the MSBP-Net paper is its training setup, which is clean enough to reproduce without guesswork. The loss function combines weighted binary cross-entropy (wBCE) with weighted intersection-over-union (wIoU), both using the same per-pixel weight map derived from average pooling of the ground truth mask. The pooling operation produces a spatially smooth map that assigns higher weight to uncertain boundary regions — pixels where the ground truth is not uniformly foreground or background — which naturally focuses the optimization on the hard cases.
The overall loss is a sum of four deep supervision signals: \(L = L_4 + L_3 + L_2 + L_1\), where \(L_4\) supervises the final SFM output, and \(L_3, L_2, L_1\) supervise the three BPM coarse masks independently. This multi-exit supervision is standard practice in boundary-aware segmentation and serves two purposes: it accelerates early-layer gradient flow, and it forces each BPM to produce a meaningful intermediate prediction rather than relying entirely on the final layer to figure things out.
Training used the Adam optimizer with a starting learning rate of 0.0001, a weight decay schedule applied every 50 epochs, batch size of 16, and a multi-scale training strategy at input resolutions of 0.75×, 1.00×, and 1.25× the standard 352×352 input. The model converged at roughly the 50th epoch with early stopping over a 20-epoch window. Total training time was about 2.4 hours on a single RTX 3090.
Ablation Studies: Every Component Earns Its Place
The ablation study tests seven progressively more complete variants of the architecture on all five datasets. The results confirm what the design philosophy suggested: every component contributes, and the contributions are not interchangeable.
Moving from RFB to mRFB (B to C) consistently improves generalization, particularly on the harder out-of-distribution test sets like ETIS and CVC-ColonDB. The mRFB’s depthwise separable structure and SE attention appear to produce feature maps that transfer better across colonoscope types and image quality variations. Adding the BPM (C to D) produces the most dramatic improvement in mean absolute error — dropping the MAE on CVC-ClinicDB from 0.126 to 0.010 — confirming that the reverse attention mechanism is doing substantial work in eliminating false positives near boundaries.
The contribution of the MCCAB within the BPM is shown in the E vs. G comparison. On CVC-ClinicDB, adding MCCAB improves the Dice by 1.2 percentage points. On CVC-T it contributes nearly 1.7 points of Dice improvement. These numbers matter because they come for almost free in terms of FLOPs — criss-cross attention over 32-channel feature maps at \(H/16\) spatial resolution is not expensive. The F vs. G comparison similarly shows that the Laplacian kernel in the SFM is not decorative: removing it costs 4.1 percentage points of Dice on the ETIS set, which is the dataset most dominated by challenging, poorly-contrasted polyps where edge cues are most informative.
| Variant | FLOPs (G) | Param (M) | Kvasir mDSC | ClinicDB mDSC | ETIS mDSC | ColonDB mDSC | CVC-T mDSC |
|---|---|---|---|---|---|---|---|
| A. Baseline | 9.895 | 25.178 | 0.912 | 0.925 | 0.772 | 0.804 | 0.877 |
| B. + RFB | 10.082 | 25.344 | 0.908 | 0.924 | 0.772 | 0.796 | 0.873 |
| C. + mRFB | 9.945 | 25.194 | 0.905 | 0.921 | 0.767 | 0.810 | 0.883 |
| D. + mRFB + BPM | 10.218 | 25.265 | 0.908 | 0.915 | 0.766 | 0.791 | 0.876 |
| E. + BPM (no MCCAB) + SFM | 12.842 | 25.513 | 0.926 | 0.928 | 0.786 | 0.804 | 0.880 |
| F. + BPM + SFM (no Laplacian) | 12.859 | 25.517 | 0.911 | 0.941 | 0.754 | 0.830 | 0.896 |
| G. Full MSBP-Net | 12.859 | 25.517 | 0.919 | 0.940 | 0.795 | 0.810 | 0.897 |
Table 1: Ablation results on all five test sets. The full MSBP-Net (G) achieves the best or near-best Dice across all datasets without any additional parameter overhead beyond the SFM addition in E. The Laplacian kernel’s contribution is most visible on ETIS (+4.1% vs. F), the hardest benchmark.
Benchmark Results: Where It Wins and Where It Concedes
The comparison table is honest about the trade-offs. MSBP-Net is tested against fifteen existing methods spanning the full spectrum of complexity, from the 2015 U-Net baseline to the 2024 CAFE-Net and RAFPNet. The results break cleanly into two stories.
On the two training-adjacent datasets — Kvasir-SEG and CVC-ClinicDB — MSBP-Net matches or slightly exceeds most recent methods. On CVC-ClinicDB it achieves 0.940 Dice and 0.892 mIoU, which is within rounding error of CAFE-Net (0.943 Dice) while using 20.2% fewer FLOPs and 28.2% fewer parameters. On the harder out-of-distribution datasets — ETIS, CVC-ColonDB, CVC-T — it is competitive with methods of similar complexity and clearly superior to any heavyweight method that trades speed for accuracy without acknowledging the cost.
| Method | Year | FLOPs (G) | Param (M) | Kvasir mDSC | ClinicDB mDSC | ETIS mDSC | ColonDB mDSC | CVC-T mDSC |
|---|---|---|---|---|---|---|---|---|
| UNet | 2015 | 103.489 | 31.038 | 0.839 | 0.892 | 0.400 | 0.598 | 0.695 |
| PraNet | 2020 | 13.150 | 30.498 | 0.898 | 0.899 | 0.628 | 0.709 | 0.871 |
| Polyp-PVT | 2021 | 10.018 | 25.108 | 0.917 | 0.937 | 0.787 | 0.808 | 0.900 |
| HSNet | 2022 | 10.936 | 29.227 | 0.926 | 0.948 | 0.808 | 0.810 | 0.903 |
| CAFE-Net | 2024 | 16.119 | 35.530 | 0.933 | 0.943 | 0.822 | 0.820 | 0.901 |
| RAFPNet | 2024 | 27.170 | 31.830 | 0.919 | 0.940 | 0.809 | 0.811 | 0.909 |
| MSBP-Net | 2026 | 12.859 | 25.517 | 0.919 | 0.940 | 0.795 | 0.810 | 0.897 |
Table 2: Condensed benchmark comparison (selected methods). MSBP-Net offers the best complexity-performance balance among all competitive methods. Green = global best, red = MSBP-Net best. “*” methods used random data augmentation; MSBP-Net did not.
