Introduction: The Future of Automated Medical Diagnosis
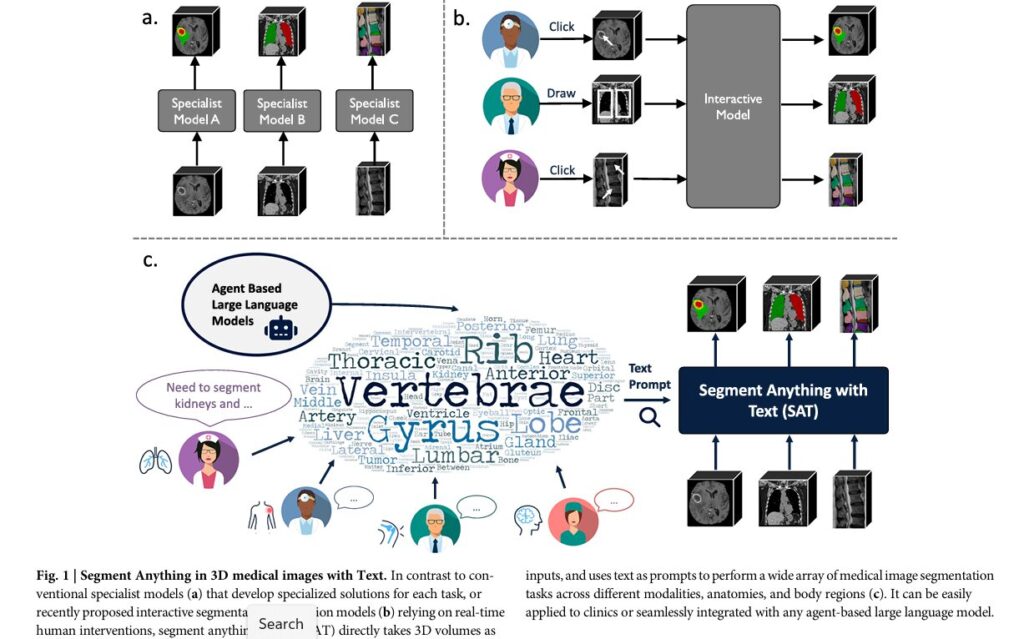
The traditional workflow in medical imaging has remained largely unchanged for decades. Radiologists manually examine thousands of scans, carefully delineating regions of interest slice by slice—a process that is both time-consuming and prone to human error. But what if an AI model could segment any anatomical structure, lesion, or tissue type across virtually any medical imaging modality using nothing more than a simple text description?
This is precisely what Segment Anything with Text (SAT) achieves, introducing the first knowledge-enhanced foundation model for 3D medical volume segmentation that accepts medical terminology as text prompts and seamlessly tackles various medical image segmentation tasks across modalities, anatomies, and body regions. This breakthrough represents far more than incremental progress in medical AI—it signals a fundamental shift in how the healthcare industry approaches medical image analysis at scale.
This article explores the transformative capabilities of SAT, its architectural innovations, the unprecedented dataset that powers it, and its implications for the future of clinical practice and medical artificial intelligence.
The Evolution of Medical Image Segmentation: From Specialists to Generalists
Understanding the Current Landscape
Medical image segmentation is the process of automatically identifying and delineating regions of interest—organs, lesions, and tissues—within medical images. This plays a crucial role in numerous clinical applications, such as disease diagnosis, treatment planning, and disease progression tracking, as well as in medical research.
Historically, the medical imaging AI community has operated under a specialist paradigm. Individual research groups develop and train specialized neural networks for specific tasks: one model for liver segmentation, another for lung cancer detection, yet another for brain tumor delineation. While these specialist models demonstrate impressive segmentation capabilities, their major drawback lies in their narrow specialization—they are designed and optimized for distinct regions of interest and imaging modalities, require distinct preprocessing methods for each dataset, and often fall short in diverse and dynamic clinical environments where adaptability to new conditions and imaging techniques is essential.
This fragmented approach creates significant practical challenges in real-world clinical deployment. Hospitals cannot maintain dozens of specialized models, each requiring unique infrastructure, monitoring, and updates. The specialist paradigm has become a bottleneck preventing AI’s broader adoption in clinical settings.
Limitations of Interactive Segmentation Approaches
Recent efforts to address this limitation have explored adapting foundation models from computer vision. Models typically perform 2D slice segmentation, which is later fused into 3D volumes through post-processing, overlooking crucial contextual information in 3D radiological imaging. Additionally, models often require point or box inputs as prompts, making them interactive segmentation models that require considerable manual effort for use in practice, and they suffer from significant domain gaps from image statistics to domain-specific medical knowledge.
While interactive approaches represent progress, they still demand substantial human intervention—physicians must click or draw on images to guide segmentation, reducing efficiency and scalability.
SAT: A Paradigm Shift in Medical Image Analysis
Core Innovation: Text-Prompted Segmentation
The first knowledge-enhanced foundation model for 3D medical volume segmentation with medical terminology as text prompt distinguishes itself from previous medical segmentation paradigms by being seamlessly applied to clinical practice or integrated with any large language model.
Key Advantages of the Text-Prompt Paradigm:
- Automatic segmentation without point-and-click interaction
- 3D volumetric processing preserving crucial spatial context
- Knowledge-enhanced representations leveraging anatomical relationships
- Large language model integration enabling agent-based clinical workflows
- Scalability across diverse imaging modalities and anatomical structures
The shift from interactive spatial prompts (points, boxes) to text prompts represents a fundamental recognition that medical knowledge is fundamentally linguistic. Radiologists think in anatomical terminology, clinical reports use standardized medical language, and integrating large language models with segmentation requires text-based communication.
Model Variants: SAT-Nano and SAT-Pro
To balance computational cost and performance, SAT includes two variants: SAT-Nano with 110M parameters and SAT-Pro with 447M parameters. This dual-model strategy addresses the practical reality that different clinical settings have varying computational constraints. Hospital systems with substantial GPU resources can deploy SAT-Pro for maximum accuracy, while resource-constrained clinics can utilize SAT-Nano with acceptable performance degradation.
This scalability approach mirrors successful strategies in foundation model development, where organizations release multiple model sizes to maximize adoption across diverse infrastructure capabilities.
Building the Foundation: SAT-DS, the Largest Medical Segmentation Dataset
Dataset Scope and Composition
The technical achievements of SAT rest upon an unprecedented foundation of curated medical imaging data. The researchers collected and integrated 72 diverse publicly available medical segmentation datasets, totaling 22,186 scans including CT, MRI, and PET, with 302,033 segmentation annotations spanning 8 different regions of the human body: brain, head and neck, upper limb, thorax, abdomen, pelvis, and lower limb.
To contextualize this achievement: The data collection is more than fourth times the size of the largest dataset (BraTS2023-GLI) in terms of volume number, and nearly triple the most comprehensive dataset (DAP Atlas) in terms of the class number.
The Challenge of Dataset Unification
Creating SAT-DS required far more than simply concatenating existing datasets. Medical datasets represent decades of collection efforts under varying clinical protocols, annotation standards, and quality criteria. The research team addressed critical challenges:
1. Unified Annotation System
- Manually verified anatomical targets across all datasets
- Standardized naming conventions (e.g., “lumbar vertebrae 1 (L1)”)
- Resolved contradictions between overlapping classes
- Merged fine-grained classes to maintain consistency
2. Image Preprocessing Standardization
- Reoriented images to specific axcodes
- Respaced to uniform voxel size of 1 × 1 × 3 mm³
- Applied modality-specific intensity normalization:
- CT images: Truncated to [-500, 1000] HU with z-score normalization
- MRI/PET images: Clipped at 0.5% and 99.5% percentiles with z-score normalization
- Cropped to non-zero regions
These preprocessing standards ensure the model learns consistent representations across diverse source datasets, preventing dataset-specific biases from dominating the learned features.
