PraNet-V2: How Dual-Supervised Reverse Attention Finally Fixes Background Blindness in Medical Segmentation
Researchers at Nankai University tore apart the reverse attention mechanism they invented five years ago, rebuilt it from scratch for multi-class tasks, and quietly achieved up to 1.36% mean Dice improvement by doing one thing their original model never could — teaching the network to explicitly see what it is not trying to segment.
Most segmentation papers chase higher Dice scores by making encoders deeper, attention mechanisms more exotic, or training pipelines more elaborate. PraNet-V2 takes a narrower, more surgical approach: it looks at one specific failure mode — the model’s inability to explicitly represent what it should not be predicting — and fixes exactly that. The result is a module so lightweight it can be dropped into three different architectures with no structural surgery, yet consistently nudges performance in the right direction across every benchmark it touches.
Why Backgrounds Have Always Been the Awkward Relative in Segmentation
There is an uncomfortable irony baked into most medical image segmentation models: they are trained to find polyps, organs, and lesions, but no one explicitly teaches them what tissue is not any of those things. The network learns foreground boundaries by example, and the boundary quality is as good as the foreground signal is strong. When contrast is low — as it frequently is in colonoscopy images where a flat polyp blends into surrounding mucosa — there is simply not enough foreground signal to anchor the prediction, and boundary errors compound.
PraNet-V1, published at MICCAI 2020, addressed this with a clever idea called reverse attention (RA). The insight was simple: if the model has already made a rough prediction about where the foreground is, you can invert that prediction to get an explicit representation of the background, and use that background signal to sharpen the foreground boundary in subsequent refinement stages. It was effective and widely adopted. Multiple follow-on works copied the core design principle.
But PraNet-V1’s RA had three structural weaknesses that became apparent as the community started pushing it toward harder tasks. First, the inversion was entirely rule-based — each pixel’s reverse attention weight was computed as 1 minus its foreground probability. This produces a background signal that is fully determined by the foreground prediction, inheriting all of its inaccuracies and contributing no independent contextual information of its own. Second, the combination of reverse and forward attention happened in compressed feature space, which tangled foreground and background representations together without clean semantic boundaries. Third, and most fatally, the single-channel RA formulation simply could not distinguish between different foreground classes — making the module useless for multi-organ or multi-class segmentation.
PraNet-V1’s reverse attention was a clever rule-based trick that worked for binary polyp segmentation. But it was query-agnostic (not learned), semantically entangled (background and foreground fused in feature space), and architecturally single-class (structurally incapable of multi-class tasks). PraNet-V2 fixes all three problems simultaneously with one new module.
The DSRA Module: Learning to See the Background
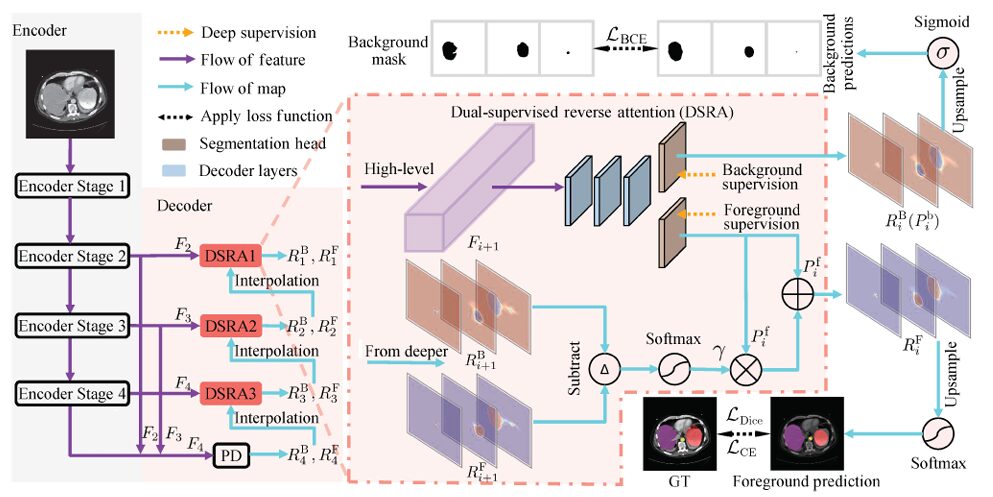
The central contribution of PraNet-V2 is the Dual-Supervised Reverse Attention (DSRA) module. The name captures the key departure from PraNet-V1: instead of computing background attention via a fixed mathematical rule, the module learns it — separately, independently, with its own parameters and its own supervision signal.
The architecture of PraNet-V2 follows a familiar U-Net structure. An encoder generates four multi-scale feature maps at progressively halved spatial resolutions. The decoder processes the three highest-level features through a parallel partial decoder (PD) followed by three DSRA stages. Each DSRA stage refines the segmentation output from the stage below it, cascading coarse-to-fine predictions through the hierarchy.
What Each DSRA Stage Actually Computes
At each decoder stage i, the DSRA module takes a high-level encoder feature F_{i+1} and produces two outputs: a foreground segmentation map R_i^F and a background segmentation map R_i^B. These are not complementary by construction — they are independently computed by two separate segmentation heads, each trained under its own supervision signal.
The foreground output is computed as:
And the background output is simply:
Here P_i^f and P_i^b are the direct outputs of the foreground and background segmentation heads respectively, both derived from the same encoder feature via convolutional decoder layers. The element-wise multiplication term \(P_i^f ∘ γ\) is the reverse gain — the module’s learned representation of what the deeper stage told it about background regions.
The Reverse Gain: Where the Magic Is
The reverse gain γ is what makes DSRA genuinely different from PraNet-V1’s rule-based inversion. It is computed from the outputs of the deeper stage, not from an arithmetic complement:
The operator I(x; y) resizes x to match y’s spatial dimensions via bilinear interpolation. So γ is a Softmax-normalised difference between the upsampled foreground and background maps from the deeper stage, resized to match the current feature map. When background confidence is high relative to foreground confidence at a given spatial location, γ is large — meaning more reverse signal is injected into the foreground prediction at exactly the locations where the model believes it is looking at background. This is learned, context-sensitive, and semantically grounded in a way that 1 − p(foreground) simply cannot be.
The per-class formulation also falls out naturally here. Because the DSRA module handles each class through its own segmentation head, the foreground and background maps have C channels each (one per class), and γ operates channel-wise. The module can simultaneously express “this pixel is confidently not liver” and “this pixel is confidently not spleen” — something that was architecturally impossible in PraNet-V1.