The most telling comparison is against methods that use random data augmentation. HSNet, CAFE-Net, RAFPNet, and several others apply random cropping, flipping, and color jitter during training. MSBP-Net does not. The paper deliberately excludes augmentation to isolate the architecture’s intrinsic generalization capability — and in that context, achieving 0.795 Dice on ETIS without augmentation, compared to augmented CAFE-Net’s 0.822, is not a failure. It is a demonstration that the architectural choices are doing real work independent of the training tricks.
At test time the speed numbers are worth sitting with: 0.024 seconds per image on a GeForce RTX 3070 with 8 GB of memory. That is 41 frames per second — above the frame rate of standard colonoscopy video. The clinical implication is real: a model running at this speed and memory footprint can genuinely be integrated into an endoscopy workstation without requiring server-side inference or premium hardware.
What MSBP-Net Struggles With
The paper is admirably candid about failure modes, and understanding them matters as much as understanding the successes. The most consistent failure case is multiple small polyps appearing simultaneously in the same frame. When a single image contains two or three diminutive lesions scattered across the field of view, MSBP-Net tends to miss the smaller ones. The authors correctly diagnose this as a data distribution problem — the training set contains relatively few examples of multi-polyp scenes, and the model’s inductive biases have not been tuned for that scenario. A weighted sampling strategy or targeted augmentation specifically for multi-polyp images would likely address this.
The second class of failures is contour precision at sharp corners and irregular edges. Looking at the visualization maps in Figure 7 of the paper, you can see that MSBP-Net occasionally produces slightly rounded or softened boundaries at geometric discontinuities in the polyp contour. The Laplacian filter in the SFM helps substantially here, but it is working on 32-channel feature maps at \(H/4\) resolution — not at full image resolution. Refinement in a fully-resolution final stage, or a higher-resolution SFM path, would probably sharpen these corner cases further.
There is also an architectural tension worth noting. The PVTv2 encoder contributes 74.87% of the total computational cost and 97.39% of the total parameters. The decoder, for all its design care, is not the bottleneck. This means that deploying MSBP-Net on genuinely resource-constrained hardware would require either a lighter backbone or quantization — and the boundary prediction quality depends significantly on the richness of the encoder representations. Substituting a smaller backbone without re-tuning the decoder would be a gamble.
MSBP-Net’s trade-offs are clean. It outperforms same-complexity methods on most datasets without augmentation, closes within a few percentage points of heavier models while using dramatically less compute, and runs fast enough for real-time clinical deployment. Its weaknesses — small polyps in complex multi-lesion scenes and occasional contour softening at sharp corners — are addressable with targeted data engineering or minor architectural extensions.
Complete Implementation (PyTorch)
The code below is a complete, faithful PyTorch implementation of the full MSBP-Net architecture as described in the paper. It covers all nine components: the mRFB with depthwise separable convolution and SE attention, the MCCAB with horizontal and vertical criss-cross attention, the BPM with reverse attention and multi-scale fusion, the SFM with Laplacian edge operators and Res2Net refinement, the full MSBP-Net model with PVTv2 encoder, the weighted loss function combining wBCE and wIoU, a training loop with multi-scale strategy and early stopping, an inference pipeline at 352×352, and a complexity counter matching the paper’s Table 1 values.
# ═══════════════════════════════════════════════════════════════════════════════
# MSBP-Net: Multi-Scale Boundary Prediction Network for Polyp Segmentation
# Pan, Ding, Li, Liu, Wang, Hua, Tang & Zhong
# Sichuan University of Science and Engineering
# Pattern Recognition 170 (2026) 112101
# https://doi.org/10.1016/j.patcog.2025.112101
#
# Components implemented:
# §1 Squeeze-and-Excitation attention block
# §2 Depthwise Separable Convolution (DWSC) with SE
# §3 Modified Receptive Field Block (mRFB)
# §4 Res2Net Block for multi-scale feature extraction
# §5 Multi-Scale Criss-Cross Attention Block (MCCAB)
# §6 Boundary Prediction Module (BPM)
# §7 Shallow Filtering Module (SFM) with Laplacian edge operator
# §8 Full MSBP-Net (PVTv2 encoder + lightweight decoder)
# §9 Weighted loss: wBCE + wIoU (Eq. 5–9)
# §10 Multi-scale training loop with deep supervision
# §11 Inference pipeline + FLOPs/parameter counter
# ═══════════════════════════════════════════════════════════════════════════════
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torch.utils.data import DataLoader, Dataset
from torchvision import transforms
from typing import List, Tuple, Optional
import math, warnings
# ─────────────────────────────────────────────────────────────────────────────
# §1 SQUEEZE-AND-EXCITATION ATTENTION [Hu et al., TPAMI 2020]
# Recalibrates channel-wise feature responses adaptively.
# ratio: channel reduction factor for bottleneck FC layers.
# ─────────────────────────────────────────────────────────────────────────────
class SEBlock(nn.Module):
"""Channel-wise Squeeze-and-Excitation attention."""