Architectural Innovation: Knowledge Injection Through Contrastive Learning
The Multimodal Knowledge Graph
The researchers constructed a multimodal knowledge tree integrating anatomical concepts and definitions throughout the human body, creating 6502 anatomical terminologies with concept-definition pairs and extensive relationships. This knowledge graph draws from two primary sources:
1. Unified Medical Language System (UMLS)
- 229,435 biomedical terminologies and definitions
- 1,048,575 relationship triplets from 60+ vocabularies
- Comprehensive but functionally-focused definitions
2. Search Engine-Augmented Knowledge
- 6,502 anatomy-specific concept definitions emphasizing morphological features
- Location, shape, appearance, structure, and spatial relationships
- 38,344 relationships extracted through GPT-4 analysis
- Authority-sourced definitions from medical institutions
Knowledge-Enhanced Representation Learning
The power of SAT derives from injecting this rich medical knowledge into neural representations through contrastive learning. The framework operates on the principle that similar anatomical concepts—whether represented as text definitions, visual atlases, or concept relationships—should produce similar embeddings in latent space.
The Contrastive Learning Objective:
For a batch of positive knowledge pairs linked by medical terminology:
\[ L_{\text{knowledge}} = -\frac{1}{N}\sum_{i=1}^{N}\left(\log\frac{\exp(z_i \cdot z’_i/\tau)}{\sum_{k=1}^{N}\mathbb{1}_{i\neq k}\exp(z_i \cdot z’_k/\tau)} + \log\frac{\exp(z_i \cdot z’_i/\tau)}{\sum_{k=1}^{N}\mathbb{1}_{i\neq k}\exp(z_k \cdot z’_i/\tau)}\right) \]Where τ = 0.07 is the temperature parameter, and ziz_i zi and zi′z’_i zi′ represent paired embeddings from potentially different modalities (text, visual) but encoding the same anatomical concept.
Key Insight: This approach differs fundamentally from standard contrastive learning by enforcing alignment between textual definitions, visual atlases, and concept relationships—creating a unified embedding space where medical knowledge manifests as geometric structure.
Ablation Study Results: The Value of Knowledge Injection
Testing with different text encoders (BERT-Base, CLIP, MedCPT, and their proprietary knowledge-enhanced encoder) demonstrated that their proposed knowledge-enhanced text encoder achieved 99.18% Recall@1 on anatomy concept-definition retrieval, compared to only 0.08% for BERT-Base, 4.13% for CLIP, and 11.19% for MedCPT.
This dramatic difference illustrates how domain-specific knowledge injection creates embeddings that genuinely capture medical concepts rather than relying on surface-level language similarity.
Performance Benchmarks: SAT Versus Established Baselines
Comparison with Specialist Models
Comparing SAT-Pro against 72 specialist nnU-Net models trained individually on each dataset, SAT-Pro shows comparable performance to the ensemble of 72 nnU-Nets over 497 categories, while using only 20% of their combined model parameters (447M vs. 2.2B+).
Benchmark Results (Internal Evaluation):
$$ \begin{array}{|l|c|c|c|c|l|} \hline \textbf{Model} & \textbf{Parameters} & \textbf{Datasets} & \textbf{Categories} & \textbf{Avg DSC} & \textbf{Notes} \\ \hline \text{SAT-Pro} & 447M & \text{All 72} & 497 & \text{Comparable} & \text{Generalist} \\ \hline \text{72 × nnU-Net} & 2.2B+ & \text{Each specialized} & \text{12 avg} & \text{Reference} & \text{Specialist ensemble} \\ \hline \text{SAT-Nano} & 110M & \text{All 72} & 497 & \text{Lower} & \text{Resource-constrained} \\ \hline \text{U-Mamba} & \approx 400M & \text{Each dataset} & \text{Variable} & \text{Comparable} & \text{Specialist} \\ \hline \text{SwinUNETR} & \approx 400M & \text{Each dataset} & \text{Variable} & \text{Lower} & \text{Specialist} \\ \hline \end{array} $$This represents a remarkable achievement: a single generalist model achieves performance parity with 72 specialized systems while consuming 1/5 the parameters and offering 40× greater semantic coverage (497 vs. 12 average classes per specialist model).
Superior Generalization on External Data
On external validation datasets (AbdomenAtlas and LiQA, which were excluded from SAT-DS training), SAT-Pro achieves higher performance than all baselines (+3.7% average DSC), demonstrating superior generalization ability. This external evaluation is particularly significant—it simulates real-world deployment scenarios where models encounter multi-center data with different acquisition protocols and patient populations.
Comparison with Interactive Segmentation: MedSAM
Compared with the interactive approach MedSAM, SAT-Pro consistently outperforms across all 7 human body regions with +7.1% average Dice Similarity Coefficient (DSC) improvement, while showing enhanced scalability and robustness.
Particularly compelling is the practical efficiency difference: MedSAM requires 10+ to 60+ prompts to segment a single target in a 3D image scan across different body regions, while SAT requires only a single text prompt to segment the entire 3D scan. For a radiologist segmenting a full abdominal CT scan with 10+ structures of interest, this translates from 100+ manual interactions to simply typing anatomical names.
Clinical Applications and LLM Integration
SAT as an Interface Between Language and Medicine
Perhaps the most transformative aspect of SAT is its potential integration with large language models. SAT can be seamlessly applied as an interface between natural language and segmentation, acting as a high-performance and efficient agent for language models. GPT-4 can analyze and extract anatomical targets of interest from real clinical reports and prompt SAT to segment them on clinical images, providing visual cues for the clinical report and enhancing its interpretability.
Three Emerging Clinical Workflows
1. Report-Guided Segmentation A radiologist writes a clinical report noting “suspicious lesion in the right lung.” GPT-4 extracts “right lung” and “lesion,” prompting SAT to automatically segment both structures. The radiologist receives immediate visual confirmation, accelerating report writing and reducing cognitive load.
2. Conversational Clinical Assistance Physicians can query SAT through natural language without memorizing anatomical terminology. “Show me the liver and any lesions within it” automatically segments appropriate structures, democratizing access to AI-assisted analysis for non-specialist clinicians.
3. Comprehensive EHR-Integrated Analysis Combining radiology images, patient complaints, and historical context, LLM-SAT systems provide automated diagnostic pipelines. A patient presenting with “abdominal pain and weight loss” receives automatic segmentation of pancreas, liver, and surrounding structures relevant to differential diagnosis.
The Long-Tail Distribution Problem: Why Knowledge Matters
Medical segmentation datasets exhibit severe long-tail distributions. In SAT-DS-Nano, the 10 “head” classes account for 12.75% of annotations, while 150 classes with minimum annotations account for only 3.25% despite comprising 34.97% of classes.
Models trained purely on image-label pairs struggle catastrophically on rare structures. Knowledge injection specifically addresses this pathology: Using their proposed knowledge-enhanced text encoder, performance improvements are most pronounced on “tail” classes, demonstrating that knowledge enhancement particularly improves rare structure segmentation where visual examples are limited.
This capability is clinically crucial—many segmentation targets are pathological lesions or anatomical variants encountered infrequently in training data but critical for clinical decision-making.
Limitations and Future Directions
No breakthrough technology emerges without acknowledged limitations. The research team identifies key areas for continued development:
- Regional Performance Gaps: SAT-Pro still lags behind nnU-Nets in some anatomical regions, suggesting scaling to larger models remains beneficial
- Unseen Categories: Generalization to previously unseen lesion types remains limited, motivating exploration of open-vocabulary segmentation
- Demographic Representation: SAT-DS remains skewed toward adult populations with limited pediatric and fetal data, potentially affecting generalization across age groups
- Clinical Deployment: While inference speed is suitable for clinical use, further optimization for standard hardware remains important
Conclusion: A New Era in Medical AI
The significance of SAT is demonstrated through multiple dimensions: it represents an important step toward a universal segmentation model in medical scenarios that can flexibly handle diverse images and targets within a single generalist model, effectively replacing the need for dozens of specialist models while achieving superior cross-center transferability.