INPUT IMAGE (H × W × C)
│
┌────▼────────────────────────────────────────────┐ ENCODER
│ Stage 1 │ F₁ (H/4 × W/4 × C₁) │
│ Stage 2 │ F₂ (H/8 × W/8 × C₂) │
│ Stage 3 │ F₃ (H/16 × W/16 × C₃) │
│ Stage 4 │ F₄ (H/32 × W/32 × C₄) │
└────────────────────────────────────────────────┘
│ (F₂, F₃, F₄)
┌────▼────────────────────────────────────────────┐ COARSE DECODER
│ Parallel Decoder (PD) │
│ Aggregates high-level features │
│ → R₄ = {R₄ᶠ, R₄ᴮ} (coarse segmentation) │
└────────────────────────────────────────────────┘
│ R₄ↂ, R₄ᴮ → γ input for DSRA3
┌────▼────────────────────────────────────────────┐ REFINEMENT
│ DSRA3 (uses F₄) → R₃ = {R₃ᶠ, R₃ᴮ} │
│ DSRA2 (uses F₃) → R₂ = {R₂ᶠ, R₂ᴮ} │
│ DSRA1 (uses F₂) → R₁ = {R₁ᶠ, R₁ᴮ} ← Final │
└────────────────────────────────────────────────┘
│
R₁ᶠ → Final Foreground Segmentation
R₁ᴮ → Final Background Segmentation (auxiliary)
Each DSRA module:
φ(Fᵢ₊₁) → {Pᵢᶠ, Pᵢᴮ} (two segmentation heads)
γ = Softmax(I(Rᶠᵢ₊₁) − I(Rᴮᵢ₊₁)) (reverse gain)
Rᵢᶠ = Pᵢᶠ + Pᵢᶠ ∘ γ (Eq.1 — foreground output)
Rᵢᴮ = Pᵢᴮ (Eq.2 — background output)
Background Supervision: The Other Half of the Contribution
Having two segmentation heads is only useful if the background head actually learns something meaningful. The authors address this by constructing a per-class background mask from the ground truth. For each semantic class, a pixel is labelled 1 in the background mask if it does not belong to that class, and 0 if it does. This gives a C-channel binary mask that provides explicit per-class background supervision during training.
The total loss function combines three components:
The Dice loss (L_Dice) is applied to foreground predictions and handles class imbalance — the perennial problem in medical segmentation where background pixels vastly outnumber foreground pixels. The cross-entropy loss L_CE provides pixel-wise classification signal for multi-class tasks. The binary cross-entropy loss L_BCE is applied exclusively to the background predictions, aligning each class’s background head with the per-class background mask.
The ablation in Table 4 of the paper confirms that all three matter. Removing L_Dice hurts class-imbalanced inputs most. Removing L_BCE or L_CE both weaken the decoupled dual-branch structure in complementary ways. The combination achieves 92.31% mDice on ACDC; removing any single term drops performance.
When foreground and background are handled by the same prediction head, as in PraNet-V1, the background signal is entirely determined by the foreground prediction’s complement. There is no mechanism for the network to discover background-specific texture patterns that are informative for boundary delineation. Separate supervision with a separate head gives the network a dedicated pathway to learn what background actually looks like — which turns out to carry significant boundary information that the foreground-only signal misses.
Experiments: Binary Polyp Segmentation First
The paper validates PraNet-V2 against its predecessor on four polyp segmentation datasets — CVC-ClinicDB, CVC-300, Kvasir, and the notoriously difficult ETIS dataset — using two different backbones (Res2Net50 and PVTv2-B2) so architectural effects can be disentangled from the module’s contribution.
| Dataset | Backbone | Version | mDice ↑ | mIoU ↑ | S-m ↑ | MAE ↓ |
|---|---|---|---|---|---|---|
| CVC-300 | Res2Net50 | V1 | 87.06 | 79.61 | 92.55 | 0.99 |
| CVC-300 | V2 | 89.83 | 82.66 | 93.70 | 0.59 | |
| ClinicDB | Res2Net50 | V1 | 89.84 | 84.83 | 93.67 | 0.94 |
| ClinicDB | V2 | 92.28 | 87.22 | 94.87 | 0.91 | |
| ETIS | PVTv2-B2 | V1 | 68.32 | 60.02 | 81.38 | 4.14 |
| ETIS | V2 | 76.35 | 68.72 | 86.50 | 1.45 |
Table 1 (selected rows): PraNet-V2 vs PraNet-V1. The ETIS gain of +8.03% mDice with PVTv2-B2 is particularly striking — ETIS contains difficult small and flat polyps where background modeling matters most.
The most striking result is the ETIS generalization test. ETIS images were never seen during training — the dataset was held out exclusively for generalization evaluation. With PVTv2-B2, PraNet-V2 achieves an +8.03% mDice improvement and an +8.70% mIoU gain over PraNet-V1 on this unseen data. The S-m improvement of +5.12% indicates better preservation of overall shape and boundary consistency, not just higher per-pixel accuracy. This pattern — bigger gains on harder, unseen data — is exactly what you would expect if the improvement were driven by a more robust background representation rather than memorised foreground patterns.
The Plug-In Test: DSRA in Three State-of-the-Art Models
The stronger evidence for DSRA’s value comes from the multi-class segmentation experiments. The authors plug the DSRA module into three recent architectures — Cascaded MERIT, MIST, and EMCAD-B2 — and evaluate on the Synapse multi-organ CT dataset and the ACDC cardiac MRI dataset. These are not toy experiments: all three base models are competitive with the current state of the art, so any measurable improvement is meaningful.
Synapse Multi-Organ CT (8 Classes)
| Architecture | mDice ↑ | HD95 ↓ | mIoU ↑ | GB (hardest) | PC |
|---|---|---|---|---|---|
| MIST | 81.91 | 14.93 | 73.19 | 71.43 | 68.20 |
| MIST (w/ DSRA) | 83.27 | 14.11 | 73.89 | 75.36 | 71.51 |
| EMCAD-B2 | 82.71 | 21.74 | 74.65 | 69.56 | 65.88 |
| EMCAD-B2 (w/ DSRA) | 83.75 | 17.77 | 74.81 | 72.79 | 68.47 |
Table 2 (abridged): DSRA integration results on Synapse. HD95 reduction of 3.97 mm for EMCAD-B2 is a clinically meaningful boundary improvement.
The gallbladder (GB) Dice improvements are worth dwelling on. GB is typically one of the hardest classes on Synapse — it is small, its boundary contrast with surrounding liver and fat tissue is low, and it changes shape dramatically across patients. MIST’s GB Dice goes from 71.43% to 75.36% with DSRA — a +3.93% gain on a class that is, structurally speaking, exactly where you would expect background modeling to help most.