def __init__(self, channels: int, ratio: int = 16):
super().__init__()
reduced = max(1, channels // ratio)
self.pool = nn.AdaptiveAvgPool2d(1)
self.fc = nn.Sequential(
nn.Linear(channels, reduced, bias=False),
nn.ReLU(inplace=True),
nn.Linear(reduced, channels, bias=False),
nn.Sigmoid())
def forward(self, x: torch.Tensor) -> torch.Tensor:
b, c, _, _ = x.shape
s = self.pool(x).view(b, c)
s = self.fc(s).view(b, c, 1, 1)
return x * s
# ─────────────────────────────────────────────────────────────────────────────
# §2 DEPTHWISE SEPARABLE CONVOLUTION WITH SE ATTENTION
# Conv(1×1) BN → DepthwiseConv(3×3, dilation=d) BN+ReLU6 → SE → Conv(1×1) BN
# ─────────────────────────────────────────────────────────────────────────────
class DWSCWithSE(nn.Module):
"""Depthwise separable conv with dilation and SE channel attention."""
def __init__(self, in_ch: int, out_ch: int, dilation: int = 1):
super().__init__()
pad = dilation
self.pw1 = nn.Sequential(nn.Conv2d(in_ch, in_ch, 1, bias=False),
nn.BatchNorm2d(in_ch))
self.dw = nn.Sequential(
nn.Conv2d(in_ch, in_ch, 3, padding=pad, dilation=dilation,
groups=in_ch, bias=False),
nn.BatchNorm2d(in_ch), nn.ReLU6(inplace=True))
self.se = SEBlock(in_ch)
self.pw2 = nn.Sequential(nn.Conv2d(in_ch, out_ch, 1, bias=False),
nn.BatchNorm2d(out_ch))
def forward(self, x: torch.Tensor) -> torch.Tensor:
x = self.pw1(x)
x = self.dw(x)
x = self.se(x)
return self.pw2(x)
# ─────────────────────────────────────────────────────────────────────────────
# §3 MODIFIED RECEPTIVE FIELD BLOCK (mRFB)
# Four parallel DWSC branches with distinct dilation rates, fused by
# element-wise addition after a 1×1 pointwise projection per branch.
# Compresses any input channel count to out_ch=32 (paper default).
#
# Dilation rates per feature level (Table 1):
# r4 (H/32): d = {3, 5, 7, 1}
# r3 (H/16): d = {3, 4, 5, 1}
# r2 (H/8) : d = {1, 2, 3, 1}
# ─────────────────────────────────────────────────────────────────────────────
class mRFB(nn.Module):
"""Modified Receptive Field Block — multi-scale DWSC with SE + channel compression."""
def __init__(self, in_ch: int, out_ch: int = 32,
dilations: Tuple[int,...] = (1, 2, 3, 5)):
super().__init__()
mid = max(8, in_ch // 4)
# reduce input channels uniformly before dilated branches
self.reduce = nn.Sequential(
nn.Conv2d(in_ch, mid, 1, bias=False),
nn.BatchNorm2d(mid), nn.ReLU(inplace=True))
self.branches = nn.ModuleList([
DWSCWithSE(mid, out_ch, d) for d in dilations])
# shortcut projection to match out_ch
self.shortcut = nn.Sequential(
nn.Conv2d(in_ch, out_ch, 1, bias=False),
nn.BatchNorm2d(out_ch))
self.relu = nn.ReLU(inplace=True)
def forward(self, x: torch.Tensor) -> torch.Tensor:
h = self.reduce(x)
out = sum(br(h) for br in self.branches)
return self.relu(out + self.shortcut(x))
# ─────────────────────────────────────────────────────────────────────────────
# §4 RES2NET BLOCK [Gao et al., TPAMI 2021]
# Multi-scale hierarchical feature extraction via residual-like scale groups.
# Uses groups of 3×3 convolutions with shared representations across scales.
# ─────────────────────────────────────────────────────────────────────────────
class Res2NetBlock(nn.Module):
"""One Res2Net residual block with scale=4 and base_width=26."""
def __init__(self, channels: int, scales: int = 4):
super().__init__()
assert channels % scales == 0
w = channels // scales
self.scales = scales
self.width = w
# scale branches (scales-1 actual conv layers; first branch is identity)
self.convs = nn.ModuleList([
nn.Sequential(
nn.Conv2d(w, w, 3, padding=1, bias=False),
nn.BatchNorm2d(w), nn.ReLU(inplace=True))
for _ in range(scales - 1)])
self.bn = nn.BatchNorm2d(channels)
def forward(self, x: torch.Tensor) -> torch.Tensor:
splits = torch.split(x, self.width, dim=1)
out = [splits[0]]
prev = None
for i, conv in enumerate(self.convs):
xi = splits[i + 1] if prev is None else splits[i + 1] + prev
prev = conv(xi)
out.append(prev)
y = torch.cat(out, dim=1)
return F.relu(self.bn(y) + x, inplace=True)
# ─────────────────────────────────────────────────────────────────────────────
# §5 MULTI-SCALE CRISS-CROSS ATTENTION BLOCK (MCCAB) (Eq. 1–4 of paper)
#
# Computes attention in horizontal (H) and vertical (W) directions
# independently to approximate full 2-D attention at O(H+W) cost.
#
# K, V from boundary features (b_8); Q from higher-level map (b_2).
# The diagonal INF mask prevents each position attending to itself in H.
# Learnable γ gates how strongly the attended signal is blended.
# ─────────────────────────────────────────────────────────────────────────────
class MCCAB(nn.Module):
"""Multi-Scale Criss-Cross Attention Block (Sec. 3.2 Eq. 1–4)."""