SAT exemplifies how foundation models, knowledge graphs, and large language models converge to create genuinely transformative medical AI. Rather than forcing clinicians to adapt to technology, SAT adapts to how clinicians think and communicate—through medical language itself.
The path forward is clear: As medical AI matures, expect the specialist paradigm to fade. Future systems will combine the knowledge of foundation models with the linguistic fluency of large language models, delivering AI that is simultaneously more capable, more accessible, and more clinically integrated than anything available today.
Take Action: Engage with the Future of Medical AI
Are you a researcher, clinician, or healthcare technologist interested in how foundation models are revolutionizing medical imaging? The implications of text-prompted segmentation extend far beyond technical achievement—they reshape how radiologists work, how hospitals deploy AI infrastructure, and how medical knowledge translates into clinical action.
We invite you to:
- Share your perspective: How might text-prompted segmentation transform your clinical workflow or research program?
- Explore the code: The SAT implementation and SAT-DS are publicly available for reproducibility and benchmarking
- Consider the implications: What new clinical applications become possible when any anatomical structure can be automatically segmented through natural language?
The democratization of medical AI through accessible, knowledge-enhanced foundation models isn’t inevitable—it requires ongoing investment in research, thoughtful clinical integration, and dialogue between technologists and medical professionals. Your insights and engagement help shape this transformation.
Subscribe to stay informed about breakthrough developments in medical AI, foundation models for healthcare, and clinical AI deployment strategies.
References and Further Reading
The original research introducing SAT was published in npj Digital Medicine by Ziheng Zhao, Yao Zhang, and colleagues, presenting comprehensive evaluation across 72 datasets with 497 anatomical categories and demonstrating superior generalization compared to specialist models, interactive segmentation approaches, and concurrent text-prompted systems. The dataset, code, and trained models are publicly available to accelerate research and clinical adoption.
Below is the Complete End-to-End Implementation of SAT Model.
SAT/
├── config/
│ ├── config.yaml
│ └── dataset_config.json
├── data/
│ ├── __init__.py
│ ├── dataset.py
│ ├── preprocessing.py
│ └── knowledge_graph.py
├── models/
│ ├── __init__.py
│ ├── encoders.py
│ ├── decoders.py
│ └── sat_model.py
├── training/
│ ├── __init__.py
│ ├── trainer.py
│ ├── losses.py
│ └── optimizers.py
├── evaluation/
│ ├── __init__.py
│ ├── metrics.py
│ └── validator.py
├── utils/
│ ├── __init__.py
│ ├── logger.py
│ ├── visualization.py
│ └── checkpoint.py
├── inference/
│ ├── __init__.py
│ └── predictor.py
├── train.py
├── evaluate.py
├── requirements.txt
└── README.md# 1. Configuration Files
# Model Configuration
model:
name: "SAT"
variant: "pro" # "nano" or "pro"
# Text Encoder
text_encoder:
type: "biobert"
pretrained_model: "dmis-lab/biobert-base-cased"
hidden_dim: 768
num_layers: 12
dropout: 0.1
freeze_layers: 0 # Number of layers to freeze
# Visual Encoder
visual_encoder:
type: "unet3d"
in_channels: 1
num_features: 64
depth: 5
# Query Decoder
query_decoder:
type: "transformer"
num_layers: 6
num_heads: 8
hidden_dim: 512
ffn_dim: 2048
dropout: 0.1
# Visual Decoder
visual_decoder:
type: "unet3d"
num_features: 64
depth: 5
out_channels: 256
# Knowledge Injection
knowledge_injection:
enabled: true
contrastive_temp: 0.07
embedding_dim: 768
# Data Configuration
data:
dataset_path: "/path/to/SAT-DS"
batch_size: 4
num_workers: 8
image_size: [288, 288, 96]
# Preprocessing
preprocessing:
normalize_ct: true
ct_hu_range: [-500, 1000]
normalize_mri: true
mri_percentile: [0.5, 99.5]
normalize_pet: true
pet_percentile: [0.5, 99.5]
# Augmentation
augmentation:
enable_random_flip: true
enable_random_rotation: true
enable_random_zoom: true
enable_intensity_shift: true
# Training Configuration
training:
num_epochs: 100
learning_rate: 1e-4
weight_decay: 1e-5
warmup_epochs: 5
# Loss weights
loss_weights:
segmentation: 1.0
knowledge: 0.5
# Optimizer
optimizer: "adamw"
# Scheduler
scheduler: "cosine"
# Precision
use_amp: true
# Checkpointing
checkpoint_interval: 5
checkpoint_dir: "./checkpoints"
# Evaluation Configuration
evaluation:
val_interval: 1
metrics:
- "dice"
- "nsd"
- "hausdorff"
# Device Configuration
device:
cuda: true
distributed: false
num_gpus: 1
# Logging
logging:
log_dir: "./logs"
log_interval: 100
save_tensorboard: true# config/dataset_config.json
{
"datasets": {
"brain": {
"modalities": ["CT", "MRI"],
"anatomies": [
"brain stem",
"superior frontal gyrus",
"lateral ventricle",
"postcentral gyrus",
"superior parietal gyrus",
"cingulate gyrus"
],
"num_samples": 3500,
"num_annotations": 45000
},
"thorax": {
"modalities": ["CT", "MRI"],
"anatomies": [
"lung",
"heart atrium",
"aorta",
"rib",
"autochthon"
],
"num_samples": 4200,
"num_annotations": 52000
},
"abdomen": {
"modalities": ["CT", "MRI"],
"anatomies": [
"liver",
"stomach",
"colon",
"kidney",
"spleen",
"pancreas"
],
"num_samples": 6800,
"num_annotations": 85000
},
"pelvis": {
"modalities": ["CT", "MRI"],
"anatomies": [
"transition zone of prostate",
"peripheral zone of prostate",
"bladder"
],
"num_samples": 1200,
"num_annotations": 15000
},
"spine": {
"modalities": ["CT", "MRI"],
"anatomies": [
"lumbar vertebrae",
"sacrum",
"intervertebral disc"
],
"num_samples": 2800,
"num_annotations": 35000
}
},
"modality_stats": {
"CT": {
"hu_range": [-500, 1000],
"typical_spacing": [1.0, 1.0, 3.0]
},
"MRI": {
"intensity_range": [0, 4096],
"typical_spacing": [1.0, 1.0, 3.0]
},
"PET": {
"intensity_range": [0, 100],
"typical_spacing": [2.0, 2.0, 2.0]
}
},
"total_scans": 22186,
"total_annotations": 302033,
"num_classes": 497
}# 2. Data Loading and Preprocessing
import numpy as np
import torch
from torch.nn import functional as F
from scipy import ndimage
from typing import Tuple, Dict, Optional
import logging
logger = logging.getLogger(__name__)
class MedicalImagePreprocessor:
"""Preprocessing pipeline for medical images."""
def __init__(self, config: Dict):
self.config = config
self.target_spacing = config.get('target_spacing', [1.0, 1.0, 3.0])
self.target_size = config.get('image_size', [288, 288, 96])
def preprocess_ct(self, image: np.ndarray) -> np.ndarray:
"""Preprocess CT image."""
# Clamp HU values
hu_min, hu_max = self.config['preprocessing']['ct_hu_range']
image = np.clip(image, hu_min, hu_max)
# Normalize
image = (image - hu_min) / (hu_max - hu_min)
return image.astype(np.float32)
def preprocess_mri(self, image: np.ndarray) -> np.ndarray:
"""Preprocess MRI image."""