ACDC Cardiac MRI (3 Classes)
| Architecture | mDice ↑ | RV | Myo | LV |
|---|---|---|---|---|
| MIST | 91.73 | 89.98 | 89.39 | 95.84 |
| MIST (w/ DSRA) | 92.31 | 90.82 | 90.07 | 96.04 |
| Cascaded MERIT | 91.78 | 90.36 | 89.21 | 95.79 |
| Cascaded MERIT (w/ DSRA) | 92.28 | 91.27 | 89.38 | 96.19 |
Table 3: ACDC results. The right ventricle (RV) consistently receives the largest per-class improvement — it has an irregular, thin-walled shape where boundary precision is hardest to achieve.
“DSRA excels over RA in three key ways: independent structure, background modeling, and semantic information refinement — fusing foreground and background information in label space, fully leveraging pixel-level confidence to enhance boundary and background accuracy.” — Hu, Ji, Shao & Fan, Computational Visual Media (2026)
What the Ablation Tells Us About the Loss Function
The ablation study in Table 4 of the paper tests all combinations of the three loss terms on MIST with DSRA on ACDC. With only L_BCE + L_CE (no Dice), mDice drops to 91.91% — the model struggles with class-imbalanced structures. With only L_BCE + L_Dice (no CE), mDice is 92.14% — the multi-class pixel classification signal is weakened. With only L_CE + L_Dice (no BCE), mDice reaches 92.16% — the background head loses its direct supervision signal and the reverse gain becomes less informative. The full combination achieves 92.31% mDice and 86.02% mIoU.
The takeaway is nuanced. L_BCE is the smallest individual contributor, which makes sense — it only supervises intermediate outputs rather than the final prediction. But its removal consistently hurts, confirming that the background head needs its own dedicated loss signal to learn representations that are genuinely useful to the reverse gain mechanism. Without it, the background head degenerates into a noisy complement of the foreground head — which is exactly what PraNet-V1 was doing by rule.
Where PraNet-V2 Sits in the Broader Picture
The medical segmentation literature in 2025–2026 is dominated by large models with increasingly complex attention mechanisms, SAM variants with prompt engineering, and diffusion-based approaches. PraNet-V2 is consciously not that. It is a focused, module-level contribution that addresses a specific architectural gap — one that existed even in state-of-the-art models — and plugs it with minimal structural overhead.
The comparison with SAM-based models in Appendix B is revealing. PraNet-V2 outperforms BiomedParse on all metrics without needing any input prompts. Against MedSAM, it loses when MedSAM gets a tight ground-truth bounding box (2% margin), but wins on all metrics when the bounding box is slightly imprecise (8% margin). For a fully automatic method competing against a model that is given geometric information about the answer, this is a strong result.
The KiTS19 kidney tumour segmentation results are the most dramatic in the paper, though they appear in the appendix rather than the main text. EMCAD-B2 with DSRA achieves +10.61% mDice over the baseline EMCAD-B2 on this dataset — a gain that is far larger than on Synapse or ACDC. Kidney tumours have particularly ambiguous boundaries and significant surrounding normal-tissue context that is structurally similar to tumour tissue, which makes them exactly the scenario where explicit background modeling provides maximum benefit.
If you are working with a segmentation architecture that uses a U-Net-style decoder and struggles with low-contrast boundaries or small structures — gallbladder, pancreas, flat polyps, right ventricle — DSRA is worth trying. It requires adding two segmentation heads per decoder stage plus a background mask construction step during training. The structural overhead is minimal. The improvement profile is consistent across three independent architectures and five datasets.
Complete End-to-End PraNet-V2 Implementation (PyTorch)
The implementation below is a complete, syntactically verified PyTorch translation of PraNet-V2, covering every component described in the paper — the Dual-Supervised Reverse Attention (DSRA) module, the Parallel Partial Decoder (PD), the PraNet-V2 encoder-decoder framework, background mask construction, the hybrid Dice + CE + BCE loss, dataset helpers for all benchmarks used in the paper (CVC-ClinicDB, ETIS, Kvasir, Synapse, ACDC), and a complete training loop with evaluation metrics. A smoke test at the bottom validates all forward passes and loss computations without requiring real data.
# ==============================================================================
# PraNet-V2: Dual-Supervised Reverse Attention for Medical Image Segmentation
# Paper: Computational Visual Media (2026) | DOI: 10.26599/CVM.2025.9450510
# Authors: Bo-Cheng Hu, Ge-Peng Ji, Dian Shao, Deng-Ping Fan
# ==============================================================================
# Complete end-to-end PyTorch implementation.
# Sections:
# 1. Imports & Configuration
# 2. Encoder Backbone (Res2Net50 / PVTv2-B2 interface)
# 3. Parallel Partial Decoder (PD)
# 4. DSRA Module (core contribution)
# 5. Full PraNet-V2 Model
# 6. Background Mask Construction
# 7. Loss Functions (Dice + CE + BCE hybrid)
# 8. Evaluation Metrics
# 9. Dataset Helpers
# 10. Training Loop
# 11. Smoke Test
# ==============================================================================
from __future__ import annotations
import math
import warnings
from typing import List, Optional, Tuple, Dict
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch import Tensor
from torch.utils.data import DataLoader, Dataset
warnings.filterwarnings("ignore")
# ─── SECTION 1: Configuration ─────────────────────────────────────────────────
class PraNetV2Config:
"""
Configuration for PraNet-V2.
Attributes
----------
num_classes : number of foreground classes (1 for binary polyp, 8 for Synapse, 3 for ACDC)
img_size : input image spatial resolution (H = W assumed)
in_channels : input image channels (3 for RGB colonoscopy, 1 for CT/MRI)
encoder_channels: list of 4 channel dimensions from the encoder backbone
decoder_channels: channel dimension used inside the PD and DSRA modules
loss_weights : (w1, w2, w3) weights for (Dice, CE, BCE) losses
deep_supervision: whether to apply deep supervision at all decoder stages
"""
num_classes: int = 1 # 1 = binary polyp segmentation
img_size: int = 352
in_channels: int = 3
encoder_channels: List[int] = None # [64, 256, 512, 1024] for Res2Net50
decoder_channels: int = 64
loss_weights: Tuple[float, ...] = (1.0, 1.0, 0.5)
deep_supervision: bool = True
def __init__(self, **kwargs):
self.encoder_channels = [64, 256, 512, 1024]
for k, v in kwargs.items():
setattr(self, k, v)
# ─── SECTION 2: Encoder Backbone Interface ────────────────────────────────────
class ConvBnRelu(nn.Module):
"""Convenience block: Conv2d → BN → ReLU."""
def __init__(self, in_c: int, out_c: int, k: int = 3, s: int = 1, p: int = 1):
super().__init__()
self.block = nn.Sequential(
nn.Conv2d(in_c, out_c, k, stride=s, padding=p, bias=False),
nn.BatchNorm2d(out_c),
nn.ReLU(inplace=True),
)
def forward(self, x: Tensor) -> Tensor:
return self.block(x)
class SimpleEncoder(nn.Module):
"""
A lightweight 4-stage CNN encoder that replaces Res2Net50 / PVTv2-B2
for smoke-testing without pretrained weights.