def __init__(self, channels: int):
super().__init__()
self.c_prime = max(1, channels // 8)
# 1×1 projections for K, V (from b_8) and Q (from b_2)
self.proj_k = nn.Conv2d(channels, self.c_prime, 1, bias=False)
self.proj_v = nn.Conv2d(channels, self.c_prime, 1, bias=False)
self.proj_q = nn.Conv2d(channels, self.c_prime, 1, bias=False)
self.gamma = nn.Parameter(torch.zeros(1))
# final 1×1 conv to restore channel dim
self.out_conv = nn.Conv2d(self.c_prime, channels, 1, bias=False)
def _criss_cross_attn(self, Q: torch.Tensor, K: torch.Tensor,
V: torch.Tensor) -> torch.Tensor:
"""
Q, K, V: (B, C', H, W)
Returns attended output: (B, C', H, W)
"""
B, Cp, H, W = Q.shape
scale = math.sqrt(Cp)
# --- Horizontal attention ---
Qh = Q.permute(0, 3, 1, 2).reshape(B * W, Cp, H) # (B*W, C', H)
Kh = K.permute(0, 3, 1, 2).reshape(B * W, Cp, H)
Vh = V.permute(0, 3, 1, 2).reshape(B * W, Cp, H)
Ah = torch.bmm(Qh.permute(0, 2, 1), Kh) / scale # (B*W, H, H)
# INF mask: prevent attending to same position (Eq. 3)
inf_mask = torch.zeros(H, H, device=Q.device)
inf_mask.fill_diagonal_(float('-inf'))
Ah = torch.softmax(Ah + inf_mask, dim=-1)
Yh = torch.bmm(Vh, Ah.permute(0, 2, 1)) # (B*W, C', H)
Yh = Yh.reshape(B, W, Cp, H).permute(0, 2, 3, 1) # (B, C', H, W)
# --- Vertical attention ---
Qw = Q.permute(0, 2, 1, 3).reshape(B * H, Cp, W)
Kw = K.permute(0, 2, 1, 3).reshape(B * H, Cp, W)
Vw = V.permute(0, 2, 1, 3).reshape(B * H, Cp, W)
Aw = torch.bmm(Qw.permute(0, 2, 1), Kw) / scale
Aw = torch.softmax(Aw, dim=-1)
Yw = torch.bmm(Vw, Aw.permute(0, 2, 1))
Yw = Yw.reshape(B, H, Cp, W).permute(0, 2, 1, 3)
return Yh + Yw # (B, C', H, W)
def forward(self, b8: torch.Tensor, b2: torch.Tensor) -> torch.Tensor:
"""
b8: boundary feature map (B, C, H, W) — K and V source
b2: higher-level context (B, C, H, W) — Q source
Returns: attended output with same shape as b8.
"""
K = self.proj_k(b8)
V = self.proj_v(b8)
Q = self.proj_q(b2)
attended = self._criss_cross_attn(Q, K, V) # (B, C', H, W)
# Eq. 3: Y = [(γ·(Yh+Yw) + b2) · b8] · b2
Y = (self.gamma * attended + b2) * b8 * b2
return self.out_conv(Y) # (B, C, H, W)
# ─────────────────────────────────────────────────────────────────────────────
# §6 BOUNDARY PREDICTION MODULE (BPM) (Sec. 3.2, Fig. 3)
#
# Inputs:
# b_prev : higher-level coarse feature (BPM3: r4; BPM2: b3↑; BPM1: b2↑)
# b_cur : current level feature (BPM3: r3; BPM2: r2; BPM1: e1)
#
# Processing steps:
# 1. 1×1 and 3×3 conv to adjust dims → b1, b2
# 2. Element-wise product+add fusion → b3
# 3. Two 3×3 convs to expand uncertainty region → b5
# 4. Reverse attention: invert b2 mask → b6; product with b5 → b7
# 5. Two Res2Net blocks for multi-scale boundary refinement → b8
# 6. MCCAB with b8 as K,V and b2 as Q → attended b8
# 7. 1×1 conv compress → add b4 → coarse mask b9
# ─────────────────────────────────────────────────────────────────────────────
class BPM(nn.Module):
"""Boundary Prediction Module with reverse attention and MCCAB."""
def __init__(self, ch: int = 32):
super().__init__()
# adjust input channels for b_prev and b_cur
self.adj_prev = nn.Sequential(
nn.Conv2d(ch, ch, 1, bias=False), nn.BatchNorm2d(ch))
self.adj_cur = nn.Sequential(
nn.Conv2d(ch, ch, 3, padding=1, bias=False), nn.BatchNorm2d(ch))
# expand uncertainty region (b3 → b5)
self.expand = nn.Sequential(
nn.Conv2d(ch, ch, 3, padding=1, bias=False), nn.BatchNorm2d(ch), nn.ReLU(True),
nn.Conv2d(ch, ch, 3, padding=1, bias=False), nn.BatchNorm2d(ch), nn.ReLU(True))
# compress b2 to single-channel for reverse attention
self.compress = nn.Conv2d(ch, 1, 1, bias=False)
# two Res2Net blocks for multi-scale boundary extraction
self.res2net = nn.Sequential(Res2NetBlock(ch), Res2NetBlock(ch))
# MCCAB for autonomous context prompts
self.mccab = MCCAB(ch)
# final compression to single channel mask
self.to_mask = nn.Conv2d(ch, 1, 1, bias=False)
def forward(self, b_prev: torch.Tensor, b_cur: torch.Tensor
) -> Tuple[torch.Tensor, torch.Tensor]:
"""
Returns:
b9 : single-channel coarse-grained mask logits (B, 1, H, W)
b2 : adjusted high-level feature map for upward passing (B, C, H, W)
"""
b_prev_up = F.interpolate(b_prev, size=b_cur.shape[-2:],
mode='bilinear', align_corners=False)
b1 = self.adj_prev(b_prev_up) # adjusted higher-level features
b2 = self.adj_cur(b_cur) # adjusted current-level features
b3 = b1 * b2 + b1 # element-wise fuse using b2 as template
b5 = self.expand(b3) # expanded uncertainty region
# reverse attention: invert b2 mask (Eq. sigmoid then 1-sigmoid)
b4 = self.compress(b2) # (B, 1, H, W)
b6 = 1.0 - torch.sigmoid(b4) # inverted mask
b7 = b5 * b6.expand_as(b5) # attend to boundary uncertainty
b8 = self.res2net(b7) # multi-scale boundary refinement
# MCCAB: b8 is K,V; b2 is Q (autonomous context prompting)
b8_attn = self.mccab(b8, b2)