# Percentile clipping
p_low, p_high = self.config['preprocessing']['mri_percentile']
v_low = np.percentile(image, p_low)
v_high = np.percentile(image, p_high)
image = np.clip(image, v_low, v_high)
image = (image - v_low) / (v_high - v_low + 1e-5)
return image.astype(np.float32)
def preprocess_pet(self, image: np.ndarray) -> np.ndarray:
"""Preprocess PET image."""
p_low, p_high = self.config['preprocessing']['pet_percentile']
v_low = np.percentile(image, p_low)
v_high = np.percentile(image, p_high)
image = np.clip(image, v_low, v_high)
image = (image - v_low) / (v_high - v_low + 1e-5)
return image.astype(np.float32)
def resample(self, image: np.ndarray,
original_spacing: Tuple,
target_spacing: Tuple) -> np.ndarray:
"""Resample image to target spacing."""
zoom_factors = tuple(o / t for o, t in zip(original_spacing, target_spacing))
resampled = ndimage.zoom(image, zoom_factors, order=1)
return resampled
def crop_or_pad(self, image: np.ndarray,
target_shape: Tuple) -> np.ndarray:
"""Crop or pad image to target shape."""
current_shape = image.shape
if all(c >= t for c, t in zip(current_shape, target_shape)):
# Crop from center
slices = tuple(
slice((c - t) // 2, (c - t) // 2 + t)
for c, t in zip(current_shape, target_shape)
)
return image[slices]
else:
# Pad
padding = tuple(
((t - c) // 2, (t - c) - (t - c) // 2)
for c, t in zip(current_shape, target_shape)
)
return np.pad(image, padding, mode='constant', constant_values=0)
def __call__(self, image: np.ndarray,
modality: str,
original_spacing: Tuple) -> np.ndarray:
"""Apply full preprocessing pipeline."""
# Preprocess based on modality
if modality.upper() == 'CT':
image = self.preprocess_ct(image)
elif modality.upper() == 'MRI':
image = self.preprocess_mri(image)
elif modality.upper() == 'PET':
image = self.preprocess_pet(image)
# Resample
image = self.resample(image, original_spacing, self.target_spacing)
# Crop or pad
image = self.crop_or_pad(image, self.target_size)
return image
class DataAugmentation:
"""Data augmentation for medical images."""
def __init__(self, config: Dict):
self.config = config['augmentation']
def random_flip(self, image: np.ndarray,
label: np.ndarray) -> Tuple[np.ndarray, np.ndarray]:
"""Random flip along axes."""
for axis in range(3):
if np.random.rand() < 0.5:
image = np.flip(image, axis=axis)
label = np.flip(label, axis=axis)
return image, label
def random_rotation(self, image: np.ndarray,
label: np.ndarray) -> Tuple[np.ndarray, np.ndarray]:
"""Random rotation."""
if np.random.rand() < 0.5:
angle = np.random.uniform(-30, 30)
image = ndimage.rotate(image, angle, reshape=False, order=1)
label = ndimage.rotate(label, angle, reshape=False, order=0)
return image, label
def random_zoom(self, image: np.ndarray,
label: np.ndarray) -> Tuple[np.ndarray, np.ndarray]:
"""Random zoom."""
if np.random.rand() < 0.5:
zoom_factor = np.random.uniform(0.9, 1.1)
image = ndimage.zoom(image, zoom_factor, order=1)
label = ndimage.zoom(label, zoom_factor, order=0)
return image, label
def intensity_shift(self, image: np.ndarray) -> np.ndarray:
"""Random intensity shift."""
if np.random.rand() < 0.5:
shift = np.random.uniform(-0.1, 0.1)
image = np.clip(image + shift, 0, 1)
return image
def __call__(self, image: np.ndarray,
label: np.ndarray) -> Tuple[np.ndarray, np.ndarray]:
"""Apply augmentations."""
if self.config.get('enable_random_flip', True):
image, label = self.random_flip(image, label)
if self.config.get('enable_random_rotation', True):
image, label = self.random_rotation(image, label)
if self.config.get('enable_random_zoom', True):
image, label = self.random_zoom(image, label)
if self.config.get('enable_intensity_shift', True):
image = self.intensity_shift(image)
return image, label# data/knowledge_graph.py
import json
from typing import Dict, List, Tuple, Set
import numpy as np
from collections import defaultdict
import logging
logger = logging.getLogger(__name__)
class AnatomyKnowledgeGraph:
"""Knowledge graph for anatomical concepts."""
def __init__(self, umls_path: str, custom_definitions_path: str):
"""
Initialize knowledge graph.
Args:
umls_path: Path to UMLS data
custom_definitions_path: Path to custom anatomical definitions
"""
self.concepts = {} # concept_id -> concept_data
self.definitions = {} # concept_id -> definition text
self.relationships = defaultdict(list) # concept_id -> [(rel_type, related_id), ...]
self.concept_to_id = {} # concept_name -> concept_id
self._load_umls(umls_path)
self._load_custom_definitions(custom_definitions_path)
def _load_umls(self, umls_path: str):
"""Load UMLS data."""
try:
with open(umls_path, 'r') as f:
umls_data = json.load(f)
for concept_id, concept_data in umls_data['concepts'].items():
self.concepts[concept_id] = concept_data
concept_name = concept_data.get('name', '')
self.concept_to_id[concept_name.lower()] = concept_id
self.definitions[concept_id] = concept_data.get('definition', '')
for concept_id, relationships in umls_data.get('relationships', {}).items():
self.relationships[concept_id] = relationships
logger.info(f"Loaded {len(self.concepts)} concepts from UMLS")
except FileNotFoundError:
logger.warning(f"UMLS file not found at {umls_path}")
def _load_custom_definitions(self, path: str):
"""Load custom anatomical definitions."""
try:
with open(path, 'r') as f:
custom_defs = json.load(f)
for concept_name, definition in custom_defs['definitions'].items():
concept_id = self.concept_to_id.get(concept_name.lower())
if concept_id:
self.definitions[concept_id] = definition
else:
# Create new concept
concept_id = f"CUSTOM_{len(self.concepts)}"
self.concepts[concept_id] = {'name': concept_name}
self.definitions[concept_id] = definition
self.concept_to_id[concept_name.lower()] = concept_id
logger.info(f"Loaded custom definitions")
except FileNotFoundError:
logger.warning(f"Custom definitions file not found at {path}")
def get_concept_definition(self, concept_name: str) -> str:
"""Get definition for a concept."""
concept_id = self.concept_to_id.get(concept_name.lower())
if concept_id:
return self.definitions.get(concept_id, "")
return ""
def get_related_concepts(self, concept_name: str,
rel_type: Optional[str] = None) -> List[str]:
"""Get related concepts."""
concept_id = self.concept_to_id.get(concept_name.lower())
if not concept_id:
return []
related = []
for rel_type_stored, related_id in self.relationships.get(concept_id, []):
if rel_type is None or rel_type == rel_type_stored:
related_concept = self.concepts.get(related_id, {})
related.append(related_concept.get('name', ''))
return related
def get_concept_hierarchy(self, concept_name: str,
max_depth: int = 3) -> Dict:
"""Get concept hierarchy (up and down)."""
concept_id = self.concept_to_id.get(concept_name.lower())
if not concept_id:
return {}
hierarchy = {
'concept': concept_name,
'definition': self.definitions.get(concept_id, ''),
'parents': self._get_parents(concept_id, max_depth),
'children': self._get_children(concept_id, max_depth),
}
return hierarchy
def _get_parents(self, concept_id: str, depth: int = 2) -> List[Dict]:
"""Get parent concepts recursively."""
if depth == 0:
return []
parents = []
for rel_type, related_id in self.relationships.get(concept_id, []):
if rel_type in ['is-a', 'parent_of']:
related_concept = self.concepts.get(related_id, {})
parents.append({
'concept': related_concept.get('name', ''),
'definition': self.definitions.get(related_id, ''),
'parents': self._get_parents(related_id, depth - 1)
})
return parents
def _get_children(self, concept_id: str, depth: int = 2) -> List[Dict]:
"""Get child concepts recursively."""
if depth == 0:
return []
children = []
for other_id in self.concepts:
for rel_type, related_id in self.relationships.get(other_id, []):
if related_id == concept_id and rel_type in ['is-a', 'parent_of']:
related_concept = self.concepts.get(other_id, {})
children.append({
'concept': related_concept.get('name', ''),
'definition': self.definitions.get(other_id, ''),
'children': self._get_children(other_id, depth - 1)
})
return children
class AtlasEmbedding:
"""Extract visual features from atlas segmentations."""
def __init__(self):
self.atlas_cache = {}
def extract_roi_features(self, image: np.ndarray,
mask: np.ndarray,
num_scales: int = 5) -> np.ndarray:
"""
Extract multi-scale ROI features.