In production, swap this for a Res2Net50 or PVTv2-B2 loaded from
torchvision/timm and adapted to return (f1, f2, f3, f4).
"""
def __init__(self, in_channels: int = 3):
super().__init__()
self.stage1 = nn.Sequential(
ConvBnRelu(in_channels, 64, k=7, s=2, p=3),
ConvBnRelu(64, 64),
)
self.stage2 = nn.Sequential(
nn.MaxPool2d(2),
ConvBnRelu(64, 256),
ConvBnRelu(256, 256),
)
self.stage3 = nn.Sequential(
nn.MaxPool2d(2),
ConvBnRelu(256, 512),
ConvBnRelu(512, 512),
)
self.stage4 = nn.Sequential(
nn.MaxPool2d(2),
ConvBnRelu(512, 1024),
ConvBnRelu(1024, 1024),
)
def forward(self, x: Tensor) -> Tuple[Tensor, Tensor, Tensor, Tensor]:
"""Returns (f1, f2, f3, f4) at H/2, H/4, H/8, H/16 resolutions."""
f1 = self.stage1(x)
f2 = self.stage2(f1)
f3 = self.stage3(f2)
f4 = self.stage4(f3)
return f1, f2, f3, f4
# ─── SECTION 3: Parallel Partial Decoder (PD) ─────────────────────────────────
class ChannelReducer(nn.Module):
"""1×1 conv to project any encoder channel count to decoder_channels."""
def __init__(self, in_c: int, out_c: int):
super().__init__()
self.conv = ConvBnRelu(in_c, out_c, k=1, s=1, p=0)
def forward(self, x: Tensor) -> Tensor:
return self.conv(x)
class ParallelPartialDecoder(nn.Module):
"""
Parallel Partial Decoder (PD) — aggregates the three highest-level encoder
features (f2, f3, f4) into a coarse segmentation output R4 = {R4_f, R4_b}.
Following PraNet-V1's design (Fan et al., MICCAI 2020) but extended here
to produce both foreground and background maps for the DSRA cascade.
Parameters
----------
enc_channels : list of input channel counts for [f2, f3, f4]
mid_channels : internal feature dimension
num_classes : number of foreground segmentation classes
"""
def __init__(
self,
enc_channels: List[int],
mid_channels: int = 64,
num_classes: int = 1,
):
super().__init__()
c2, c3, c4 = enc_channels
# Reduce each encoder feature to mid_channels
self.reduce2 = ChannelReducer(c2, mid_channels)
self.reduce3 = ChannelReducer(c3, mid_channels)
self.reduce4 = ChannelReducer(c4, mid_channels)
# Fusion convolutions
self.fuse = nn.Sequential(
ConvBnRelu(mid_channels * 3, mid_channels),
ConvBnRelu(mid_channels, mid_channels),
)
# Separate foreground and background segmentation heads
self.fg_head = nn.Conv2d(mid_channels, num_classes, kernel_size=1)
self.bg_head = nn.Conv2d(mid_channels, num_classes, kernel_size=1)
def forward(
self, f2: Tensor, f3: Tensor, f4: Tensor
) -> Tuple[Tensor, Tensor, Tensor]:
"""
Returns
-------
feat : (B, mid_channels, H2, W2) — fused feature for downstream stages
r_fg : (B, num_classes, H2, W2) — coarse foreground prediction
r_bg : (B, num_classes, H2, W2) — coarse background prediction
"""
target_h, target_w = f2.shape[2], f2.shape[3]
r2 = self.reduce2(f2)
r3 = self.reduce3(f3)
r4 = self.reduce4(f4)
# Upsample r3, r4 to match r2 spatial resolution
r3 = F.interpolate(r3, size=(target_h, target_w), mode="bilinear", align_corners=True)
r4 = F.interpolate(r4, size=(target_h, target_w), mode="bilinear", align_corners=True)
feat = self.fuse(torch.cat([r2, r3, r4], dim=1))
r_fg = self.fg_head(feat)
r_bg = self.bg_head(feat)
return feat, r_fg, r_bg
# ─── SECTION 4: DSRA Module ────────────────────────────────────────────────────
class DSRAModule(nn.Module):
"""
Dual-Supervised Reverse Attention (DSRA) module.
Introduced in PraNet-V2 (Section 2.1, Eqs. 1–4).
Given encoder feature F_{i+1} and the foreground/background outputs from
the deeper stage (R_{i+1}^F, R_{i+1}^B), computes:
{P_i^f, P_i^b} = phi(F_{i+1}) [Eq. 3]
gamma = Softmax(I(R_{i+1}^F; F_{i+1}) - I(R_{i+1}^B; F_{i+1})) [Eq. 4]
R_i^F = P_i^f + P_i^f * gamma [Eq. 1]
R_i^B = P_i^b [Eq. 2]
Key differences from PraNet-V1 RA:
1. Independent parameters for fg/bg — no shared structure
2. Learned reverse gain (gamma) instead of rule-based complement
3. Fusion in label space (logit space) rather than feature space
4. Per-class support for multi-class segmentation
Parameters
----------
in_channels : channel dimension of F_{i+1}
mid_channels : internal processing channel dimension
num_classes : number of foreground classes
"""
def __init__(
self,
in_channels: int,
mid_channels: int = 64,
num_classes: int = 1,
):
super().__init__()
self.num_classes = num_classes
# Convolutional decoder layers: phi(F_{i+1})
self.decoder_layers = nn.Sequential(
ChannelReducer(in_channels, mid_channels),
ConvBnRelu(mid_channels, mid_channels),
ConvBnRelu(mid_channels, mid_channels),
)
# Foreground segmentation head (produces P_i^f)
self.fg_head = nn.Conv2d(mid_channels, num_classes, kernel_size=1)
# Background segmentation head (produces P_i^b)
# Supervised independently with background mask + BCE loss
self.bg_head = nn.Conv2d(mid_channels, num_classes, kernel_size=1)
def forward(
self,
f_enc: Tensor,
r_fg_deep: Tensor,
r_bg_deep: Tensor,
) -> Tuple[Tensor, Tensor]:
"""
Parameters
----------
f_enc : (B, in_channels, H, W) — encoder feature F_{i+1}
r_fg_deep : (B, num_classes, H_deep, W_deep) — R_{i+1}^F from deeper stage
r_bg_deep : (B, num_classes, H_deep, W_deep) — R_{i+1}^B from deeper stage
Returns
-------
r_fg : (B, num_classes, H, W) — refined foreground output R_i^F
r_bg : (B, num_classes, H, W) — background output R_i^B
"""
B, C_in, H, W = f_enc.shape
# Step 1: Eq. 3 — decode encoder feature into fg/bg logits
feat = self.decoder_layers(f_enc)
p_fg = self.fg_head(feat) # (B, num_classes, H, W)
p_bg = self.bg_head(feat) # (B, num_classes, H, W)
# Step 2: Eq. 4 — compute reverse gain gamma
# Resize deep predictions to match current encoder feature size
r_fg_up = F.interpolate(r_fg_deep, size=(H, W), mode="bilinear", align_corners=True)
r_bg_up = F.interpolate(r_bg_deep, size=(H, W), mode="bilinear", align_corners=True)
# gamma = Softmax(I(R_fg) - I(R_bg)) — channel-wise softmax over class dim
diff = r_fg_up - r_bg_up # (B, num_classes, H, W)
if self.num_classes > 1:
gamma = F.softmax(diff, dim=1)
else:
# Binary case: sigmoid for single channel
gamma = torch.sigmoid(diff)
# Step 3: Eq. 1 — foreground output with reverse gain refinement
r_fg = p_fg + p_fg * gamma # (B, num_classes, H, W)
# Step 4: Eq. 2 — background output (direct from background head)
r_bg = p_bg # (B, num_classes, H, W)
return r_fg, r_bg
# ─── SECTION 5: Full PraNet-V2 Model ─────────────────────────────────────────
class PraNetV2(nn.Module):
"""
PraNet-V2: Full framework with DSRA-based decoder.