b9 = self.to_mask(b8_attn) + b4 # Eq. 4: b9 = Conv(Y) + b4
return b9, b2
# ─────────────────────────────────────────────────────────────────────────────
# §7 SHALLOW FILTERING MODULE (SFM) (Sec. 3.3, Fig. 4)
#
# Mines fine-grained boundary cues from the input image and shallow
# encoder layer e1 to sharpen the coarse-grained mask b0.
#
# Steps:
# s1 = Laplacian(x) — classical edge detection
# s2 = Res2NetLayer1(s1) — learned edge representation (H/2)
# s3 = Concat(DownSample(e1), s2) — multi-scale shallow features
# s4 = s3 + Expand(b0) — guided by coarse mask
# s5 = Res2NetBlocks(s4)
# s6 = Conv(s5) → Upsample → fine-grained mask
# ─────────────────────────────────────────────────────────────────────────────
class LaplacianEdge(nn.Module):
"""Fixed Laplacian edge detector applied to raw input image (Eq. above Sec. 3.3).
Kernel: 1/3 × [[-1,-1,-1],[-1,8,-1],[-1,-1,-1]]
The 1/3 coefficient suppresses noise vs. the unnormalized operator."""
def __init__(self):
super().__init__()
kernel = torch.tensor([[-1, -1, -1],
[-1, 8, -1],
[-1, -1, -1]], dtype=torch.float32) / 3.0
# shape: (1, 1, 3, 3) → expand for 3 input channels
self.register_buffer('weight', kernel.unsqueeze(0).unsqueeze(0))
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""x: (B, 3, H, W) RGB image. Returns (B, 3, H, W) edge map."""
w = self.weight.expand(3, 1, 3, 3)
return F.conv2d(x, w, padding=1, groups=3)
class SFM(nn.Module):
"""Shallow Filtering Module combining Laplacian edge cues with e1 features."""
def __init__(self, e1_ch: int = 64, out_ch: int = 64):
super().__init__()
self.laplacian = LaplacianEdge()
# s1 → s2: Res2Net Layer1 as shallow feature extractor
self.res2_layer1 = nn.Sequential(
nn.Conv2d(3, out_ch, 3, stride=2, padding=1, bias=False),
nn.BatchNorm2d(out_ch), nn.ReLU(True),
Res2NetBlock(out_ch), Res2NetBlock(out_ch))
# s3: concat(e1_down, s2) → reduce channels
self.fuse_conv = nn.Sequential(
nn.Conv2d(e1_ch + out_ch, out_ch, 1, bias=False),
nn.BatchNorm2d(out_ch), nn.ReLU(True))
# s4 = s3 + expand(b0) → refine with Res2Net blocks
self.b0_gate = nn.Conv2d(1, out_ch, 1, bias=False)
self.refine = nn.Sequential(
Res2NetBlock(out_ch), Res2NetBlock(out_ch), Res2NetBlock(out_ch),
nn.Conv2d(out_ch, 1, 1, bias=False))
def forward(self, x: torch.Tensor, e1: torch.Tensor,
b0: torch.Tensor) -> torch.Tensor:
"""
x : (B, 3, H, W) raw image
e1 : (B, 64, H/4, W/4) first encoder output
b0 : (B, 1, H/4, W/4) coarse mask from BPM fusion
Returns: (B, 1, H, W) fine-grained mask logits.
"""
H, W = x.shape[-2:]
# s1: Laplacian edge map
s1 = self.laplacian(x) # (B, 3, H, W)
# s2: Res2Net shallow features at H/2
s2 = self.res2_layer1(s1) # (B, 64, H/2, W/2)
# down-sample e1 to H/4 if needed (already at H/4)
e1_d = F.avg_pool2d(e1, 2) if s2.shape[-2:] != e1.shape[-2:] else e1
# s3: multi-scale shallow features
s3 = self.fuse_conv(torch.cat([e1_d, s2], dim=1)) # (B, 64, H/4, W/4)
# expand b0 to spatial size of s3
b0_up = F.interpolate(b0, size=s3.shape[-2:],
mode='bilinear', align_corners=False)
gate = torch.sigmoid(self.b0_gate(b0_up)) # soft gate
s4 = s3 * gate + s3 # s4 = gated fusion
# refine and upsample to full resolution
s6 = self.refine(s4) # (B, 1, H/4, W/4)
return F.interpolate(s6, size=(H, W), mode='bilinear', align_corners=False)
# ─────────────────────────────────────────────────────────────────────────────
# §8 FULL MSBP-NET (Sec. 3.1, Fig. 2)
#
# Encoder: PVTv2-B2 (or a lightweight substitute if timm unavailable)
# Decoder: 3 × mRFB → 3 × BPM → SFM
#
# Forward pass returns:
# (s6_logits, b1_logits, b2_logits, b3_logits)
# corresponding to L4, L1, L2, L3 deep supervision outputs.
# ─────────────────────────────────────────────────────────────────────────────
def _load_pvtv2():
"""Attempt to load PVTv2-B2 via timm; fall back to a mock encoder."""
try:
import timm
model = timm.create_model('pvt_v2_b2', pretrained=True,
features_only=True)
return model, [64, 128, 320, 512]
except Exception:
warnings.warn("timm not available or PVTv2-B2 not found. Using CNN mock encoder.")
return None, [64, 128, 320, 512]
class MockPVTv2(nn.Module):
"""Lightweight CNN mock of PVTv2's 4-level feature pyramid for testing."""
def __init__(self):
super().__init__()
def stage(ic, oc, stride=2):
return nn.Sequential(
nn.Conv2d(ic, oc, 3, stride=stride, padding=1, bias=False),
nn.BatchNorm2d(oc), nn.GELU(),
nn.Conv2d(oc, oc, 3, padding=1, bias=False),
nn.BatchNorm2d(oc), nn.GELU())
self.s1 = stage(3, 64, stride=4)
self.s2 = stage(64, 128)
self.s3 = stage(128, 320)
self.s4 = stage(320, 512)
def forward(self, x):
e1 = self.s1(x)
e2 = self.s2(e1)
e3 = self.s3(e2)
e4 = self.s4(e3)
return [e1, e2, e3, e4]
class MSBPNet(nn.Module):
"""
Multi-Scale Boundary Prediction Network (MSBP-Net).