Args:
image: Image array
mask: Binary mask
num_scales: Number of scales to extract
Returns:
Feature vector
"""
features = []
# Get bounding box
positions = np.where(mask > 0)
if len(positions[0]) == 0:
return np.zeros(256)
# Extract multi-scale features
for scale in range(num_scales):
scale_factor = 2 ** scale
pooled = self._adaptive_pool(image, mask, scale_factor)
features.append(pooled)
# Concatenate features
features = np.concatenate(features)
return features.astype(np.float32)
def _adaptive_pool(self, image: np.ndarray,
mask: np.ndarray,
pool_size: int) -> np.ndarray:
"""Adaptive pooling."""
masked_image = image * mask
pool = np.mean(masked_image[mask > 0])
return np.array([pool])# data/dataset.py
import torch
from torch.utils.data import Dataset
import numpy as np
import os
import json
from typing import Dict, Tuple, List, Optional
import logging
logger = logging.getLogger(__name__)
class SAT_Dataset(Dataset):
"""SAT Dataset for 3D medical image segmentation."""
def __init__(self,
dataset_dir: str,
split: str = 'train',
config: Dict = None,
preprocessor = None,
augmentor = None,
knowledge_graph = None):
"""
Initialize dataset.
Args:
dataset_dir: Path to dataset directory
split: 'train', 'val', or 'test'
config: Configuration dictionary
preprocessor: Preprocessing pipeline
augmentor: Data augmentation pipeline
knowledge_graph: Anatomical knowledge graph
"""
self.dataset_dir = dataset_dir
self.split = split
self.config = config or {}
self.preprocessor = preprocessor
self.augmentor = augmentor
self.knowledge_graph = knowledge_graph
self.samples = []
self._load_samples()
logger.info(f"Loaded {len(self.samples)} samples for {split} split")
def _load_samples(self):
"""Load sample metadata."""
split_file = os.path.join(self.dataset_dir, f'{self.split}_split.json')
try:
with open(split_file, 'r') as f:
split_data = json.load(f)
for sample_id, sample_info in split_data.items():
self.samples.append({
'id': sample_id,
'image_path': sample_info['image_path'],
'label_path': sample_info['label_path'],
'modality': sample_info.get('modality', 'CT'),
'anatomy_labels': sample_info.get('anatomy_labels', []),
'spacing': sample_info.get('spacing', [1.0, 1.0, 3.0])
})
except FileNotFoundError:
logger.error(f"Split file not found: {split_file}")
raise
def __len__(self) -> int:
return len(self.samples)
def __getitem__(self, idx: int) -> Dict:
"""Get dataset item."""
sample = self.samples[idx]
# Load image and label
image = np.load(sample['image_path']).astype(np.float32)
label = np.load(sample['label_path']).astype(np.int32)
# Preprocess
if self.preprocessor:
image = self.preprocessor(
image,
sample['modality'],
sample['spacing']
)
# Augmentation (train only)
if self.split == 'train' and self.augmentor:
image, label = self.augmentor(image, label)
# Convert to tensor
image = torch.from_numpy(image).unsqueeze(0) # Add channel dimension
label = torch.from_numpy(label).long()
# Get text prompts and definitions
text_prompts = sample['anatomy_labels']
definitions = []
if self.knowledge_graph:
definitions = [
self.knowledge_graph.get_concept_definition(prompt)
for prompt in text_prompts
]
return {
'image': image,
'label': label,
'text_prompts': text_prompts,
'definitions': definitions,
'modality': sample['modality'],
'sample_id': sample['id']
}
class DataCollator:
"""Custom collate function for batching."""
def __call__(self, batch: List[Dict]) -> Dict:
"""Collate batch."""
images = torch.stack([item['image'] for item in batch])
labels = torch.stack([item['label'] for item in batch])
text_prompts = [item['text_prompts'] for item in batch]
definitions = [item['definitions'] for item in batch]
modalities = [item['modality'] for item in batch]
sample_ids = [item['sample_id'] for item in batch]
return {
'images': images,
'labels': labels,
'text_prompts': text_prompts,
'definitions': definitions,
'modalities': modalities,
'sample_ids': sample_ids
}# 3. Model Architecture
import torch
import torch.nn as nn
import torch.nn.functional as F
from transformers import AutoTokenizer, AutoModel
import logging
logger = logging.getLogger(__name__)
class TextEncoder(nn.Module):
"""Knowledge-enhanced text encoder."""
def __init__(self, config: Dict):
super().__init__()
self.config = config
self.hidden_dim = config['hidden_dim']
self.embedding_dim = config.get('embedding_dim', 768)
# Load pretrained BioBERT
model_name = config.get('pretrained_model', 'dmis-lab/biobert-base-cased')
self.tokenizer = AutoTokenizer.from_pretrained(model_name)
self.encoder = AutoModel.from_pretrained(model_name)
# Projection layer
self.projection = nn.Linear(self.hidden_dim, self.embedding_dim)
self.norm = nn.LayerNorm(self.embedding_dim)
# Freeze layers if specified
num_freeze = config.get('freeze_layers', 0)
if num_freeze > 0:
for i, param in enumerate(self.encoder.parameters()):
if i < num_freeze * len(list(self.encoder.parameters())) // config['num_layers']:
param.requires_grad = False
def forward(self, texts: List[str]) -> torch.Tensor:
"""
Encode text prompts.
Args:
texts: List of text prompts
Returns:
Encoded text embeddings [batch_size, embedding_dim]
"""
# Tokenize
encoded = self.tokenizer(
texts,
padding=True,
truncation=True,
max_length=512,
return_tensors='pt'
)
# Encode with BERT
outputs = self.encoder(
input_ids=encoded['input_ids'],
attention_mask=encoded['attention_mask'],
return_dict=True
)
# Use [CLS] token representation
cls_output = outputs.last_hidden_state[:, 0, :] # [batch_size, hidden_dim]
# Project to embedding dimension
embeddings = self.projection(cls_output)
embeddings = self.norm(embeddings)
return embeddings
class VisualEncoder(nn.Module):
"""3D U-Net visual encoder."""
def __init__(self, config: Dict):
super().__init__()
self.config = config
self.in_channels = config.get('in_channels', 1)
self.num_features = config.get('num_features', 64)
self.depth = config.get('depth', 5)
self.encoder_blocks = nn.ModuleList()
self.pool = nn.MaxPool3d(2, 2)
in_channels = self.in_channels
for i in range(self.depth):
out_channels = self.num_features * (2 ** i)
self.encoder_blocks.append(
ConvBlock3D(in_channels, out_channels)
)
in_channels = out_channels
def forward(self, x: torch.Tensor) -> List[torch.Tensor]:
"""
Encode image.