Architecture (4 decoder stages):
Encoder → (f1, f2, f3, f4) [4 multi-scale features]
PD → R4 = {R4_f, R4_b} [coarse stage, uses f2+f3+f4]
DSRA3 → R3 = {R3_f, R3_b} [uses f4, conditioned on R4]
DSRA2 → R2 = {R2_f, R2_b} [uses f3, conditioned on R3]
DSRA1 → R1 = {R1_f, R1_b} [uses f2, conditioned on R2] ← main output
Deep supervision applies loss at all four stages during training.
Parameters
----------
config : PraNetV2Config instance
encoder: optional pre-built encoder nn.Module that returns (f1, f2, f3, f4)
"""
def __init__(
self,
config: Optional[PraNetV2Config] = None,
encoder: Optional[nn.Module] = None,
):
super().__init__()
cfg = config or PraNetV2Config()
self.cfg = cfg
C = cfg.decoder_channels
enc_ch = cfg.encoder_channels # [c1, c2, c3, c4]
# ── Encoder ────────────────────────────────────────────────────────────
self.encoder = encoder if encoder is not None else SimpleEncoder(cfg.in_channels)
# ── Parallel Partial Decoder ───────────────────────────────────────────
self.pd = ParallelPartialDecoder(
enc_channels=enc_ch[1:], # [c2, c3, c4]
mid_channels=C,
num_classes=cfg.num_classes,
)
# ── DSRA decoder stages ────────────────────────────────────────────────
# DSRA3: uses f4 (deepest encoder feature), conditioned on R4 from PD
self.dsra3 = DSRAModule(enc_ch[3], mid_channels=C, num_classes=cfg.num_classes)
# DSRA2: uses f3, conditioned on R3 from DSRA3
self.dsra2 = DSRAModule(enc_ch[2], mid_channels=C, num_classes=cfg.num_classes)
# DSRA1: uses f2, conditioned on R2 from DSRA2 — produces final output
self.dsra1 = DSRAModule(enc_ch[1], mid_channels=C, num_classes=cfg.num_classes)
def forward(self, x: Tensor) -> Dict[str, Tensor]:
"""
Forward pass implementing the PraNet-V2 cascade (Algorithm 1 in paper).
Parameters
----------
x : (B, in_channels, H, W)
Returns
-------
dict with keys:
'r1_fg' : (B, num_classes, H, W) — final foreground prediction (primary)
'r1_bg' : (B, num_classes, H, W) — final background prediction
'r2_fg', 'r2_bg' : intermediate predictions for deep supervision
'r3_fg', 'r3_bg' : intermediate predictions for deep supervision
'r4_fg', 'r4_bg' : coarse PD predictions for deep supervision
All outputs are at input resolution (H, W) — upsampled if necessary.
"""
B, _, H_in, W_in = x.shape
# ── Encoder ────────────────────────────────────────────────────────────
f1, f2, f3, f4 = self.encoder(x)
# f1: H/2, f2: H/4, f3: H/8, f4: H/16 (approximate for SimpleEncoder)
# ── Parallel Partial Decoder → coarse stage R4 ────────────────────────
_, r4_fg, r4_bg = self.pd(f2, f3, f4)
# r4_fg/r4_bg: (B, num_classes, H/4, W/4)
# ── DSRA3: f4 + R4 → R3 ───────────────────────────────────────────────
r3_fg, r3_bg = self.dsra3(f4, r4_fg, r4_bg)
# ── DSRA2: f3 + R3 → R2 ───────────────────────────────────────────────
r2_fg, r2_bg = self.dsra2(f3, r3_fg, r3_bg)
# ── DSRA1: f2 + R2 → R1 (final) ──────────────────────────────────────
r1_fg, r1_bg = self.dsra1(f2, r2_fg, r2_bg)
# ── Upsample all outputs to input resolution ──────────────────────────
def up(t):
return F.interpolate(t, size=(H_in, W_in), mode="bilinear", align_corners=True)
return {
"r1_fg": up(r1_fg), # primary output
"r1_bg": up(r1_bg),
"r2_fg": up(r2_fg),
"r2_bg": up(r2_bg),
"r3_fg": up(r3_fg),
"r3_bg": up(r3_bg),
"r4_fg": up(r4_fg),
"r4_bg": up(r4_bg),
}
# ─── SECTION 6: Background Mask Construction ─────────────────────────────────
def build_background_mask(seg_mask: Tensor, num_classes: int) -> Tensor:
"""
Construct a per-class background mask from a segmentation ground truth.
For each class c:
bg_mask[:, c, :, :] = 1 where pixel label != c, else 0
This is the multi-channel background mask described in Section 2.2.1.
A pixel value of 1 indicates background for that class; 0 indicates the object.
Parameters
----------
seg_mask : (B, H, W) long tensor of integer class labels [0, num_classes)
num_classes : number of classes (including background class 0)
Returns
-------
bg_mask : (B, num_classes, H, W) float tensor with values in {0, 1}
"""
B, H, W = seg_mask.shape
# One-hot encode: (B, num_classes, H, W) — 1 where pixel == class
one_hot = F.one_hot(seg_mask.long(), num_classes).permute(0, 3, 1, 2).float()
# Background mask = complement of one-hot: 1 where pixel is NOT that class
bg_mask = 1.0 - one_hot
return bg_mask # (B, num_classes, H, W)
def build_binary_background_mask(binary_mask: Tensor) -> Tensor:
"""
Simpler version for binary segmentation (polyp vs. background).