Architecture:
Encoder : PVTv2-B2 (or mock CNN)
Decoder : 3×mRFB → 3×BPM (with MCCAB) → SFM (with Laplacian)
"""
def __init__(self, use_pretrained: bool = True):
super().__init__()
pvt, ch = _load_pvtv2() if use_pretrained else (None, [64, 128, 320, 512])
self.encoder = pvt if pvt is not None else MockPVTv2()
e1_ch, e2_ch, e3_ch, e4_ch = ch
# mRFBs — dilation rates follow paper Table 1
self.mrfb4 = mRFB(e4_ch, 32, dilations=(3, 5, 7, 1))
self.mrfb3 = mRFB(e3_ch, 32, dilations=(3, 4, 5, 1))
self.mrfb2 = mRFB(e2_ch, 32, dilations=(1, 2, 3, 1))
# BPMs — all operate on 32-channel compressed maps
self.bpm3 = BPM(32)
self.bpm2 = BPM(32)
self.bpm1 = BPM(32)
# SFM — e1 has e1_ch channels; we project to 64 internally
self.sfm = SFM(e1_ch=e1_ch, out_ch=64)
def forward(self, x: torch.Tensor
) -> Tuple[torch.Tensor, torch.Tensor,
torch.Tensor, torch.Tensor]:
"""
x: (B, 3, H, W) input image (H=W=352 recommended)
Returns: (s6, b1, b2, b3) — four deep supervision mask logits
s6: (B, 1, H, W) final fine-grained mask
b1: (B, 1, H/4, W/4) BPM1 coarse mask
b2: (B, 1, H/8, W/8) BPM2 coarse mask
b3: (B, 1, H/16, W/16) BPM3 coarse mask
"""
# ── Encoder ─────────────────────────────────────────────────────────
feats = self.encoder(x) # [e1, e2, e3, e4]
e1, e2, e3, e4 = feats[0], feats[1], feats[2], feats[3]
# ── mRFB compression ────────────────────────────────────────────────
r4 = self.mrfb4(e4) # (B, 32, H/32, W/32)
r3 = self.mrfb3(e3) # (B, 32, H/16, W/16)
r2 = self.mrfb2(e2) # (B, 32, H/8, W/8 )
# ── BPM cascade: r4→BPM3→BPM2→BPM1 ────────────────────────────────
b3_mask, b3_feat = self.bpm3(r4, r3) # coarse at H/16
b2_mask, b2_feat = self.bpm2(b3_feat, r2) # coarse at H/8
b1_mask, b1_feat = self.bpm1(b2_feat, e1) # coarse at H/4
# ── Fuse coarse masks: b0 = b3↑ + b2↑ + b1 (at H/4) ────────────────
b3_up = F.interpolate(b3_mask, size=b1_mask.shape[-2:],
mode='bilinear', align_corners=False)
b2_up = F.interpolate(b2_mask, size=b1_mask.shape[-2:],
mode='bilinear', align_corners=False)
b0 = b3_up + b2_up + b1_mask # (B, 1, H/4, W/4)
# ── SFM: fine-grained mask via Laplacian + shallow features ─────────
s6 = self.sfm(x, e1, b0) # (B, 1, H, W)
return s6, b1_mask, b2_mask, b3_mask
# ─────────────────────────────────────────────────────────────────────────────
# §9 WEIGHTED LOSS: wBCE + wIoU (Eq. 5–9)
#
# Both losses use the same weight map:
# w_ij = 1 + 5 · |AvgPool(Mask_ij) − Mask_ij|
# Avg pool: kernel=31×31, stride=1, padding=15
# ─────────────────────────────────────────────────────────────────────────────
class WeightedBCEIoULoss(nn.Module):
"""Combined weighted BCE + weighted IoU loss (Eq. 5–8 in paper)."""
def __init__(self):
super().__init__()
self.avg_pool = nn.AvgPool2d(kernel_size=31, stride=1, padding=15)
def _weight_map(self, mask: torch.Tensor) -> torch.Tensor:
"""Eq. 5: w_ij = 1 + 5·|AvgPool(mask) − mask|"""
avg = self.avg_pool(mask)
return 1.0 + 5.0 * (avg - mask).abs()
def forward(self, pred_logits: torch.Tensor,
mask: torch.Tensor) -> torch.Tensor:
"""
pred_logits: raw sigmoid inputs (B, 1, H, W)
mask : ground truth (B, 1, H, W), values in {0, 1}
"""
pred = torch.sigmoid(pred_logits)
w = self._weight_map(mask)
# Weighted BCE (Eq. 6)
bce = -(mask * torch.log(pred + 1e-6) +
(1 - mask) * torch.log(1 - pred + 1e-6))
wbce = (w * bce).mean()
# Weighted IoU (Eq. 7)
inter = (w * pred * mask).sum(dim=(-1, -2))
union = (w * (pred + mask - pred * mask)).sum(dim=(-1, -2))
wiou = 1.0 - (inter + 1.0) / (union + 1.0)
wiou = wiou.mean()
return wbce + wiou
# ─────────────────────────────────────────────────────────────────────────────
# §10 MULTI-SCALE TRAINING LOOP (Sec. 4.2)
#
# Config:
# optimizer : Adam, lr=1e-4
# lr decay : ×0.1 every 50 epochs
# batch size : 16
# input size : 352×352
# scales : [0.75, 1.00, 1.25] applied per batch
# early stop : patience=20 epochs on validation Dice
# deep sup. : L = L4 + L3 + L2 + L1 (s6, b1, b2, b3)
# ─────────────────────────────────────────────────────────────────────────────
class PolyDataset(Dataset):
"""Minimal polyp segmentation dataset wrapper.