Args:
x: Input image [batch_size, 1, D, H, W]
Returns:
List of multi-scale features
"""
features = []
for i, block in enumerate(self.encoder_blocks):
x = block(x)
features.append(x)
if i < len(self.encoder_blocks) - 1:
x = self.pool(x)
return features
class ConvBlock3D(nn.Module):
"""3D Convolution block."""
def __init__(self, in_channels: int, out_channels: int):
super().__init__()
self.conv1 = nn.Conv3d(in_channels, out_channels, 3, padding=1)
self.bn1 = nn.BatchNorm3d(out_channels)
self.conv2 = nn.Conv3d(out_channels, out_channels, 3, padding=1)
self.bn2 = nn.BatchNorm3d(out_channels)
self.relu = nn.ReLU(inplace=True)
def forward(self, x: torch.Tensor) -> torch.Tensor:
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.conv2(x)
x = self.bn2(x)
x = self.relu(x)
return x
import torch
import torch.nn as nn
import torch.nn.functional as F
from typing import List, Dict
class QueryDecoder(nn.Module):
"""Transformer-based query decoder."""
def __init__(self, config: Dict):
super().__init__()
self.config = config
self.hidden_dim = config.get('hidden_dim', 512)
self.num_heads = config.get('num_heads', 8)
self.num_layers = config.get('num_layers', 6)
self.ffn_dim = config.get('ffn_dim', 2048)
self.dropout = config.get('dropout', 0.1)
self.transformer_decoder = nn.TransformerDecoder(
nn.TransformerDecoderLayer(
d_model=self.hidden_dim,
nhead=self.num_heads,
dim_feedforward=self.ffn_dim,
dropout=self.dropout,
batch_first=True
),
num_layers=self.num_layers
)
def forward(self,
text_embedding: torch.Tensor,
visual_features: List[torch.Tensor]) -> torch.Tensor:
"""
Decode query with visual context.
Args:
text_embedding: Text embedding [batch_size, embedding_dim]
visual_features: Multi-scale visual features
Returns:
Updated query embedding [batch_size, embedding_dim]
"""
# Prepare visual key-value
visual_kv = []
for feature in visual_features:
# Flatten spatial dimensions
batch_size, channels, *spatial = feature.shape
feature_flat = feature.reshape(batch_size, channels, -1).permute(0, 2, 1)
visual_kv.append(feature_flat)
# Concatenate visual features
visual_kv = torch.cat(visual_kv, dim=1) # [batch_size, total_tokens, channels]
# Use transformer decoder
# text_embedding as query, visual_features as key and value
text_embedding = text_embedding.unsqueeze(1) # [batch_size, 1, embedding_dim]
query = self.transformer_decoder(
text_embedding,
visual_kv,
memory_key_padding_mask=None
)
return query.squeeze(1) # [batch_size, embedding_dim]
class VisualDecoder(nn.Module):
"""3D U-Net visual decoder."""
def __init__(self, config: Dict):
super().__init__()
self.config = config
self.num_features = config.get('num_features', 64)
self.depth = config.get('depth', 5)
self.out_channels = config.get('out_channels', 256)
self.decoder_blocks = nn.ModuleList()
self.upsample = nn.Upsample(scale_factor=2, mode='trilinear', align_corners=True)
for i in range(self.depth - 1, 0, -1):
in_channels = self.num_features * (2 ** i)
out_channels = self.num_features * (2 ** (i - 1))
self.decoder_blocks.append(
ConvBlock3D(in_channels + out_channels, out_channels)
)
# Final dense feature layer
self.final_conv = nn.Sequential(
nn.Conv3d(self.num_features, self.out_channels, 1),
nn.BatchNorm3d(self.out_channels),
nn.ReLU(inplace=True)
)
def forward(self, encoder_features: List[torch.Tensor]) -> torch.Tensor:
"""
Decode features.
Args:
encoder_features: List of encoder features from coarse to fine
Returns:
Dense feature map [batch_size, out_channels, D, H, W]
"""
x = encoder_features[-1]
for i, decoder_block in enumerate(self.decoder_blocks):
x = self.upsample(x)
skip = encoder_features[-(i + 2)]
x = torch.cat([x, skip], dim=1)
x = decoder_block(x)
x = self.final_conv(x)
return x# models/sat_model.py
import torch
import torch.nn as nn
import torch.nn.functional as F
from typing import Dict, List, Tuple, Optional
import logging
from .encoders import TextEncoder, VisualEncoder
from .decoders import QueryDecoder, VisualDecoder
logger = logging.getLogger(__name__)
class SAT(nn.Module):
"""Segment Anything with Text - Main Model."""
def __init__(self, config: Dict):
super().__init__()
self.config = config
# Initialize encoders
self.text_encoder = TextEncoder(config['model']['text_encoder'])
self.visual_encoder = VisualEncoder(config['model']['visual_encoder'])
# Initialize decoders
self.query_decoder = QueryDecoder(config['model']['query_decoder'])
self.visual_decoder = VisualDecoder(config['model']['visual_decoder'])
# Mask generator
self.mask_generator = nn.Sequential(
nn.Linear(config['model']['visual_decoder']['out_channels'], 256),
nn.ReLU(),
nn.Linear(256, 1),
nn.Sigmoid()
)
# Knowledge injection parameters
self.use_knowledge = config['model']['knowledge_injection']['enabled']
self.contrastive_temp = config['model']['knowledge_injection']['contrastive_temp']
def forward(self,
images: torch.Tensor,
text_prompts: List[str],
definitions: Optional[List[str]] = None,
labels: Optional[torch.Tensor] = None) -> Dict:
"""
Forward pass.
Args:
images: Image batch [batch_size, 1, D, H, W]
text_prompts: Text prompts for segmentation
definitions: Anatomical definitions (for knowledge injection)
labels: Ground truth labels (optional)
Returns:
Dictionary with predictions and losses
"""
batch_size = images.shape[0]
device = images.device
# Encode text
text_embeddings = self.text_encoder(text_prompts) # [batch_size, embedding_dim]
# Encode image
visual_features = self.visual_encoder(images) # List of features
# Decode query with visual context
query_features = self.query_decoder(
text_embeddings,
visual_features
) # [batch_size, embedding_dim]
# Decode visual features
dense_features = self.visual_decoder(visual_features) # [batch_size, out_channels, D, H, W]
# Generate segmentation
out_channels = dense_features.shape[1]
dense_features_reshaped = dense_features.permute(0, 2, 3, 4, 1).reshape(-1, out_channels)
query_features_repeated = query_features.unsqueeze(-1).unsqueeze(-1).unsqueeze(-1)
query_features_repeated = query_features_repeated.repeat(
1, 1, dense_features.shape[2], dense_features.shape[3], dense_features.shape[4]
)
query_features_reshaped = query_features_repeated.permute(0, 2, 3, 4, 1).reshape(-1, out_channels)
# Element-wise multiplication
dot_product = (dense_features_reshaped * query_features_reshaped).sum(dim=1)
mask = torch.sigmoid(dot_product)
# Reshape to spatial dimensions
spatial_shape = dense_features.shape[2:]
mask = mask.reshape(batch_size, *spatial_shape)
output = {
'mask': mask,
'text_embedding': text_embeddings,
'query_feature': query_features,
'dense_features': dense_features
}
# Calculate losses if labels provided
if labels is not None:
losses = self._calculate_losses(
mask, labels, text_embeddings, definitions
)
output['losses'] = losses
return output
def _calculate_losses(self,
mask: torch.Tensor,
labels: torch.Tensor,
text_embeddings: torch.Tensor,
definitions: Optional[List[str]]) -> Dict:
"""Calculate segmentation and knowledge losses."""
losses = {}
# Segmentation loss
mask_flat = mask.flatten()
labels_flat = labels.flatten().float()
bce_loss = F.binary_cross_entropy(mask_flat, labels_flat)
# Dice loss
intersection = (mask_flat * labels_flat).sum()
union = mask_flat.sum() + labels_flat.sum()
dice_loss = 1 - (2 * intersection + 1e-6) / (union + 1e-6)
segmentation_loss = bce_loss + dice_loss
losses['segmentation'] = segmentation_loss
# Knowledge loss (if using knowledge injection)
if self.use_knowledge and definitions:
knowledge_loss = self._calculate_knowledge_loss(
text_embeddings, definitions
)
losses['knowledge'] = knowledge_loss
return losses
def _calculate_knowledge_loss(self,
text_embeddings: torch.Tensor,
definitions: List[str]) -> torch.Tensor:
"""Calculate contrastive knowledge loss."""