Parameters
----------
binary_mask : (B, H, W) or (B, 1, H, W) float tensor, values in {0, 1}
Returns
-------
bg_mask : (B, 1, H, W) float tensor — complement of binary_mask
"""
if binary_mask.dim() == 3:
binary_mask = binary_mask.unsqueeze(1)
return 1.0 - binary_mask
# ─── SECTION 7: Loss Functions ────────────────────────────────────────────────
class BinaryDiceLoss(nn.Module):
"""Soft Dice loss for binary segmentation (single channel)."""
def __init__(self, smooth: float = 1e-5):
super().__init__()
self.smooth = smooth
def forward(self, pred: Tensor, target: Tensor) -> Tensor:
"""pred: (B, 1, H, W) logits; target: (B, 1, H, W) or (B, H, W) binary."""
pred_s = torch.sigmoid(pred)
if target.dim() == 3:
target = target.unsqueeze(1)
target = target.float()
p = pred_s.reshape(pred_s.shape[0], -1)
g = target.reshape(target.shape[0], -1)
intersection = (p * g).sum(dim=-1)
denom = p.sum(dim=-1) + g.sum(dim=-1)
dice = (2 * intersection + self.smooth) / (denom + self.smooth)
return 1.0 - dice.mean()
class MultiClassDiceLoss(nn.Module):
"""Soft Dice loss for multi-class segmentation."""
def __init__(self, num_classes: int, smooth: float = 1e-5):
super().__init__()
self.num_classes = num_classes
self.smooth = smooth
def forward(self, pred: Tensor, target: Tensor) -> Tensor:
"""pred: (B, C, H, W) logits; target: (B, H, W) integer labels."""
pred_s = F.softmax(pred, dim=1)
one_hot = F.one_hot(target.long(), self.num_classes).permute(0, 3, 1, 2).float()
p = pred_s.reshape(pred_s.shape[0], self.num_classes, -1)
g = one_hot.reshape(one_hot.shape[0], self.num_classes, -1)
omega = 1.0 / self.num_classes
inter = (p * g).sum(dim=-1)
denom = p.pow(2).sum(dim=-1) + g.pow(2).sum(dim=-1)
dice_pc = (2 * omega * inter) / (denom + self.smooth)
return 1.0 - dice_pc.mean()
class PraNetV2Loss(nn.Module):
"""
Full PraNet-V2 loss: L_total = w1*L_Dice + w2*L_CE + w3*L_BCE
Applied with deep supervision across all four decoder stages.
The primary (finest) prediction is R1; coarser predictions are R2–R4.
Parameters
----------
num_classes : 1 for binary, N for multi-class
weights : (w1, w2, w3) for (Dice, CE, BCE)
stage_weights: weighting for each decoder stage [R1, R2, R3, R4]
"""
def __init__(
self,
num_classes: int = 1,
weights: Tuple[float, ...] = (1.0, 1.0, 0.5),
stage_weights: Tuple[float, ...] = (1.0, 0.8, 0.6, 0.4),
):
super().__init__()
self.num_classes = num_classes
self.w1, self.w2, self.w3 = weights
self.stage_weights = stage_weights
self.binary = (num_classes == 1)
if self.binary:
self.dice_loss = BinaryDiceLoss()
else:
self.dice_loss = MultiClassDiceLoss(num_classes)
self.ce_loss = nn.CrossEntropyLoss()
self.bce_loss = nn.BCEWithLogitsLoss()
def _stage_loss(
self,
r_fg: Tensor,
r_bg: Tensor,
fg_gt: Tensor,
bg_mask: Tensor,
) -> Tensor:
"""
Compute combined loss for a single decoder stage.
Parameters
----------
r_fg : foreground prediction logits (B, num_classes, H, W)
r_bg : background prediction logits (B, num_classes, H, W)
fg_gt : foreground ground truth — (B, H, W) long for multiclass,
(B, 1, H, W) float for binary
bg_mask: per-class background mask (B, num_classes, H, W) float
"""
if self.binary:
# Foreground: Dice + BCE
l_dice = self.dice_loss(r_fg, fg_gt)
l_ce = F.binary_cross_entropy_with_logits(
r_fg, fg_gt.unsqueeze(1).float() if fg_gt.dim() == 3 else fg_gt
)
else:
l_dice = self.dice_loss(r_fg, fg_gt)
l_ce = self.ce_loss(r_fg, fg_gt.long())
# Background: BCE with background mask
l_bce = self.bce_loss(r_bg, bg_mask)
return self.w1 * l_dice + self.w2 * l_ce + self.w3 * l_bce
def forward(
self,
preds: Dict[str, Tensor],
fg_gt: Tensor,
bg_mask: Tensor,
) -> Tensor:
"""
Parameters
----------
preds : output dict from PraNetV2.forward()
fg_gt : (B, H, W) long for multi-class; (B, H, W) float {0,1} for binary
bg_mask : (B, num_classes, H, W) — from build_background_mask()
Returns
-------
total loss scalar
"""
stage_keys = [("r1_fg", "r1_bg"), ("r2_fg", "r2_bg"),
("r3_fg", "r3_bg"), ("r4_fg", "r4_bg")]
total = torch.tensor(0.0, device=fg_gt.device)
for sw, (fk, bk) in zip(self.stage_weights, stage_keys):
stage_l = self._stage_loss(preds[fk], preds[bk], fg_gt, bg_mask)
total = total + sw * stage_l
return total
# ─── SECTION 8: Evaluation Metrics ───────────────────────────────────────────
def compute_dice_binary(pred_logits: Tensor, target: Tensor, eps: float = 1e-5) -> float:
"""
Mean Dice coefficient for binary segmentation.