Expected directory structure:
root/
images/ *.png
masks/ *.png (binary, same filename as image)
"""
def __init__(self, img_paths: List[str], mask_paths: List[str],
img_size: int = 352):
self.imgs = img_paths
self.masks = mask_paths
self.size = img_size
self.to_tensor = transforms.ToTensor()
self.normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
def __len__(self): return len(self.imgs)
def __getitem__(self, idx):
from PIL import Image
img = Image.open(self.imgs[idx]).convert('RGB').resize(
(self.size, self.size), Image.BILINEAR)
mask = Image.open(self.masks[idx]).convert('L').resize(
(self.size, self.size), Image.NEAREST)
img = self.normalize(self.to_tensor(img))
mask = (self.to_tensor(mask) > 0.5).float()
return img, mask
def multiscale_batch(imgs: torch.Tensor, masks: torch.Tensor,
scales: List[float] = [0.75, 1.0, 1.25],
base: int = 352) -> Tuple[torch.Tensor, torch.Tensor]:
"""Apply random multi-scale resizing per batch (Sec. 4.2)."""
s = np.random.choice(scales)
sz = int(base * s)
sz = sz + (sz % 2) # ensure even size
out_imgs = F.interpolate(imgs, (sz, sz), mode='bilinear', align_corners=False)
out_masks = F.interpolate(masks, (sz, sz), mode='nearest')
return out_imgs, out_masks
def dice_score(pred_logits: torch.Tensor, mask: torch.Tensor,
threshold: float = 0.5) -> float:
"""Compute Dice Similarity Coefficient for a batch."""
pred = (torch.sigmoid(pred_logits) > threshold).float()
inter = (pred * mask).sum()
union = pred.sum() + mask.sum()
return ((2 * inter + 1e-6) / (union + 1e-6)).item()
def train_msbpnet(
train_imgs: List[str], train_masks: List[str],
val_imgs: List[str], val_masks: List[str],
save_path: str = 'msbpnet_best.pth',
max_epochs: int = 80,
patience: int = 20,
batch_size: int = 16,
lr: float = 1e-4,
img_size: int = 352,
device: str = 'cuda'):
"""
Full training loop as described in Sec. 4.2 of the paper.
Args:
train_imgs : list of absolute paths to training images
train_masks : list of absolute paths to training binary masks
val_imgs : list of absolute paths to validation images
val_masks : list of absolute paths to validation binary masks
save_path : path to save best model checkpoint
max_epochs : maximum training epochs (paper: ~50, early stopped)
patience : early stopping patience in epochs
batch_size : training batch size (paper: 16)
lr : initial learning rate (paper: 1e-4)
img_size : input spatial resolution (paper: 352)
device : 'cuda' or 'cpu'
"""
dev = torch.device(device if torch.cuda.is_available() else 'cpu')
model = MSBPNet(use_pretrained=True).to(dev)
crit = WeightedBCEIoULoss().to(dev)
optim = torch.optim.Adam(model.parameters(), lr=lr)
sched = torch.optim.lr_scheduler.StepLR(optim, step_size=50, gamma=0.1)
train_ds = PolyDataset(train_imgs, train_masks, img_size)
val_ds = PolyDataset(val_imgs, val_masks, img_size)
train_dl = DataLoader(train_ds, batch_size=batch_size,
shuffle=True, num_workers=4, pin_memory=True)
val_dl = DataLoader(val_ds, batch_size=1,
shuffle=False, num_workers=2, pin_memory=True)
best_dice = 0.0
no_improve = 0
for epoch in range(max_epochs):
model.train()
total_loss = 0.0
for imgs, masks in train_dl:
imgs, masks = imgs.to(dev), masks.to(dev)
# multi-scale resize per batch (Sec. 4.2)
imgs, masks = multiscale_batch(imgs, masks)
optim.zero_grad(set_to_none=True)
s6, b1, b2, b3 = model(imgs)
# deep supervision: resize all mask predictions to input size
target = F.interpolate(masks, size=imgs.shape[-2:],
mode='nearest')
L4 = crit(s6, target)
L1 = crit(F.interpolate(b1, size=imgs.shape[-2:],
mode='bilinear', align_corners=False), target)
L2 = crit(F.interpolate(b2, size=imgs.shape[-2:],
mode='bilinear', align_corners=False), target)
L3 = crit(F.interpolate(b3, size=imgs.shape[-2:],
mode='bilinear', align_corners=False), target)
loss = L4 + L3 + L2 + L1 # Eq. 9: L = L4 + L3 + L2 + L1
loss.backward()
optim.step()
total_loss += loss.item()
sched.step()
# ── Validation ───────────────────────────────────────────────────────
model.eval()
val_dice = 0.0
with torch.no_grad():
for imgs, masks in val_dl:
imgs, masks = imgs.to(dev), masks.to(dev)
s6, _, _, _ = model(imgs)
val_dice += dice_score(s6, masks)
val_dice /= len(val_dl)
avg_loss = total_loss / len(train_dl)
print(f"Epoch {epoch+1:03d}/{max_epochs} "
f"Loss: {avg_loss:.4f} Val Dice: {val_dice:.4f} "
f"LR: {sched.get_last_lr()[0]:.6f}")
if val_dice > best_dice:
best_dice = val_dice
no_improve = 0
torch.save(model.state_dict(), save_path)
print(f" ✓ Saved new best model (Dice={best_dice:.4f})")
else:
no_improve += 1
if no_improve >= patience:
print(f"\nEarly stopping at epoch {epoch+1} "
f"(no improvement for {patience} epochs).")
break
print(f"\nTraining complete. Best Val Dice: {best_dice:.4f}")
return model
# ─────────────────────────────────────────────────────────────────────────────
# §11 INFERENCE PIPELINE + COMPLEXITY COUNTER
# ─────────────────────────────────────────────────────────────────────────────
def predict_mask(model: MSBPNet, img_path: str,
device: str = 'cuda',
threshold: float = 0.5) -> np.ndarray:
"""
Run inference on a single colonoscopy image.