# Encode definitions
def_embeddings = self.text_encoder(definitions)
# Normalize embeddings
text_norm = F.normalize(text_embeddings, dim=-1)
def_norm = F.normalize(def_embeddings, dim=-1)
# Compute similarity matrix
similarity = torch.matmul(text_norm, def_norm.T) # [batch_size, batch_size]
similarity = similarity / self.contrastive_temp
# Contrastive loss
batch_size = text_embeddings.shape[0]
labels = torch.arange(batch_size, device=text_embeddings.device)
loss_forward = F.cross_entropy(similarity, labels)
loss_backward = F.cross_entropy(similarity.T, labels)
loss = (loss_forward + loss_backward) / 2
return loss
class SAT_Nano(SAT):
"""SAT-Nano variant with reduced parameters."""
def __init__(self, config: Dict):
# Reduce model sizes
config['model']['text_encoder']['hidden_dim'] = 512
config['model']['visual_encoder']['num_features'] = 32
config['model']['query_decoder']['hidden_dim'] = 256
config['model']['visual_decoder']['out_channels'] = 128
super().__init__(config)
class SAT_Pro(SAT):
"""SAT-Pro variant with full capacity."""
def __init__(self, config: Dict):
# Use default full sizes
super().__init__(config)# 4. Training Pipeline
import torch
import torch.nn as nn
import torch.nn.functional as F
from typing import Dict
class DiceLoss(nn.Module):
"""Dice Loss for segmentation."""
def __init__(self, smooth: float = 1e-6):
super().__init__()
self.smooth = smooth
def forward(self, pred: torch.Tensor, target: torch.Tensor) -> torch.Tensor:
pred_flat = pred.flatten()
target_flat = target.flatten().float()
intersection = (pred_flat * target_flat).sum()
union = pred_flat.sum() + target_flat.sum()
dice = (2 * intersection + self.smooth) / (union + self.smooth)
return 1 - dice
class CombinedSegmentationLoss(nn.Module):
"""Combined BCE + Dice loss."""
def __init__(self, bce_weight: float = 0.5, dice_weight: float = 0.5):
super().__init__()
self.bce_weight = bce_weight
self.dice_weight = dice_weight
self.bce_loss = nn.BCELoss()
self.dice_loss = DiceLoss()
def forward(self, pred: torch.Tensor, target: torch.Tensor) -> torch.Tensor:
bce = self.bce_loss(pred, target.float())
dice = self.dice_loss(pred, target)
return self.bce_weight * bce + self.dice_weight * dice
class ContrastiveKnowledgeLoss(nn.Module):
"""Contrastive learning loss for knowledge injection."""
def __init__(self, temperature: float = 0.07):
super().__init__()
self.temperature = temperature
def forward(self,
text_embeddings: torch.Tensor,
definition_embeddings: torch.Tensor) -> torch.Tensor:
"""
Contrastive loss between text and definition embeddings.
"""
# Normalize
text_norm = F.normalize(text_embeddings, dim=-1)
def_norm = F.normalize(definition_embeddings, dim=-1)
# Similarity matrix
similarity = torch.matmul(text_norm, def_norm.T) / self.temperature
batch_size = text_embeddings.shape[0]
labels = torch.arange(batch_size, device=text_embeddings.device)
loss_forward = F.cross_entropy(similarity, labels)
loss_backward = F.cross_entropy(similarity.T, labels)
return (loss_forward + loss_backward) / 2
class CombinedLoss(nn.Module):
"""Combined segmentation and knowledge loss."""
def __init__(self, config: Dict):
super().__init__()
self.segmentation_loss = CombinedSegmentationLoss()
self.knowledge_loss = ContrastiveKnowledgeLoss(
temperature=config['model']['knowledge_injection']['contrastive_temp']
)
self.seg_weight = config['training']['loss_weights']['segmentation']
self.know_weight = config['training']['loss_weights']['knowledge']
self.use_knowledge = config['model']['knowledge_injection']['enabled']
def forward(self,
predictions: Dict,
targets: torch.Tensor,
definition_embeddings: torch.Tensor = None) -> torch.Tensor:
"""Compute combined loss."""
seg_loss = self.segmentation_loss(predictions['mask'], targets)
total_loss = self.seg_weight * seg_loss
if self.use_knowledge and definition_embeddings is not None:
know_loss = self.knowledge_loss(
predictions['text_embedding'],
definition_embeddings
)
total_loss += self.know_weight * know_loss
return total_loss
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
import logging
from pathlib import Path
from typing import Dict, Optional
import json
from ..models.sat_model import SAT, SAT_Pro, SAT_Nano
from .losses import CombinedLoss
from ..evaluation.metrics import DiceScore, NormalizedSurfaceDistance
logger = logging.getLogger(__name__)
class Trainer:
"""Training framework for SAT model."""
def __init__(self, config: Dict, device: str = 'cuda'):
self.config = config
self.device = device
# Initialize model
model_variant = config['model'].get('variant', 'pro')
if model_variant == 'nano':
self.model = SAT_Nano(config).to(device)
else:
self.model = SAT_Pro(config).to(device)
# Initialize loss
self.criterion = CombinedLoss(config)
# Initialize optimizer
self._setup_optimizer()
# Initialize scheduler
self._setup_scheduler()
# Metrics
self.dice_metric = DiceScore()
self.nsd_metric = NormalizedSurfaceDistance()
# Checkpointing
self.checkpoint_dir = Path(config['training']['checkpoint_dir'])
self.checkpoint_dir.mkdir(exist_ok=True)
def _setup_optimizer(self):
"""Setup optimizer."""
optimizer_name = self.config['training'].get('optimizer', 'adamw')
lr = self.config['training']['learning_rate']
wd = self.config['training']['weight_decay']
if optimizer_name == 'adamw':
self.optimizer = optim.AdamW(
self.model.parameters(),
lr=lr,
weight_decay=wd
)
elif optimizer_name == 'sgd':
self.optimizer = optim.SGD(
self.model.parameters(),
lr=lr,
weight_decay=wd,
momentum=0.9
)
else:
raise ValueError(f"Unknown optimizer: {optimizer_name}")
def _setup_scheduler(self):
"""Setup learning rate scheduler."""
scheduler_name = self.config['training'].get('scheduler', 'cosine')
num_epochs = self.config['training']['num_epochs']
warmup_epochs = self.config['training']['warmup_epochs']
if scheduler_name == 'cosine':
self.scheduler = optim.lr_scheduler.CosineAnnealingLR(
self.optimizer,
T_max=num_epochs - warmup_epochs
)
elif scheduler_name == 'linear':
self.scheduler = optim.lr_scheduler.LinearLR(
self.optimizer,
total_iters=num_epochs
)
else:
raise ValueError(f"Unknown scheduler: {scheduler_name}")
def train_epoch(self, train_loader: DataLoader) -> Dict:
"""Train one epoch."""
self.model.train()
losses = []
dice_scores = []
for batch_idx, batch in enumerate(train_loader):
images = batch['images'].to(self.device)
labels = batch['labels'].to(self.device)
text_prompts = batch['text_prompts']
definitions = batch['definitions']
# Forward pass
outputs = self.model(images, text_prompts, definitions, labels)
# Calculate loss
loss = self.criterion(outputs, labels)
# Backward pass
self.optimizer.zero_grad()
loss.backward()
torch.nn.utils.clip_grad_norm_(self.model.parameters(), 1.0)
self.optimizer.step()
losses.append(loss.item())
# Calculate metrics
dice = self.dice_metric(outputs['mask'], labels)
dice_scores.append(dice)
if batch_idx % self.config['logging']['log_interval'] == 0:
logger.info(
f"Batch {batch_idx}/{len(train_loader)}: "
f"Loss = {loss.item():.4f}, Dice = {dice:.4f}"
)
return {
'loss': sum(losses) / len(losses),
'dice': sum(dice_scores) / len(dice_scores)
}
def validate(self, val_loader: DataLoader) -> Dict:
"""Validate model."""