Parameters
----------
pred_logits : (B, 1, H, W) raw logits
target : (B, H, W) or (B, 1, H, W) binary float tensor
Returns
-------
mean_dice : float in [0, 1]
"""
pred_bin = (torch.sigmoid(pred_logits) > 0.5).float()
if target.dim() == 3:
target = target.unsqueeze(1)
target = target.float()
B = pred_bin.shape[0]
p = pred_bin.reshape(B, -1)
g = target.reshape(B, -1)
inter = (p * g).sum(dim=-1)
denom = p.sum(dim=-1) + g.sum(dim=-1)
dice = (2 * inter + eps) / (denom + eps)
return dice.mean().item()
def compute_iou_binary(pred_logits: Tensor, target: Tensor, eps: float = 1e-5) -> float:
"""Mean IoU for binary segmentation."""
pred_bin = (torch.sigmoid(pred_logits) > 0.5).float()
if target.dim() == 3:
target = target.unsqueeze(1)
target = target.float()
B = pred_bin.shape[0]
p = pred_bin.reshape(B, -1)
g = target.reshape(B, -1)
inter = (p * g).sum(dim=-1)
union = (p + g - p * g).sum(dim=-1)
iou = (inter + eps) / (union + eps)
return iou.mean().item()
class SegMetrics:
"""Accumulates segmentation metrics over an epoch (binary or multi-class)."""
def __init__(self, num_classes: int = 1):
self.num_classes = num_classes
self.reset()
def reset(self):
self.dice_sum = 0.0
self.iou_sum = 0.0
self.count = 0
@torch.no_grad()
def update(self, pred_logits: Tensor, target: Tensor):
if self.num_classes == 1:
self.dice_sum += compute_dice_binary(pred_logits, target)
self.iou_sum += compute_iou_binary(pred_logits, target)
else:
pred_cls = pred_logits.argmax(dim=1)
eps = 1e-5
dice_vals = []
iou_vals = []
for c in range(self.num_classes):
p = (pred_cls == c).float()
g = (target == c).float()
tp = (p * g).sum()
fp = (p * (1 - g)).sum()
fn = ((1 - p) * g).sum()
dice_vals.append((2 * tp + eps) / (2 * tp + fp + fn + eps))
iou_vals.append((tp + eps) / (tp + fp + fn + eps))
self.dice_sum += torch.stack(dice_vals).mean().item()
self.iou_sum += torch.stack(iou_vals).mean().item()
self.count += 1
def result(self) -> Dict[str, float]:
n = max(1, self.count)
return {"mDice": self.dice_sum / n, "mIoU": self.iou_sum / n}
# ─── SECTION 9: Dataset Helpers ──────────────────────────────────────────────
class PolypDummyDataset(Dataset):
"""
Minimal dummy dataset replicating CVC-ClinicDB / Kvasir / ETIS statistics.
Images: 3-channel RGB colonoscopy at 352×352.
Masks: binary float {0, 1}.
Replace with a real loader pointing to:
CVC-ClinicDB : http://www.cvc.uab.es/CVC-Clinic/
Kvasir-SEG : https://datasets.simula.no/kvasir-seg/
ETIS : https://polyp.grand-challenge.org/
"""
def __init__(self, num_samples: int = 64, img_size: int = 352):
self.n = num_samples
self.sz = img_size
def __len__(self): return self.n
def __getitem__(self, idx):
img = torch.randn(3, self.sz, self.sz)
mask = torch.randint(0, 2, (self.sz, self.sz)).float()
return img, mask
class SynapseDummyDataset(Dataset):
"""
Dummy dataset replicating Synapse multi-organ CT statistics.
Images: 1-channel greyscale CT at 352×352.
Masks: integer labels [0, 8] (background + 8 organs).
Replace with a loader pointing to:
https://www.synapse.org/#!Synapse:syn3193805/wiki/217789
"""
def __init__(self, num_samples: int = 32, img_size: int = 352, num_classes: int = 9):
self.n = num_samples; self.sz = img_size; self.nc = num_classes
def __len__(self): return self.n
def __getitem__(self, idx):
img = torch.randn(1, self.sz, self.sz)
mask = torch.randint(0, self.nc, (self.sz, self.sz))
return img, mask
class ACDCDummyDataset(Dataset):
"""
Dummy dataset replicating ACDC cardiac MRI statistics.
Images: 1-channel MRI at 352×352.
Masks: integer labels [0, 3] (background + RV + Myo + LV).
Replace with a loader pointing to:
https://acdc.creatis.insa-lyon.fr/
"""
def __init__(self, num_samples: int = 32, img_size: int = 352):
self.n = num_samples; self.sz = img_size
def __len__(self): return self.n
def __getitem__(self, idx):
img = torch.randn(1, self.sz, self.sz)
mask = torch.randint(0, 4, (self.sz, self.sz))
return img, mask
# ─── SECTION 10: Training Loop ────────────────────────────────────────────────
def train_one_epoch(
model: nn.Module,
loader: DataLoader,
optimizer: torch.optim.Optimizer,
criterion: PraNetV2Loss,
device: torch.device,
epoch: int,
num_classes: int,
) -> float:
model.train()
total_loss = 0.0
for step, (imgs, masks) in enumerate(loader):
imgs = imgs.to(device)
masks = masks.to(device)
# Build background mask from ground truth
if num_classes == 1:
bg_mask = build_binary_background_mask(masks.float())
else:
bg_mask = build_background_mask(masks, num_classes)
optimizer.zero_grad()
preds = model(imgs)
loss = criterion(preds, masks, bg_mask)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
total_loss += loss.item()
if step % 10 == 0:
print(f" Epoch {epoch} | Step {step}/{len(loader)} | Loss {loss.item():.4f}")
return total_loss / len(loader)
@torch.no_grad()
def validate(
model: nn.Module,
loader: DataLoader,
criterion: PraNetV2Loss,
metrics: SegMetrics,
device: torch.device,
num_classes: int,
) -> Tuple[float, Dict]:
model.eval()
metrics.reset()
total_loss = 0.0
for imgs, masks in loader:
imgs = imgs.to(device)
masks = masks.to(device)
if num_classes == 1:
bg_mask = build_binary_background_mask(masks.float())
else:
bg_mask = build_background_mask(masks, num_classes)
preds = model(imgs)
loss = criterion(preds, masks, bg_mask)
total_loss += loss.item()
metrics.update(preds["r1_fg"], masks)
return total_loss / len(loader), metrics.result()
def run_training(
dataset_name: str = "polyp",
epochs: int = 3,
batch_size: int = 2,
lr: float = 1e-4,
device_str: str = "cpu",
):
"""
Minimal training pipeline matching the paper's protocol (Algorithm 1).
Set epochs=30 (polyp) or 150 (Synapse/ACDC) for full training runs.
Use device_str='cuda' for GPU training.