Returns binary mask as numpy array (H, W) with values in {0, 1}.
"""
from PIL import Image
dev = torch.device(device if torch.cuda.is_available() else 'cpu')
model.eval().to(dev)
to_tensor = transforms.ToTensor()
normalizer = transforms.Normalize(
mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
img_orig = Image.open(img_path).convert('RGB')
orig_size = img_orig.size # (W, H) for PIL
img_r = img_orig.resize((352, 352), Image.BILINEAR)
tensor = normalizer(to_tensor(img_r)).unsqueeze(0).to(dev)
with torch.no_grad():
s6, _, _, _ = model(tensor) # (1, 1, 352, 352)
prob = torch.sigmoid(s6).squeeze().cpu().numpy() # (352, 352)
# resize back to original image dimensions
from PIL import Image as PILImage
prob_img = PILImage.fromarray((prob * 255).astype(np.uint8))
prob_orig = np.array(prob_img.resize(orig_size, PILImage.BILINEAR)) / 255.0
return (prob_orig >= threshold).astype(np.uint8)
def count_complexity(img_size: int = 352) -> None:
"""Print FLOPs and parameter counts matching paper Table 1."""
model = MSBPNet(use_pretrained=False)
total_params = sum(p.numel() for p in model.parameters()) / 1e6
print(f"Total parameters: {total_params:.3f} M")
try:
from thop import profile
dummy = torch.zeros(1, 3, img_size, img_size)
macs, _ = profile(model, inputs=(dummy,), verbose=False)
print(f"Total FLOPs: {macs / 1e9:.3f} G (at {img_size}×{img_size})")
except ImportError:
print("thop not installed — install with: pip install thop")
# Paper values for reference (Table 1)
print("\nPaper reported values (Table 1):")
print(" Total decoder FLOPs : 3.231 G")
print(" Total decoder Params: 0.667 M")
print(" Total model FLOPs : 12.859 G (encoder 9.628 + decoder 3.231)")
print(" Total model Params: 25.517 M (encoder 24.850 + decoder 0.667)")
# ─────────────────────────────────────────────────────────────────────────────
# ENTRY POINT
# ─────────────────────────────────────────────────────────────────────────────
if __name__ == "__main__":
print("=" * 70)
print(" MSBP-Net: Multi-Scale Boundary Prediction Network")
print(" Pan, Ding, Li, Liu, Wang, Hua, Tang & Zhong")
print(" Pattern Recognition 170 (2026) 112101")
print("=" * 70)
# ── Sanity-check forward pass ────────────────────────────────────────
model = MSBPNet(use_pretrained=False) # use mock encoder for testing
model.eval()
dummy = torch.randn(2, 3, 352, 352) # batch of 2 images
with torch.no_grad():
s6, b1, b2, b3 = model(dummy)
print("\n── Output shapes ─────────────────────────────────────────────────")
print(f" s6 (final mask) : {tuple(s6.shape)}")
print(f" b1 (BPM1 coarse) : {tuple(b1.shape)}")
print(f" b2 (BPM2 coarse) : {tuple(b2.shape)}")
print(f" b3 (BPM3 coarse) : {tuple(b3.shape)}")
# ── Loss computation ─────────────────────────────────────────────────
crit = WeightedBCEIoULoss()
target = torch.randint(0, 2, (2, 1, 352, 352)).float()
L4 = crit(s6, target)
L1 = crit(F.interpolate(b1, (352, 352), mode='bilinear', align_corners=False), target)
L2 = crit(F.interpolate(b2, (352, 352), mode='bilinear', align_corners=False), target)
L3 = crit(F.interpolate(b3, (352, 352), mode='bilinear', align_corners=False), target)
total = L4 + L3 + L2 + L1
print(f"\n── Deep supervision losses ───────────────────────────────────────")
print(f" L4 (SFM final) : {L4.item():.4f}")
print(f" L3 (BPM3 coarse): {L3.item():.4f}")
print(f" L2 (BPM2 coarse): {L2.item():.4f}")
print(f" L1 (BPM1 coarse): {L1.item():.4f}")
print(f" L (total) : {total.item():.4f}")
# ── Parameter count ──────────────────────────────────────────────────
count_complexity(img_size=352)
# ── Dice on random predictions ────────────────────────────────────────
d = dice_score(s6, target[:, :, :s6.shape[-2], :s6.shape[-1]])
print(f"\n── Dice (random weights, untrained): {d:.4f}")
print("\nForward pass complete — architecture instantiated correctly.\n")
Access the Paper and Resources
The full MSBP-Net framework, ablation results, and five-dataset benchmark comparison are available in the published article. This research was conducted by Pan, Ding, Li, Liu, Wang, Hua, Tang, and Zhong at the Sichuan University of Science and Engineering, published in Pattern Recognition, 2026.
Pan, X.-L., Ding, J.-R., Li, X., Liu, S., Wang, J., Hua, B., Tang, G.-Z., & Zhong, C.-H. (2026). MSBP-Net: A multi-scale boundary prediction network for automated polyp segmentation. Pattern Recognition, 170, 112101. https://doi.org/10.1016/j.patcog.2025.112101
This article is an independent editorial analysis of peer-reviewed research published in Pattern Recognition (Elsevier). The views and commentary expressed here reflect the editorial perspective of this site and do not represent the views of the original authors or their institutions. Code is provided for educational purposes to illustrate technical concepts and has not been validated against the authors’ original implementation. Always refer to the original publication for authoritative details.
Explore More on AI Research
If this analysis sparked your interest, here is more of what we cover across the site—from foundational tutorials to the latest breakthroughs in medical imaging, multimodal learning, and knowledge distillation.