self.model.eval()
losses = []
dice_scores = []
nsd_scores = []
with torch.no_grad():
for batch in val_loader:
images = batch['images'].to(self.device)
labels = batch['labels'].to(self.device)
text_prompts = batch['text_prompts']
definitions = batch['definitions']
outputs = self.model(images, text_prompts, definitions)
# Evaluate
dice = self.dice_metric(outputs['mask'], labels)
nsd = self.nsd_metric(outputs['mask'], labels)
dice_scores.append(dice)
nsd_scores.append(nsd)
return {
'dice': sum(dice_scores) / len(dice_scores),
'nsd': sum(nsd_scores) / len(nsd_scores)
}
def train(self, train_loader: DataLoader, val_loader: DataLoader):
"""Full training loop."""
num_epochs = self.config['training']['num_epochs']
best_dice = 0
for epoch in range(num_epochs):
logger.info(f"Epoch {epoch + 1}/{num_epochs}")
# Train
train_metrics = self.train_epoch(train_loader)
logger.info(f"Train - Loss: {train_metrics['loss']:.4f}, "
f"Dice: {train_metrics['dice']:.4f}")
# Validate
if (epoch + 1) % self.config['evaluation']['val_interval'] == 0:
val_metrics = self.validate(val_loader)
logger.info(f"Val - Dice: {val_metrics['dice']:.4f}, "
f"NSD: {val_metrics['nsd']:.4f}")
# Save checkpoint
if val_metrics['dice'] > best_dice:
best_dice = val_metrics['dice']
self._save_checkpoint(epoch, val_metrics)
# Update scheduler
self.scheduler.step()
def _save_checkpoint(self, epoch: int, metrics: Dict):
"""Save model checkpoint."""
checkpoint = {
'epoch': epoch,
'model_state': self.model.state_dict(),
'optimizer_state': self.optimizer.state_dict(),
'metrics': metrics,
'config': self.config
}
path = self.checkpoint_dir / f"checkpoint_epoch_{epoch}.pt"
torch.save(checkpoint, path)
logger.info(f"Saved checkpoint: {path}")
def load_checkpoint(self, path: str):
"""Load model checkpoint."""
checkpoint = torch.load(path, map_location=self.device)
self.model.load_state_dict(checkpoint['model_state'])
self.optimizer.load_state_dict(checkpoint['optimizer_state'])
logger.info(f"Loaded checkpoint from {path}")
return checkpoint['epoch']# 5. Evaluation and Inference
import torch
import torch.nn.functional as F
import numpy as np
from scipy.spatial.distance import directed_hausdorff
from typing import Tuple
class DiceScore:
"""Dice Similarity Coefficient."""
def __init__(self, smooth: float = 1e-6):
self.smooth = smooth
def __call__(self, pred: torch.Tensor, target: torch.Tensor) -> float:
"""Calculate Dice score."""
pred_binary = (pred > 0.5).float()
target_binary = target.float()
intersection = (pred_binary * target_binary).sum()
union = pred_binary.sum() + target_binary.sum()
dice = (2 * intersection + self.smooth) / (union + self.smooth)
return dice.item()
class NormalizedSurfaceDistance:
"""Normalized Surface Distance metric."""
def __init__(self, tau: int = 1):
self.tau = tau
def __call__(self, pred: torch.Tensor, target: torch.Tensor) -> float:
"""Calculate NSD metric."""
pred_np = pred.cpu().numpy()
target_np = target.cpu().numpy()
# Get boundaries
pred_boundary = self._get_boundary(pred_np > 0.5)
target_boundary = self._get_boundary(target_np > 0)
if len(pred_boundary) == 0 or len(target_boundary) == 0:
return 0.0
# Calculate directed Hausdorff distances
dist1 = directed_hausdorff(pred_boundary, target_boundary)[0]
dist2 = directed_hausdorff(target_boundary, pred_boundary)[0]
# Normalized surface distance
denom = max(dist1, dist2)
if denom == 0:
return 1.0
nsd = 1 - min(dist1, dist2) / denom
return nsd
def _get_boundary(self, mask: np.ndarray) -> np.ndarray:
"""Get boundary voxels."""
boundary = np.zeros_like(mask)
for i in range(mask.shape[0]):
for j in range(mask.shape[1]):
for k in range(mask.shape[2]):
if mask[i, j, k]:
# Check if on boundary
neighbors = mask[
max(0, i-1):min(mask.shape[0], i+2),
max(0, j-1):min(mask.shape[1], j+2),
max(0, k-1):min(mask.shape[2], k+2)
]
if not neighbors.all():
boundary[i, j, k] = True
return np.argwhere(boundary)
import torch
import yaml
import logging
import json
from pathlib import Path
from torch.utils.data import DataLoader
from config import load_config
from data.dataset import SAT_Dataset, DataCollator
from data.preprocessing import MedicalImagePreprocessor, DataAugmentation
from data.knowledge_graph import AnatomyKnowledgeGraph
from training.trainer import Trainer
from utils.logger import setup_logging
def main():
"""Main training function."""
# Setup logging
setup_logging()
logger = logging.getLogger(__name__)
# Load configuration
config = load_config('config/config.yaml')
logger.info(f"Loaded configuration")
# Setup device
device = 'cuda' if torch.cuda.is_available() and config['device']['cuda'] else 'cpu'
logger.info(f"Using device: {device}")
# Initialize preprocessor and augmentor
preprocessor = MedicalImagePreprocessor(config['data'])
augmentor = DataAugmentation(config['data'])
# Initialize knowledge graph
kg = AnatomyKnowledgeGraph(
umls_path='data/umls_data.json',
custom_definitions_path='data/anatomical_definitions.json'
)
# Create datasets
logger.info("Creating datasets...")
train_dataset = SAT_Dataset(
dataset_dir=config['data']['dataset_path'],
split='train',
config=config,
preprocessor=preprocessor,
augmentor=augmentor,
knowledge_graph=kg
)
val_dataset = SAT_Dataset(
dataset_dir=config['data']['dataset_path'],
split='val',
config=config,
preprocessor=preprocessor,
augmentor=None,
knowledge_graph=kg
)
# Create dataloaders
collate_fn = DataCollator()
train_loader = DataLoader(
train_dataset,
batch_size=config['data']['batch_size'],
shuffle=True,
num_workers=config['data']['num_workers'],
collate_fn=collate_fn
)
val_loader = DataLoader(
val_dataset,
batch_size=config['data']['batch_size'],
shuffle=False,
num_workers=config['data']['num_workers'],
collate_fn=collate_fn
)
logger.info(f"Train loader: {len(train_loader)} batches")
logger.info(f"Val loader: {len(val_loader)} batches")
# Initialize trainer
trainer = Trainer(config, device=device)
# Train
logger.info("Starting training...")
trainer.train(train_loader, val_loader)
logger.info("Training completed!")
if __name__ == '__main__':
main()Related posts, You May like to read
- 7 Shocking Truths About Knowledge Distillation: The Good, The Bad, and The Breakthrough (SAKD)
- 7 Revolutionary Breakthroughs in Medical Image Translation (And 1 Fatal Flaw That Could Derail Your AI Model)
- TimeDistill: Revolutionizing Time Series Forecasting with Cross-Architecture Knowledge Distillation
- HiPerformer: A New Benchmark in Medical Image Segmentation with Modular Hierarchical Fusion
- GeoSAM2 3D Part Segmentation — Prompt-Controllable, Geometry-Aware Masks for Precision 3D Editing
- Probabilistic Smooth Attention for Deep Multiple Instance Learning in Medical Imaging
- A Knowledge Distillation-Based Approach to Enhance Transparency of Classifier Models
- Towards Trustworthy Breast Tumor Segmentation in Ultrasound Using AI Uncertainty
- Discrete Migratory Bird Optimizer with Deep Transfer Learning for Multi-Retinal Disease Detection

Thank you for your sharing. I am worried that I lack creative ideas. It is your article that makes me full of hope. Thank you. But, I have a question, can you help me? https://accounts.binance.com/register-person?ref=IXBIAFVY