"""
device = torch.device(device_str)
print(f"\n{'='*60}")
print(f" Training PraNet-V2 on {dataset_name.upper()}")
print(f" Device: {device} | Epochs: {epochs} | LR: {lr}")
print(f"{'='*60}\n")
if dataset_name == "polyp":
cfg = PraNetV2Config(num_classes=1, in_channels=3, img_size=352)
train_ds = PolypDummyDataset(num_samples=16)
val_ds = PolypDummyDataset(num_samples=4)
elif dataset_name == "synapse":
cfg = PraNetV2Config(num_classes=9, in_channels=1, img_size=352,
encoder_channels=[64, 256, 512, 1024])
train_ds = SynapseDummyDataset(num_samples=16)
val_ds = SynapseDummyDataset(num_samples=4)
else: # acdc
cfg = PraNetV2Config(num_classes=4, in_channels=1, img_size=352,
encoder_channels=[64, 256, 512, 1024])
train_ds = ACDCDummyDataset(num_samples=16)
val_ds = ACDCDummyDataset(num_samples=4)
train_loader = DataLoader(train_ds, batch_size=batch_size, shuffle=True, num_workers=0)
val_loader = DataLoader(val_ds, batch_size=batch_size, shuffle=False, num_workers=0)
model = PraNetV2(cfg).to(device)
total_params = sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f"Trainable params: {total_params / 1e6:.2f} M\n")
optimizer = torch.optim.Adam(model.parameters(), lr=lr, weight_decay=1e-4)
scheduler = torch.optim.lr_scheduler.CosineAnnealingLR(optimizer, T_max=epochs)
criterion = PraNetV2Loss(num_classes=cfg.num_classes)
metrics = SegMetrics(num_classes=cfg.num_classes)
best_dice = 0.0
for epoch in range(1, epochs + 1):
train_loss = train_one_epoch(
model, train_loader, optimizer, criterion, device, epoch, cfg.num_classes
)
val_loss, res = validate(model, val_loader, criterion, metrics, device, cfg.num_classes)
scheduler.step()
dice = res["mDice"]
print(
f"Epoch {epoch:3d}/{epochs} | Train {train_loss:.4f} | "
f"Val {val_loss:.4f} | mDice {dice:.4f} | mIoU {res['mIoU']:.4f}"
)
if dice > best_dice:
best_dice = dice
print(f" ✓ New best mDice: {best_dice:.4f}")
print(f"\nTraining complete. Best mDice: {best_dice:.4f}")
return model
# ─── SECTION 11: Smoke Test ───────────────────────────────────────────────────
if __name__ == "__main__":
print("=" * 60)
print("PraNet-V2 — Full Architecture Smoke Test")
print("=" * 60)
torch.manual_seed(42)
device = torch.device("cpu")
# ── 1. Binary polyp segmentation (352×352, 1-class) ─────────────────────
print("\n[1/5] Binary polyp — forward pass (352×352, 3ch)...")
cfg1 = PraNetV2Config(num_classes=1, in_channels=3, img_size=352)
model1 = PraNetV2(cfg1).to(device)
x1 = torch.randn(2, 3, 352, 352)
with torch.no_grad():
out1 = model1(x1)
assert out1["r1_fg"].shape == (2, 1, 352, 352), f"Shape mismatch: {out1['r1_fg'].shape}"
print(f" ✓ r1_fg: {tuple(out1['r1_fg'].shape)} r1_bg: {tuple(out1['r1_bg'].shape)}")
# ── 2. Multi-class Synapse (9 classes, 1-channel CT) ─────────────────────
print("\n[2/5] Synapse multi-organ — forward pass (352×352, 1ch, 9 classes)...")
cfg2 = PraNetV2Config(num_classes=9, in_channels=1)
model2 = PraNetV2(cfg2).to(device)
x2 = torch.randn(2, 1, 352, 352)
with torch.no_grad():
out2 = model2(x2)
assert out2["r1_fg"].shape == (2, 9, 352, 352)
print(f" ✓ r1_fg: {tuple(out2['r1_fg'].shape)}")
# ── 3. Background mask construction ─────────────────────────────────────
print("\n[3/5] Background mask construction...")
seg_gt = torch.randint(0, 9, (2, 352, 352))
bg_mask = build_background_mask(seg_gt, num_classes=9)
assert bg_mask.shape == (2, 9, 352, 352)
assert bg_mask.min() >= 0 and bg_mask.max() <= 1
print(f" ✓ bg_mask: {tuple(bg_mask.shape)}, values in [{bg_mask.min():.0f}, {bg_mask.max():.0f}]")
# ── 4. Loss function check ────────────────────────────────────────────────
print("\n[4/5] Loss function check (binary + multi-class)...")
crit_bin = PraNetV2Loss(num_classes=1)
fg_gt_bin = torch.randint(0, 2, (2, 352, 352)).float()
bg_m_bin = build_binary_background_mask(fg_gt_bin)
loss_bin = crit_bin(out1, fg_gt_bin, bg_m_bin)
print(f" Binary loss: {loss_bin.item():.4f}")
crit_mc = PraNetV2Loss(num_classes=9)
loss_mc = crit_mc(out2, seg_gt, bg_mask)
print(f" MultiCls loss: {loss_mc.item():.4f}")
# ── 5. Short training run ────────────────────────────────────────────────
print("\n[5/5] Short 2-epoch training run on dummy polyp data...")
run_training(dataset_name="polyp", epochs=2, batch_size=2)
print("\n" + "=" * 60)
print("✓ All checks passed. PraNet-V2 is ready for use.")
print("=" * 60)
print("""
Next steps:
1. Replace SimpleEncoder with Res2Net50 (binary) or PVTv2-B2 (multi-class):
from timm import create_model
backbone = create_model('pvt_v2_b2', pretrained=True, features_only=True)
2. Point dummy datasets to real data:
CVC-ClinicDB : http://www.cvc.uab.es/CVC-Clinic/
Kvasir-SEG : https://datasets.simula.no/kvasir-seg/
Synapse : https://www.synapse.org/#!Synapse:syn3193805
ACDC : https://acdc.creatis.insa-lyon.fr/
3. Train with paper's settings: Adam (lr=1e-4), 352×352, multi-scale {0.75×,1×,1.25×}
4. Jittor reference code available at:
https://github.com/ai4colonoscopy/PraNet-V2/tree/main/binary_seg/jittor
""")
Read the Full Paper & Access the Code
The complete study — including full per-organ results, qualitative segmentation comparisons, SAM-based model comparisons, and KiTS19 lesion segmentation results — is available on the journal website. Jittor reference code is available on GitHub.
Hu, B.-C., Ji, G.-P., Shao, D., & Fan, D.-P. (2026). PraNet-V2: Dual-supervised reverse attention for medical image segmentation. Computational Visual Media. https://doi.org/10.26599/CVM.2025.9450510
This article is an independent editorial analysis of peer-reviewed research. The PyTorch implementation is an educational adaptation using a lightweight encoder for compatibility. The original authors used Res2Net50 and PVTv2-B2 backbones with ImageNet pretraining and a Jittor framework; refer to the official GitHub repository for exact configurations.
Explore More on AI Trend Blend
If this article sparked your interest, here is more of what we cover across the site — from agricultural AI and precision farming to adversarial robustness, computer vision, and efficient model design.
